| interleukin 2 receptor, alpha chain | |||||||
|---|---|---|---|---|---|---|---|
 | |||||||
| Identifiers | |||||||
| Symbol | IL2RA | ||||||
| Alt. symbols | IL2R CD25 | ||||||
| NCBI gene | 3559 | ||||||
| HGNC | 6008 | ||||||
| OMIM | 147730 | ||||||
| RefSeq | NM_000417 | ||||||
| UniProt | P01589 | ||||||
| Other data | |||||||
| Locus | Chr. 10 p15.1 | ||||||
| |||||||
| interleukin 2 receptor, beta chain | |||||||
|---|---|---|---|---|---|---|---|
 | |||||||
| Identifiers | |||||||
| Symbol | IL2RB | ||||||
| Alt. symbols | CD122 | ||||||
| NCBI gene | 3560 | ||||||
| HGNC | 6009 | ||||||
| OMIM | 146710 | ||||||
| RefSeq | NM_000878 | ||||||
| UniProt | P14784 | ||||||
| Other data | |||||||
| Locus | Chr. 22 q13 | ||||||
| |||||||
| interleukin 2 receptor, gamma chain ( severe combined immunodeficiency) | |||||||
|---|---|---|---|---|---|---|---|
 | |||||||
| Identifiers | |||||||
| Symbol | IL2RG | ||||||
| Alt. symbols | SCIDX1, IMD4, CD132 | ||||||
| NCBI gene | 3561 | ||||||
| HGNC | 6010 | ||||||
| OMIM | 308380 | ||||||
| RefSeq | NM_000206 | ||||||
| UniProt | P31785 | ||||||
| Other data | |||||||
| Locus | Chr. X q13 | ||||||
| |||||||
The interleukin-2 receptor (IL-2R) is a heterotrimeric protein expressed on the surface of certain immune cells, such as lymphocytes, that binds and responds to a cytokine called IL-2.
Composition
IL-2 binds to the IL-2 receptor, which has three forms, generated by different combinations of three different proteins, often referred to as "chains": α (alpha) (also called IL-2Rα, CD25, or Tac antigen), β (beta) (also called IL-2Rβ, or CD122), and γ (gamma) (also called IL-2Rγ, γc, common gamma chain, or CD132); these subunits are also parts of receptors for other cytokines. [2]: 713 The β and γ chains of the IL-2R are members of the type I cytokine receptor family. [3]
Structure-activity relationships of the IL-2/IL-2R interaction
The three receptor chains are expressed separately and differently on various cell types and can assemble in different combinations and orders to generate low, intermediate, and high affinity IL-2 receptors.
The α chain binds IL-2 with low affinity, the combination of β and γ together form a complex that binds IL-2 with intermediate affinity, primarily on memory T cells and NK cells; and all three receptor chains form a complex that binds IL-2 with high affinity (Kd ~ 10−11 M) on activated T cells and regulatory T cells. The intermediate and high affinity receptor forms are functional and cause changes in the cell when IL-2 binds to them. [3]
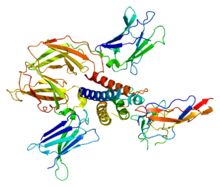
The structure of the stable complex formed when IL-2 binds to the high affinity receptor has been determined using X-ray crystallography. The structure supports a model wherein IL-2 initially binds to the α chain, then the β is recruited, and finally γ. [3] [4] [5]
Signaling
The three IL-2 receptor chains span the cell membrane and extend into the cell, thereby delivering biochemical signals to the cell interior. The alpha chain does not participate in signaling, but the beta chain is complexed with an enzyme called Janus kinase 1 (JAK1), that is capable of adding phosphate groups to molecules. Similarly the gamma chain complexes with another tyrosine kinase called JAK3. These enzymes are activated by IL-2 binding to the external domains of the IL-2R. As a consequence, three intracellular signaling pathways are initiated, the MAP kinase pathway, the Phosphoinositide 3-kinase (PI3K) pathway, and the JAK-STAT pathway. [3] [4]
Once IL-2 binds to the high affinity receptor, the complex is rapidly internalized and has only a short time to signal. IL-2, IL-2Rβ, and γc are rapidly degraded, but IL-2Rα is recycled to the cell surface. Thus, the concentration of IL-2 and its receptor available determines the tempo, magnitude and extent of T cell immune responses. [3] [4]
IL-2 and its receptor have key roles in key functions of the immune system, tolerance and immunity, primarily via their direct effects on T cells. In the thymus, where T cells mature, they prevent autoimmune diseases by promoting the differentiation of certain immature T cells into regulatory T cells, which kill off other T cells that are primed to attack normal healthy cells in the body. IL-2/IL2R also promotes the differentiation of T cells into effector T cells and into memory T cells when the initial T cells is also stimulated by an antigen, thus helping the body fight off infections. [3] Through their role in the development of T cell immunologic memory, which depends upon the expansion of the number and function of antigen-selected T cell clones, they also have a key role in enduring cell-mediated immunity. [3] [4]
Clinical implications
Drugs that inhibit IL-2 receptors, such as basiliximab and daclizumab are used in conjunction with other drugs to prevent immune rejection of transplants. [6]
History
According to an immunology textbook: "IL-2 is particularly important historically, as it is the first type I cytokine that was cloned, the first type I cytokine for which a receptor component was cloned, and was the first short-chain type I cytokine whose receptor structure was solved. Many general principles have been derived from studies of this cytokine, including its being the first cytokine demonstrated to act in a growth factor–like fashion through specific high-affinity receptors, analogous to the growth factors being studied by endocrinologists and biochemists". [2]: 712
See also
References
- ^ PDB: 2B5IWang X, Rickert M, Garcia KC (November 2005). "Structure of the quaternary complex of interleukin-2 with its alpha, beta, and gammac receptors". Science. 310 (5751): 1159–63. Bibcode: 2005Sci...310.1159W. doi: 10.1126/science.1117893. PMID 16293754. S2CID 85394260.
- ^ a b Leonard WJ (2008). "Chapter 23: Type I Cytokines and Interferons and Their Receptors.". In Paul WE (ed.). Fundamental Immunology (6th ed.). Philadelphia: Wolters Kluwer/Lippincott Williams & Wilkins. ISBN 9780781765190.
- ^ a b c d e f g Liao W, Lin JX, Leonard WJ (October 2011). "IL-2 family cytokines: new insights into the complex roles of IL-2 as a broad regulator of T helper cell differentiation". Current Opinion in Immunology. 23 (5): 598–604. doi: 10.1016/j.coi.2011.08.003. PMC 3405730. PMID 21889323.
- ^ a b c d Malek TR, Castro I (August 2010). "Interleukin-2 receptor signaling: at the interface between tolerance and immunity". Immunity. 33 (2): 153–65. doi: 10.1016/j.immuni.2010.08.004. PMC 2946796. PMID 20732639.
- ^ Metz A, Ciglia E, Gohlke H (2012). "Modulating protein-protein interactions: from structural determinants of binding to druggability prediction to application". Current Pharmaceutical Design. 18 (30): 4630–47. doi: 10.2174/138161212802651553. PMID 22650257.
- ^ Hardinger KL, Brennan DC, Klein CL (July 2013). "Selection of induction therapy in kidney transplantation". Transplant International. 26 (7): 662–72. doi: 10.1111/tri.12043. PMID 23279211. S2CID 3296555.
External links
- Interleukin-2+Receptors at the U.S. National Library of Medicine Medical Subject Headings (MeSH)