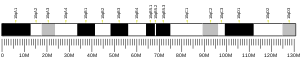
Ankyrin-3 (ANK-3), also known as ankyrin-G, is a protein from ankyrin family that in humans is encoded by the ANK3 gene. [5] [6]
Function
The protein encoded by this gene, ankyrin-3 is an immunologically distinct gene product from ankyrins ANK1 and ANK2, and was originally found at the axonal initial segment and nodes of Ranvier of neurons in the central and peripheral nervous systems. Alternatively spliced variants may be expressed in other tissues. Although multiple transcript variants encoding several different isoforms have been found for this gene, the full-length nature of only two have been characterized. [5]
Within the nervous system, ankyrin-G is specifically localized to the neuromuscular junction, the axon initial segment and the Nodes of Ranvier. [7] Within the nodes of Ranvier where action potentials are actively propagated, ankyrin-G has long been thought to be the intermediate binding partner to neurofascin and voltage-gated sodium channels. [8] The genetic deletion of ankyrin-G from multiple neuron types has shown that ankyrin-G is required for the normal clustering of voltage-gated sodium channels at the axon hillock and for action potential firing. [9] [10]
Disease linkage
The ANK3 protein associates with the cardiac sodium channel Nav1.5 ( SCN5A). Both proteins are highly expressed at ventricular intercalated disc and T-tubule membranes in cardiomyocytes. A mutation in the Nav1.5 protein blocks interaction with ANK3 binding and therefore disrupts surface expression of Nav1.5 in cardiomyocytes resulting in Brugada syndrome, a type of cardiac arrhythmia. [11]
Other mutations in the ANK3 gene may be involved in the bipolar disorder and intellectual disability. [12] [13] [14] [15]
Ankyrin family
The protein encoded by the ANK3 gene is a member of the ankyrin family of proteins that link the integral membrane proteins to the underlying spectrin- actin cytoskeleton. Ankyrins play key roles in activities such as cell motility, activation, proliferation, contact and the maintenance of specialized membrane domains. Most ankyrins are typically composed of three structural domains: an amino-terminal domain containing multiple ankyrin repeats; a central region with a highly conserved spectrin binding domain; and a carboxy-terminal regulatory domain which is the least conserved and subject to variation. [5]
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000151150 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000069601 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b c "Entrez Gene: ANK2 ankyrin 3, node of Ranvier".
- ^ Kapfhamer D, Miller DE, Lambert S, Bennett V, Glover TW, Burmeister M (May 1995). "Chromosomal localization of the ankyrin-G gene (ANK3/Ank3) to human 10q21 and mouse 10". Genomics. 27 (1): 189–91. doi: 10.1006/geno.1995.1023. PMID 7665168.
- ^ Lambert S, Davis JQ, Bennett V (September 1997). "Morphogenesis of the node of Ranvier: co-clusters of ankyrin and ankyrin-binding integral proteins define early developmental intermediates". J. Neurosci. 17 (18): 7025–36. doi: 10.1523/JNEUROSCI.17-18-07025.1997. PMC 6573274. PMID 9278538.
- ^ Srinivasan Y, Lewallen M, Angelides KJ (April 1992). "Mapping the binding site on ankyrin for the voltage-dependent sodium channel from brain". J Biol Chem. 267 (11): 7483–9. doi: 10.1016/S0021-9258(18)42543-4. PMID 1313804.
- ^ Zhou D, Lambert S, Malen PL, Carpenter S, Boland LM, Bennett V (November 1998). "AnkyrinG Is Required for Clustering of Voltage-gated Na Channels at Axon Initial Segments and for Normal Action Potential Firing". J. Cell Biol. 143 (5): 1295–1304. doi: 10.1083/jcb.143.5.1295. PMC 2133082. PMID 9832557.
- ^ Hedstrom KL, Xu X, Ogawa Y, Frischknecht R, Seidenbecher CI, Shrager P, Rasband MN (August 2007). "Neurofascin assembles a specialized extracellular matrix at the axon initial segment". J. Cell Biol. 178 (5): 875–886. doi: 10.1083/jcb.200705119. PMC 2064550. PMID 17709431.
- ^ Mohler PJ, Rivolta I, Napolitano C, LeMaillet G, Lambert S, Priori SG, Bennett V (December 2004). "Nav1.5 E1053K mutation causing Brugada syndrome blocks binding to ankyrin-G and expression of Nav1.5 on the surface of cardiomyocytes". Proc. Natl. Acad. Sci. U.S.A. 101 (50): 17533–8. Bibcode: 2004PNAS..10117533M. doi: 10.1073/pnas.0403711101. PMC 536011. PMID 15579534.
- ^ "Bipolar Disorder Discovery at the Nano Level". Archived from the original on 2014-12-04. Retrieved 2014-12-01.
- ^ Ferreira MA, O'Donovan MC, Meng YA, et al. (August 2008). "Collaborative genome-wide association analysis supports a role for ANK3 and CACNA1C in bipolar disorder". Nat. Genet. 40 (9): 1056–8. doi: 10.1038/ng.209. PMC 2703780. PMID 18711365.
- ^ "Channeling Mental Illness: GWAS Links Ion Channels, Bipolar Disorder". Schizophrenia Research Forum: News. schizophreniaforum.org. 2008-08-19. Archived from the original on 2010-12-18. Retrieved 2008-08-21.
- ^ Iqbal, Zafar; Vandeweyer, Geert; van der Voet, Monique; Waryah, Ali Muhammad; Zahoor, Muhammad Yasir; Besseling, Judith A.; Roca, Laura Tomas; Vulto-van Silfhout, Anneke T.; Nijhof, Bonnie; Kramer, Jamie M.; Van der Aa, Nathalie; Ansar, Muhammad; Peeters, Hilde; Helsmoortel, Celine; Gilissen, Christian; Vissers, Lisenka; Veltman, Joris A.; de Brouwer, Arjan P. M.; Kooy, R. Frank; Riazuddin, Sheikh; Schenck, Annette; van Bokhoven, Hans; Rooms, Liesbeth (2013). "Homozygous and heterozygous disruptions of ANK3: at the crossroads of neurodevelopmental and psychiatric disorders". Human Molecular Genetics. 22 (10): 1960–1970. doi: 10.1093/hmg/ddt043. ISSN 0964-6906. PMID 23390136.
Further reading
- Lopez C, Métral S, Eladari D, et al. (2005). "The ammonium transporter RhBG: requirement of a tyrosine-based signal and ankyrin-G for basolateral targeting and membrane anchorage in polarized kidney epithelial cells". J. Biol. Chem. 280 (9): 8221–8. doi: 10.1074/jbc.M413351200. PMID 15611082.
- Kizhatil K, Yoon W, Mohler PJ, et al. (2007). "Ankyrin-G and beta2-spectrin collaborate in biogenesis of lateral membrane of human bronchial epithelial cells". J. Biol. Chem. 282 (3): 2029–37. doi: 10.1074/jbc.M608921200. PMID 17074766.
- Weimer JM, Chattopadhyay S, Custer AW, Pearce DA (2005). "Elevation of Hook1 in a disease model of Batten disease does not affect a novel interaction between Ankyrin G and Hook1". Biochem. Biophys. Res. Commun. 330 (4): 1176–81. doi: 10.1016/j.bbrc.2005.03.103. PMID 15823567.
- Shirahata E, Iwasaki H, Takagi M, et al. (2006). "Ankyrin-G regulates inactivation gating of the neuronal sodium channel, Nav1.6". J. Neurophysiol. 96 (3): 1347–57. doi: 10.1152/jn.01264.2005. PMID 16775201.
- Schulze TG, Detera-Wadleigh SD, Akula N, et al. (2009). "Two variants in Ankyrin 3 (ANK3) are independent genetic risk factors for bipolar disorder". Mol. Psychiatry. 14 (5): 487–91. doi: 10.1038/mp.2008.134. PMC 2793269. PMID 19088739.
- Morgan AR, Turic D, Jehu L, et al. (2007). "Association studies of 23 positional/functional candidate genes on chromosome 10 in late-onset Alzheimer's disease". Am. J. Med. Genet. B Neuropsychiatr. Genet. 144B (6): 762–70. doi: 10.1002/ajmg.b.30509. PMID 17373700. S2CID 26081707.
- Morgan AR, Hamilton G, Turic D, et al. (2008). "Association analysis of 528 intra-genic SNPs in a region of chromosome 10 linked to late onset Alzheimer's disease". Am. J. Med. Genet. B Neuropsychiatr. Genet. 147B (6): 727–31. doi: 10.1002/ajmg.b.30670. PMID 18163421. S2CID 13916214.
- Grupe A, Li Y, Rowland C, et al. (2006). "A Scan of Chromosome 10 Identifies a Novel Locus Showing Strong Association with Late-Onset Alzheimer Disease". Am. J. Hum. Genet. 78 (1): 78–88. doi: 10.1086/498851. PMC 1380225. PMID 16385451.
- Stabach PR, Devarajan P, Stankewich MC, et al. (2008). "Ankyrin facilitates intracellular trafficking of α1-Na+-K+-ATPase in polarized cells". Am. J. Physiol., Cell Physiol. 295 (5): C1202–14. doi: 10.1152/ajpcell.00273.2008. PMC 2584975. PMID 18768923.
- Sohet F, Colin Y, Genetet S, et al. (2008). "Phosphorylation and ankyrin-G binding of the C-terminal domain regulate targeting and function of the ammonium transporter RhBG". J. Biol. Chem. 283 (39): 26557–67. doi: 10.1074/jbc.M803120200. PMC 3258915. PMID 18635543.
- Ignatiuk A, Quickfall JP, Hawrysh AD, et al. (2006). "The smaller isoforms of ankyrin 3 bind to the p85 subunit of phosphatidylinositol 3'-kinase and enhance platelet-derived growth factor receptor down-regulation". J. Biol. Chem. 281 (9): 5956–64. doi: 10.1074/jbc.M510032200. PMID 16377635.
- Colland F, Jacq X, Trouplin V, et al. (2004). "Functional Proteomics Mapping of a Human Signaling Pathway". Genome Res. 14 (7): 1324–32. doi: 10.1101/gr.2334104. PMC 442148. PMID 15231748.
- Glinsky GV, Berezovska O, Glinskii AB (2005). "Microarray analysis identifies a death-from-cancer signature predicting therapy failure in patients with multiple types of cancer". J. Clin. Invest. 115 (6): 1503–21. doi: 10.1172/JCI23412. PMC 1136989. PMID 15931389.
- McEwen DP, Meadows LS, Chen C, et al. (2004). "Sodium channel beta1 subunit-mediated modulation of Nav1.2 currents and cell surface density is dependent on interactions with contactin and ankyrin". J. Biol. Chem. 279 (16): 16044–9. doi: 10.1074/jbc.M400856200. PMID 14761957.
- Ota T, Suzuki Y, Nishikawa T, et al. (2004). "Complete sequencing and characterization of 21,243 full-length human cDNAs". Nat. Genet. 36 (1): 40–5. doi: 10.1038/ng1285. PMID 14702039.
- Wang J, Robinson JF, O'Neil CH, et al. (2006). "Ankyrin G overexpression in Hutchinson–Gilford progeria syndrome fibroblasts identified through biological filtering of expression profiles". J. Hum. Genet. 51 (11): 934–42. doi: 10.1007/s10038-006-0042-0. PMID 17033732.
- Kizhatil K, Davis JQ, Davis L, et al. (2007). "Ankyrin-G is a molecular partner of E-cadherin in epithelial cells and early embryos". J. Biol. Chem. 282 (36): 26552–61. doi: 10.1074/jbc.M703158200. PMID 17620337.
- Kizhatil K, Bennett V (2004). "Lateral membrane biogenesis in human bronchial epithelial cells requires 190-kDa ankyrin-G". J. Biol. Chem. 279 (16): 16706–14. doi: 10.1074/jbc.M314296200. PMID 14757759.
External links
- ANK3+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Human ANK3 genome location and ANK3 gene details page in the UCSC Genome Browser.
This article incorporates text from the United States National Library of Medicine, which is in the public domain.