Rev-Erb alpha (Rev-Erbɑ), also known as nuclear receptor subfamily 1 group D member 1 (NR1D1), is one of two Rev-Erb proteins in the nuclear receptor (NR) family of intracellular transcription factors. In humans, REV-ERBɑ is encoded by the NR1D1 gene, which is highly conserved across animal species. [5]
Rev-Erbɑ plays an important role in regulation of the core circadian clock through repression of the positive clock element Bmal1. It also regulates several physiological processes under circadian control, including metabolic and immune pathways. [6] [7] Rev-Erbɑ mRNA demonstrates circadian oscillation in its expression, and it is highly expressed in mammals in the brain and metabolic tissues such as skeletal muscle, adipose tissue, and liver. [6] [8]
Discovery
Rev-Erbɑ was discovered in 1989 by Nobuyuki Miyajima and colleagues, who identified two erbA homologs on human chromosome 17 that were transcribed from opposite DNA strands in the same locus. One of the genes encoded a protein that was highly similar to chicken thyroid hormone receptor, and the other, which they termed ear-1, would later be described as Rev-Erbɑ. [9] The protein was first referenced by the name Rev-Erbɑ in 1990 by Mitchell A. Lazar, Karen E. Jones, and William W. Chin, who isolated Rev-Erbɑ complementary DNA from a human fetal skeletal muscle library. Similar to the gene in rats, they found that human Rev-Erbɑ was transcribed from the strand opposite human thyroid hormone receptor alpha (THRA, c-erbAα). [10]
Rev-Erbɑ was first implicated in circadian control in 1998, when Aurelio Balsalobre, Francesca Damiola, and Ueli Schibler demonstrated that expression of Rev-Erbɑ in rat fibroblasts showed daily rhythms. [11] Rev-Erbɑ was first identified as a key player in the transcription translation feedback loop (TTFL) in 2002, when experiments demonstrated that Rev-Erbɑ acted to repress transcription of the Bmal1 gene, and Rev-Erbɑ expression was controlled by other TTFL components. This established Rev-Erbɑ as the link between the positive and negative loops of the TTFL. [12]
Genetics and evolution
The NR1D1 (nuclear receptor subfamily 1 group D member 1) gene, located on chromosome 17, encodes the protein REV-ERBɑ in humans. It is transcribed from the opposite strand of the human thyroid hormone receptor alpha (THRA, c-erbAα) so that NR1D1 and THRA cDNA are complementary on 269 bases. [10] The gene consists of 7,797 bases with 8 exons, forming only 1 splice variant. [5] The NR1D1 promoter itself contains a REV-ERB response element (RevRE), which allows for regulation of gene expression both through autoregulation and regulation by retinoic acid receptor-related orphan receptor alpha (RORɑ), another nuclear receptor transcription factor. [8] NR1D1 also contains an E-box at its promoter, which allows for regulation by BMAL1. In humans, NR1D1 (REV-ERBɑ) is highly expressed in the brain and metabolic tissues, including skeletal muscle, adipose tissue, and the liver. [8] [6]
Genomic analysis suggests that the NR1D1 gene was present in the most recent common ancestor of all animals, with orthologs present in 378 species tested, including chimpanzees, dogs, mice, rats, chickens, zebrafish, frogs, and fruit flies. [13] Comparison to the rat ortholog, Nr1d1, indicates high conservation in the DNA binding and carboxy-terminal domains, as well as conservation of transcription of c-erbA alpha-2 and Rev-Erbɑ on opposite strands. [10] In humans, NR1D1 has only one paralog, NR1D2 ( REV-ERBβ), which is located on chromosome 3 and likely arose from a duplication event. [14] However, both NR1D1 and NR1D2 are members of the nuclear receptor family, indicating they share common ancestry. As such, NR1D1 is functionally related to other nuclear receptor genes, such as peroxisome proliferator activated receptor delta (PPARD) and retinoic acid receptor alpha (RARA). [13] Furthermore, studies have shown that the NR1D1/THRA genetic locus is genetically linked to the RARA gene. [6] [15]
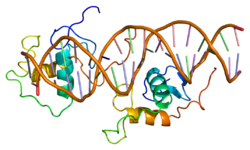
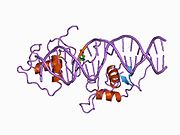
Protein structure
The human NR1D1 gene produces a protein product (REV-ERBα) of 614 amino acids. [5] REV-ERBα has 3 major functional domains, including a DNA-binding domain (DBD) and a ligand-binding domain (LBD) at the C-terminus, and a N-terminus domain which allows for activity modulation. [16] [17] These three domains are a common feature of nuclear receptor proteins. [8]
The Rev-Erb proteins are unique from other nuclear receptors in that they do not have a helix in the C-terminal that is necessary for coactivator recruitment and activation by nuclear receptors via their LBD. [18] Instead, Rev-Erbα interacts via its LBD with Nuclear Receptor Co-Repressor (NCoR) and another closely related co-repressor Silencing Mediator of Retinoid and Thyroid Receptors (SMRT), although the interaction with NCoR is stronger due to its structural compatibility. [18] Heme, an endogenous ligand of Rev-Erbα, further stabilizes the interaction with NCoR. [18] [8] The repression by Rev-Erbα also requires interaction with the class I histone deactylase 3 (HDAC3) - NCoR complex. The catalytic activity of HDAC3 is activated only when it complexes with NCoR or SMRT, so Rev-Erbα must interact with this complex in order for gene repression to occur via histone deacetylation. [6] It is still unknown whether other HDACs play a role in the function of Rev-Erbα. [6] Rev-Erbα recruits the NCoR-HDAC3 complex through binding a specific DNA sequence commonly referred to as RORE due to its interaction with the transcriptional activator Retinoic Acid Receptor-related Orphan Receptor (ROR). This sequence consists of an "AGGTCA" half-site preceded by an A/T sequence.. [18] Rev-Erbα binds in the major groove of this sequence via its DBD domain, which contains two C4-type zinc fingers. [18] Rev-Erbα can repress gene activation as a monomer through competitive binding at this RORE site, but two Rev-Erbα molecules are required for interaction with NCoR and active gene repression. This can occur by two Rev-Erbα molecules binding separate ROREs or as a stronger interaction through binding a response element that is a direct repeat of the RORE (RevDR2). [18]
In mice, it has been shown that the N-terminal regulatory domain contains an important site for phosphorylation by casein kinase 1 epsilon (Csnk1e), which aids in proper localization of Rev-Erbα, and furthermore, that this domain is necessary for activation of the gap junction protein 1 (GJA1) gene. [19] [20]
Function

Circadian oscillator
Rev-Erbα has been proposed to coordinate circadian metabolic responses. [21] Circadian rhythms are driven by interlocking transcription/translation feedback regulatory loops (TTFLs) that generate and maintain these daily rhythms, and Rev-Erbα is involved in a secondary TTFL in mammals. The primary TTFL features transcriptional activator proteins CLOCK and BMAL1 that contribute to the rhythmic expression of genes within this loop, notably per and cry. [22] The expression of these genes then act through negative feedback to inhibit CLOCK:BMAL1 transcription. [12] The secondary TTFL, featuring Rev-Erbα working in conjunction with Rev-Erbβ and the orphan receptor RORα, is thought to strengthen this primary TTFL by further regulating BMAL1. [23] RORα shares the same response elements as Rev-Erbα but exerts opposite effects on gene transcription; BMAL1 expression is repressed by Rev-Erbα and activated by RORα. [24] CLOCK:BMAL1 expression activates the transcription of NR1D1, encoding the Rev-Erbα protein. Increased Rev-Erbα expression in turn, represses transcription of BMAL1, stabilizing the loop. [25] The oscillating expression of RORα and Rev-Erbα in the suprachiasmatic nucleus, the principal circadian timekeeper in mammals, [26] leads to the circadian pattern of BMAL1 expression. The occupancy of the BMAL1 promoter by these two receptors is key for proper timing of the core clock machinery in mammals. [21]
Metabolism
Rev-erbα plays a role in the regulation of whole body metabolism through controlling lipid metabolism, bile acid metabolism, and glucose metabolism. [27] Rev-Erbα relays circadian signals into metabolic and inflammatory regulatory responses and vice versa, although the precise mechanisms underlying this relationship are not entirely understood. [21]
Rev-erbα regulates the expression of liver apolipoproteins, sterol regulatory element binding protein, and the fatty acid elongase elovl3 through its repressional activity [28] [29] [30] In addition, the silencing of Rev-erbα is associated with the reduction of fatty acid synthase, a key regulator of lipogenesis. [30] Rev-erbα deficient mice exhibit dyslipidemia due to elevated triglyceride levels [31] and Rev-erbα polymorphisms in humans have been associated with obesity. [32] Rev-erbα also regulates adipogenesis of white and brown adipocytes. [9] Rev-Erbα transcription is induced during the adipogenic process, and over-expression of Rev-erbα enhances adipogenesis. Researchers have proposed that Rev-erbα's role in adipocyte function may affect the timing of processes such as lipid storage and lipolysis, contributing to long term issues with BMI control. [28] Rev-erbα also regulates bile acid metabolism by indirectly down-regulating Cyp7A1, which encodes the first and rate controlling enzyme of the major bile acid biosynthetic pathway. [21]
Rev-erbα plays both indirect and direct roles in glucose metabolism. BMAL1 heavily influences glucose production and glycogen synthesis, thus through the regulation of BMAL1, Rev-erbα indirectly regulates glucose synthesis. [33] More directly, Rev-erbα's expression in the pancreas regulates the function of α-cells and β-cells, which produce glucagon and insulin, respectively. [34]
Muscle and cartilage
Rev-erbα plays a role in myogenesis through interaction with the transcription complex Nuclear Factor-T. [29] It also represses the expression of genes involved in muscle cell differentiation and is expressed in a circadian manner in mouse skeletal muscle. Loss of Rev-erbα function reduces mitochondrial content and function, leading to an impaired exercise capacity. Over-expression leads to improvement. [34] [30]
This protein has also been implicated in the integrity of cartilage. Out of all known nuclear receptors, Rev-erbα is the most highly expressed in osteoarthritic cartilage. [35] One study found that in patients with osteoarthritis has reduced Rev-erbα levels compared to normal cartilage. [36] Research on rheumatoid arthritis (RA) has implicated the potential for treatment with Rev-erbα agonists to RA patients due to their suppression of bone and cartilage destruction. [37]
Immune system
Rev-erbα contributes to the inflammatory response in mammals. [34] In mouse smooth muscle cells, the protein up-regulates expression of interleukin 6 (IL-6) and cyclooxygenase-2. In humans, it controls the lipopolysaccharide (LPS) induced endotoxic response through repressing toll-like receptor (TLR-4), which triggers the immune response to LPS. [28] [34] In the brain, Rev-erbα deletion causes a disruption in the oscillation of microglial activation and increases the expression of pro-inflammatory transcripts. [19]
Many immune and inflammatory proteins exhibit circadian oscillatory behavior, and research has shown that Rev-erbα deficient mice no longer exhibit these oscillations, notably in IL-6, IL-12, CCL5, and CXCL1, and CCL2. [38] Rev-erbα has also been implicated in the development of group 3 innate lymphoid cells (ILC3), which play a role in regulating intestinal health and are responsible for lymphoid development. REV-ERBα promotes RORγt expression, and RORγt is required for ILC3 expression. Rev-erbα is highly expressed in ILC3 subsets. [39]
Mood and behavior
Rev-erbα has been implicated in the regulation of memory and mood. Rev-erbα knockout mice are deficient in short term, long term, and contextual memories, showing deficits in the function of their hippocampus. [40] In addition, Rev-erbα has been proposed to play a role in the regulation of midbrain dopamine production and mood-related behavior in mice through repression of tyrosine hydroxylase gene transcription. [41] Dopamine related dysfunction is associated with mood disorders, notably major depressive disorder, seasonal affective disorder, and bipolar disorder. Genetic variations in human NR1D1 loci are also associated with bipolar disorder onset. [41]
Rev-erbα has been proposed as a target in the treatment of bipolar disorder through lithium, which indirectly regulates the protein at a post-translational level. Lithium inhibits glycogen synthase kinase (GSK 3β), an enzyme that phosphorylates and stabilizes Rev-erbα. Lithium binding to GSK 3β then destabilizes and alters the function of Rev-erbα. [41] This research has been implicated in the development of therapeutic agents for affective disorders, such as lithium for bipolar disorder. [30]
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000126368 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000020889 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b c "NR1D1 Gene | NR1D1 Protein | NR1D1 Antibody". GeneCards. Retrieved 2021-05-06.
- ^ a b c d e f Yin L, Wu N, Lazar MA (April 2010). "Nuclear receptor Rev-erbalpha: a heme receptor that coordinates circadian rhythm and metabolism". Nuclear Receptor Signaling. 8 (1): e001. doi: 10.1621/nrs.08001. PMC 2858265. PMID 20414452.
- ^ Wang S, Li F, Lin Y, Wu B (2020). "Targeting REV-ERBα for therapeutic purposes: promises and challenges". Theranostics. 10 (9): 4168–4182. doi: 10.7150/thno.43834. PMC 7086371. PMID 32226546.
- ^ a b c d e Burris TP (July 2008). "Nuclear hormone receptors for heme: REV-ERBalpha and REV-ERBbeta are ligand-regulated components of the mammalian clock". Molecular Endocrinology. 22 (7): 1509–20. doi: 10.1210/me.2007-0519. PMC 5419435. PMID 18218725.
- ^ a b Miyajima N, Horiuchi R, Shibuya Y, Fukushige S, Matsubara K, Toyoshima K, Yamamoto T (April 1989). "Two erbA homologs encoding proteins with different T3 binding capacities are transcribed from opposite DNA strands of the same genetic locus". Cell. 57 (1): 31–9. doi: 10.1016/0092-8674(89)90169-4. PMID 2539258. S2CID 19135678.
- ^ a b c Lazar MA, Hodin RA, Darling DS, Chin WW (March 1989). "A novel member of the thyroid/steroid hormone receptor family is encoded by the opposite strand of the rat c-erbA alpha transcriptional unit". Molecular and Cellular Biology. 9 (3): 1128–36. doi: 10.1128/MCB.9.3.1128. PMC 362703. PMID 2542765.
- ^ Balsalobre A, Damiola F, Schibler U (June 1998). "A serum shock induces circadian gene expression in mammalian tissue culture cells". Cell. 93 (6): 929–37. doi: 10.1016/s0092-8674(00)81199-x. PMID 9635423. S2CID 12445337.
- ^ a b Preitner N, Damiola F, Lopez-Molina L, Zakany J, Duboule D, Albrecht U, Schibler U (July 2002). "The orphan nuclear receptor REV-ERBalpha controls circadian transcription within the positive limb of the mammalian circadian oscillator". Cell. 110 (2): 251–60. doi: 10.1016/s0092-8674(02)00825-5. PMID 12150932. S2CID 15224136.
- ^ a b Yates AD, Achuthan P, Akanni W, Allen J, Allen J, Alvarez-Jarreta J, et al. (January 2020). "Ensembl 2020". Nucleic Acids Research. 48 (D1): D682–D688. doi: 10.1093/nar/gkz966. PMC 7145704. PMID 31691826.
- ^ Koh YS, Moore DD (April 1999). "Linkage of the nuclear hormone receptor genes NR1D2, THRB, and RARB: evidence for an ancient, large-scale duplication". Genomics. 57 (2): 289–92. doi: 10.1006/geno.1998.5683. PMID 10198169.
- ^ Kainu, Tommi; Enmark, Eva; Gustafsson, Jan-Åke; Pelto-Huikko, Markku (1996-12-16). "Localization of the Rev-ErbA orphan receptors in the brain". Brain Research. 743 (1–2): 315–319. doi: 10.1016/S0006-8993(96)00507-0. ISSN 0006-8993. PMID 9017260. S2CID 43554925.
- ^ Bateman, Alex; et al. (January 2021). "UniProt: the universal protein knowledgebase in 2021". Nucleic Acids Research. 49 (D1): D480–D489. doi: 10.1093/nar/gkaa1100. PMC 7778908. PMID 33237286.
- ^ "HomoloGene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2021-05-06.
- ^ a b c d e f Everett LJ, Lazar MA (November 2014). "Nuclear receptor Rev-erbα: up, down, and all around". Trends in Endocrinology and Metabolism. 25 (11): 586–92. doi: 10.1016/j.tem.2014.06.011. PMC 4252361. PMID 25066191.
- ^ a b Griffin P, Dimitry JM, Sheehan PW, Lananna BV, Guo C, Robinette ML, et al. (March 2019). "Circadian clock protein Rev-erbα regulates neuroinflammation". Proceedings of the National Academy of Sciences of the United States of America. 116 (11): 5102–5107. Bibcode: 2019PNAS..116.5102G. doi: 10.1073/pnas.1812405116. PMC 6421453. PMID 30792350.
- ^ Negoro H, Okinami T, Kanematsu A, Imamura M, Tabata Y, Ogawa O (January 2013). "Role of Rev-erbα domains for transactivation of the connexin43 promoter with Sp1". FEBS Letters. 587 (1): 98–103. Bibcode: 2013FEBSL.587...98N. doi: 10.1016/j.febslet.2012.11.021. hdl: 2433/168617. PMID 23201262. S2CID 30249508.
- ^ a b c d Duez H, Staels B (December 2009). "Rev-erb-alpha: an integrator of circadian rhythms and metabolism". Journal of Applied Physiology. 107 (6): 1972–80. doi: 10.1152/japplphysiol.00570.2009. PMC 2966474. PMID 19696364.
- ^ Andreani TS, Itoh TQ, Yildirim E, Hwangbo DS, Allada R (December 2015). "Genetics of Circadian Rhythms". Sleep Medicine Clinics. 10 (4): 413–21. doi: 10.1016/j.jsmc.2015.08.007. PMC 4758938. PMID 26568119.
- ^ Guillaumond F, Dardente H, Giguère V, Cermakian N (October 2005). "Differential control of Bmal1 circadian transcription by REV-ERB and ROR nuclear receptors". Journal of Biological Rhythms. 20 (5): 391–403. doi: 10.1177/0748730405277232. PMID 16267379. S2CID 33279857.
- ^ Solt LA, Kojetin DJ, Burris TP (April 2011). "The REV-ERBs and RORs: molecular links between circadian rhythms and lipid homeostasis". Future Medicinal Chemistry. 3 (5): 623–38. doi: 10.4155/fmc.11.9. PMC 3134326. PMID 21526899.
- ^ Oishi Y, Hayashi S, Isagawa T, Oshima M, Iwama A, Shimba S, et al. (August 2017). "Bmal1 regulates inflammatory responses in macrophages by modulating enhancer RNA transcription". Scientific Reports. 7 (1): 7086. Bibcode: 2017NatSR...7.7086O. doi: 10.1038/s41598-017-07100-3. PMC 5539165. PMID 28765524.
- ^ Hastings MH, Maywood ES, Brancaccio M (August 2018). "Generation of circadian rhythms in the suprachiasmatic nucleus". Nature Reviews. Neuroscience. 19 (8): 453–469. doi: 10.1038/s41583-018-0026-z. PMID 29934559. S2CID 49357675.
- ^ Duez H, Staels B (January 2008). "Rev-erb alpha gives a time cue to metabolism". FEBS Letters. 582 (1): 19–25. Bibcode: 2008FEBSL.582...19D. doi: 10.1016/j.febslet.2007.08.032. PMID 17765229. S2CID 84204023.
- ^ a b c Duez H, Staels B (August 2010). "Nuclear receptors linking circadian rhythms and cardiometabolic control". Arteriosclerosis, Thrombosis, and Vascular Biology. 30 (8): 1529–34. doi: 10.1161/ATVBAHA.110.209098. PMC 3056213. PMID 20631353.
- ^ a b Welch RD, Billon C, Kameric A, Burris TP, Flaveny CA (2020-05-14). "Rev-erbα heterozygosity produces a dose-dependent phenotypic advantage in mice". PLOS ONE. 15 (5): e0227720. Bibcode: 2020PLoSO..1527720W. doi: 10.1371/journal.pone.0227720. PMC 7224546. PMID 32407314.
- ^ a b c d Marciano DP, Chang MR, Corzo CA, Goswami D, Lam VQ, Pascal BD, Griffin PR (February 2014). "The therapeutic potential of nuclear receptor modulators for treatment of metabolic disorders: PPARγ, RORs, and Rev-erbs". Cell Metabolism. 19 (2): 193–208. doi: 10.1016/j.cmet.2013.12.009. PMID 24440037.
- ^ Raspé E, Duez H, Mansén A, Fontaine C, Fiévet C, Fruchart JC, et al. (December 2002). "Identification of Rev-erbalpha as a physiological repressor of apoC-III gene transcription". Journal of Lipid Research. 43 (12): 2172–9. doi: 10.1194/jlr.M200386-JLR200. PMID 12454280.
- ^ Ruano EG, Canivell S, Vieira E (2014-08-04). "REV-ERB ALPHA polymorphism is associated with obesity in the Spanish obese male population". PLOS ONE. 9 (8): e104065. Bibcode: 2014PLoSO...9j4065R. doi: 10.1371/journal.pone.0104065. PMC 4121274. PMID 25089907.
- ^ Kalsbeek A, la Fleur S, Fliers E (July 2014). "Circadian control of glucose metabolism". Molecular Metabolism. 3 (4): 372–83. doi: 10.1016/j.molmet.2014.03.002. PMC 4060304. PMID 24944897.
- ^ a b c d Lazar MA (2016). "Rev-erbs: Integrating Metabolism Around the Clock". In Sassone-Corsi P, Christen Y (eds.). A Time for Metabolism and Hormones. Research and Perspectives in Endocrine Interactions. Cham: Springer International Publishing. pp. 63–70. doi: 10.1007/978-3-319-27069-2_7. ISBN 978-3-319-27068-5. PMID 28892343.
- ^ Marks R (2018). "Circadian Clock: Potential Role in Cartilage Integrity and Disruption". International Journal of Orthopaedics. 5 (4): 936–942. doi: 10.17554/j.issn.2311-5106.2018.05.280. ISSN 2311-5106.
- ^ Yin L, Lazar MA (June 2005). "The orphan nuclear receptor Rev-erbalpha recruits the N-CoR/histone deacetylase 3 corepressor to regulate the circadian Bmal1 gene". Molecular Endocrinology. 19 (6): 1452–9. doi: 10.1210/me.2005-0057. PMID 15761026.
- ^ Fontaine C, Dubois G, Duguay Y, Helledie T, Vu-Dac N, Gervois P, et al. (September 2003). "The orphan nuclear receptor Rev-Erbalpha is a peroxisome proliferator-activated receptor (PPAR) gamma target gene and promotes PPARgamma-induced adipocyte differentiation". The Journal of Biological Chemistry. 278 (39): 37672–80. doi: 10.1074/jbc.M304664200. PMID 12821652. S2CID 82056456.
- ^ Scheiermann C, Kunisaki Y, Frenette PS (March 2013). "Circadian control of the immune system". Nature Reviews. Immunology. 13 (3): 190–8. doi: 10.1038/nri3386. PMC 4090048. PMID 23391992.
- ^ Wang Q, Robinette ML, Billon C, Collins PL, Bando JK, Fachi JL, et al. (October 2019). "Circadian rhythm-dependent and circadian rhythm-independent impacts of the molecular clock on type 3 innate lymphoid cells". Science Immunology. 4 (40): eaay7501. doi: 10.1126/sciimmunol.aay7501. PMC 6911370. PMID 31586012.
- ^ Jager J, O'Brien WT, Manlove J, Krizman EN, Fang B, Gerhart-Hines Z, et al. (April 2014). "Behavioral changes and dopaminergic dysregulation in mice lacking the nuclear receptor Rev-erbα". Molecular Endocrinology. 28 (4): 490–8. doi: 10.1210/me.2013-1351. PMC 3968406. PMID 24552589.
- ^ a b c Chung S, Lee EJ, Yun S, Choe HK, Park SB, Son HJ, et al. (May 2014). "Impact of circadian nuclear receptor REV-ERBα on midbrain dopamine production and mood regulation". Cell. 157 (4): 858–68. doi: 10.1016/j.cell.2014.03.039. PMID 24813609. S2CID 3334962.
Further reading
- Laudet V, Begue A, Henry-Duthoit C, Joubel A, Martin P, Stehelin D, Saule S (Mar 1991). "Genomic organization of the human thyroid hormone receptor alpha (c-erbA-1) gene". Nucleic Acids Research. 19 (5): 1105–12. doi: 10.1093/nar/19.5.1105. PMC 333788. PMID 1850510.
- Miyajima N, Horiuchi R, Shibuya Y, Fukushige S, Matsubara K, Toyoshima K, Yamamoto T (Apr 1989). "Two erbA homologs encoding proteins with different T3 binding capacities are transcribed from opposite DNA strands of the same genetic locus". Cell. 57 (1): 31–9. doi: 10.1016/0092-8674(89)90169-4. PMID 2539258. S2CID 19135678.
- Adelmant G, Bègue A, Stéhelin D, Laudet V (Apr 1996). "A functional Rev-erb alpha responsive element located in the human Rev-erb alpha promoter mediates a repressing activity". Proceedings of the National Academy of Sciences of the United States of America. 93 (8): 3553–8. Bibcode: 1996PNAS...93.3553A. doi: 10.1073/pnas.93.8.3553. PMC 39648. PMID 8622974.
- Downes M, Burke LJ, Bailey PJ, Muscat GE (Nov 1996). "Two receptor interaction domains in the corepressor, N-CoR/RIP13, are required for an efficient interaction with Rev-erbA alpha and RVR: physical association is dependent on the E region of the orphan receptors". Nucleic Acids Research. 24 (22): 4379–86. doi: 10.1093/nar/24.22.4379. PMC 146280. PMID 8948627.
- Burke LJ, Downes M, Laudet V, Muscat GE (Feb 1998). "Identification and characterization of a novel corepressor interaction region in RVR and Rev-erbA alpha". Molecular Endocrinology. 12 (2): 248–62. doi: 10.1210/mend.12.2.0061. PMID 9482666.
- Zhao Q, Khorasanizadeh S, Miyoshi Y, Lazar MA, Rastinejad F (May 1998). "Structural elements of an orphan nuclear receptor-DNA complex". Molecular Cell. 1 (6): 849–61. doi: 10.1016/S1097-2765(00)80084-2. PMID 9660968.
- Sierk ML, Zhao Q, Rastinejad F (Oct 2001). "DNA deformability as a recognition feature in the reverb response element". Biochemistry. 40 (43): 12833–43. doi: 10.1021/bi011086r. PMID 11669620.
- Coste H, Rodríguez JC (Jul 2002). "Orphan nuclear hormone receptor Rev-erbalpha regulates the human apolipoprotein CIII promoter". The Journal of Biological Chemistry. 277 (30): 27120–9. doi: 10.1074/jbc.M203421200. hdl: 2445/105711. PMID 12021280.
- Delerive P, Chin WW, Suen CS (Sep 2002). "Identification of Reverb(alpha) as a novel ROR(alpha) target gene". The Journal of Biological Chemistry. 277 (38): 35013–8. doi: 10.1074/jbc.M202979200. PMID 12114512.
- Raspè E, Mautino G, Duval C, Fontaine C, Duez H, Barbier O, Monte D, Fruchart J, Fruchart JC, Staels B (Dec 2002). "Transcriptional regulation of human Rev-erbalpha gene expression by the orphan nuclear receptor retinoic acid-related orphan receptor alpha". The Journal of Biological Chemistry. 277 (51): 49275–81. doi: 10.1074/jbc.M206215200. PMID 12377782.
- Raspé E, Duez H, Mansén A, Fontaine C, Fiévet C, Fruchart JC, Vennström B, Staels B (Dec 2002). "Identification of Rev-erbalpha as a physiological repressor of apoC-III gene transcription". Journal of Lipid Research. 43 (12): 2172–9. doi: 10.1194/jlr.M200386-JLR200. PMID 12454280.
- Chopin-Delannoy S, Thénot S, Delaunay F, Buisine E, Begue A, Duterque-Coquillaud M, Laudet V (Apr 2003). "A specific and unusual nuclear localization signal in the DNA binding domain of the Rev-erb orphan receptors". Journal of Molecular Endocrinology. 30 (2): 197–211. doi: 10.1677/jme.0.0300197. PMID 12683943.
- Fontaine C, Dubois G, Duguay Y, Helledie T, Vu-Dac N, Gervois P, Soncin F, Mandrup S, Fruchart JC, Fruchart-Najib J, Staels B (Sep 2003). "The orphan nuclear receptor Rev-Erbalpha is a peroxisome proliferator-activated receptor (PPAR) gamma target gene and promotes PPARgamma-induced adipocyte differentiation". The Journal of Biological Chemistry. 278 (39): 37672–80. doi: 10.1074/jbc.M304664200. PMID 12821652.
- Johnson JM, Castle J, Garrett-Engele P, Kan Z, Loerch PM, Armour CD, Santos R, Schadt EE, Stoughton R, Shoemaker DD (Dec 2003). "Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays". Science. 302 (5653): 2141–4. Bibcode: 2003Sci...302.2141J. CiteSeerX 10.1.1.1017.9438. doi: 10.1126/science.1090100. PMID 14684825. S2CID 10007258.
- Migita H, Morser J, Kawai K (Mar 2004). "Rev-erbalpha upregulates NF-kappaB-responsive genes in vascular smooth muscle cells". FEBS Letters. 561 (1–3): 69–74. Bibcode: 2004FEBSL.561...69M. doi: 10.1016/S0014-5793(04)00118-8. PMID 15013753. S2CID 84456190.
- Cheng H, Khanna H, Oh EC, Hicks D, Mitton KP, Swaroop A (Aug 2004). "Photoreceptor-specific nuclear receptor NR2E3 functions as a transcriptional activator in rod photoreceptors". Human Molecular Genetics. 13 (15): 1563–75. doi: 10.1093/hmg/ddh173. PMID 15190009.
- Beausoleil SA, Jedrychowski M, Schwartz D, Elias JE, Villén J, Li J, Cohn MA, Cantley LC, Gygi SP (Aug 2004). "Large-scale characterization of HeLa cell nuclear phosphoproteins". Proceedings of the National Academy of Sciences of the United States of America. 101 (33): 12130–5. Bibcode: 2004PNAS..10112130B. doi: 10.1073/pnas.0404720101. PMC 514446. PMID 15302935.
External links
- NR1D1+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)