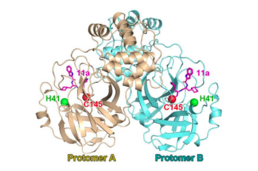
SARS-CoV-2 main proteinase
dimer with the
catalytic dyad (
H41;
C145) in complex with a
covalent
peptidomimetic protease inhibitor ("11a", magenta). From
PDB:
6LZE.
[1] | |||||||||
| Identifiers | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| EC no. | 3.4.22.69 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
| Peptidase C30, Coronavirus endopeptidase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Peptidase_C30 | ||||||||
| Pfam | PF05409 | ||||||||
| InterPro | IPR008740 | ||||||||
| PROSITE | PS51442 | ||||||||
| MEROPS | C30 | ||||||||
| SCOP2 | d1q2wb1 / SCOPe / SUPFAM | ||||||||
| |||||||||
The 3C-like protease (3CLpro) or main protease (Mpro), formally known as C30 endopeptidase or 3-chymotrypsin-like protease, [2] is the main protease found in coronaviruses. It cleaves the coronavirus polyprotein at eleven conserved sites. It is a cysteine protease and a member of the PA clan of proteases. It has a cysteine-histidine catalytic dyad at its active site and cleaves a Gln–( Ser/ Ala/ Gly) peptide bond.
The Enzyme Commission refers to this family as SARS coronavirus main proteinase (Mpro; EC 3.4.22.69). The 3CL protease corresponds to coronavirus nonstructural protein 5 (nsp5). The "3C" in the common name refers to the 3C protease (3Cpro) which is a homologous protease found in picornaviruses.
Function
The 3C-like protease is able to catalytically cleave a peptide bond between a glutamine at position P1 and a small amino acid ( serine, alanine, or glycine) at position P1'. The SARS coronavirus 3CLpro can for instance self-cleave the following peptides: [3] [4] [5]
TSAVL Q- SGFRK-NH2 and SGVTF Q- GKFKK are the two peptides corresponding to the two self-cleavage sites of the SARS 3C-like proteinase
The protease is important in the processing of the coronavirus replicase polyprotein ( P0C6U8). It is the main protease in coronaviruses and corresponds to nonstructural protein 5 (nsp5). [6] It cleaves the coronavirus polyprotein at 11 conserved sites. The 3CL protease has a cysteine-histidine catalytic dyad at its active site. [4] The sulfur of the cysteine acts as a nucleophile and the imidazole ring of the histidine as a general base. [7]
| Position | Substrate preference |
|---|---|
| P5 | No strong preference |
| P4 | Small hydrophobic residues |
| P3 | Positively charged residue |
| P2 | High hydrophobicity and absence of beta-branch |
| P1 | Glutamine |
| P1' | Small residues |
| P2' | Small residues |
| P3' | No strong preference |
Nomenclature
Alternative names provided by the EC include 3CLpro, 3C-like protease, coronavirus 3C-like protease, Mpro, SARS 3C-like protease, SARS coronavirus 3CL protease, SARS coronavirus main peptidase, SARS coronavirus main protease, SARS-CoV 3CLpro enzyme, SARS-CoV main protease, SARS-CoV Mpro and severe acute respiratory syndrome coronavirus main protease.
As a treatment target

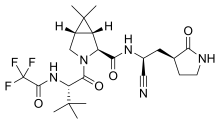
The protease 3CLpro is used as a drug target for coronavirus infections due to its essential role in processing the polyproteins that are translated from the viral RNA. [11] [12] The X-ray structures of the unliganded SARS-CoV-2 protease 3CLpro and its complex with an α-ketoamide inhibitor provides a basis for design of α-ketoamide inhibitors [13] for a treatment of SARS-CoV-2 infection. [14] [15] [16] [17] [18]
A number of protease inhibitors are being developed targeting 3CLpro and homologous 3Cpro, including CLpro-1, GC376, rupintrivir, lufotrelvir, PF-07321332, and AG7404. [19] [20] [21] [22] [1] The intravenous administered prodrug PF-07304814 (lufotrelvir) entered clinical trials in September 2020. [23]
After clinical trials, in December 2021, the oral medication nirmatrelvir (formerly PF-07321332) became commercially available under emergency use authorizations (EUA), as part of the nirmatrelvir/ritonavir combination therapy (brand name Paxlovid). [24] [25] In May 2023, the medication got full FDA approval for high-risk adults, while children 12–18 were still covered under the EUA. [26]
The 3C-like protease inhibitor ensitrelvir received authorization to treat COVID-19 in Japan in 2022. [27] [28]
In 2022, an ultralarge virtual screening campaign of 235 million molecules was able to identify a novel broad-spectrum inhibitor targeting the main protease of several coronaviruses. It is unusually not a peptidomimetic. [29]

Other 3C(-like) proteases
3C-like proteases (3C(L)pro) are widely found in (+)ssRNA viruses. All of them are cysteine proteases with a chymotrypsin-like fold (PA clan), using a catalytic dyad or triad. They share some general similarities on substrate specificity and inhibitor effectiveness. They are divided into subfamilies by sequence similarity, corresponding to the family of viruses they are found in: [30]
- This entry is the coronavirus 3CLpro.
- Picornaviridae have a picornavirus 3Cpro ( EC 3.4.22.28; InterPro: IPR000199; MEROPS C03). This is the earliest-studied family. Examples include the ones found in poliovirus and in rhinovirus (both are members of genus Enterovirus).
- Caliciviridae have a 3CLpro ( InterPro: IPR001665; MEROPS C37). Examples include the one found in Norwalk virus.
Additional members are known from Potyviridae and non-Coronaviridae Nidovirales. [31]
See also
- 3CLpro-1
- Carmofur
- COVID Moonshot
- Ebselen
- EDP-235
- GC376
- GRL-0617
- Nirmatrelvir
- Olgotrelvir
- RAY1216
- Rupintrivir
- SIM0417
- Theaflavin digallate
- Tollovir
- Y180
- Tetrahydrocannabutol
References
- ^ a b c Dai W, Zhang B, Jiang XM, Su H, Li J, Zhao Y, et al. (June 2020). "Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease". Science. 368 (6497): 1331–1335. Bibcode: 2020Sci...368.1331D. doi: 10.1126/science.abb4489. PMC 7179937. PMID 32321856.
- ^ Ahmad B, Batool M, Ain QU, Kim MS, Choi S (August 2021). "Exploring the Binding Mechanism of PF-07321332 SARS-CoV-2 Protease Inhibitor through Molecular Dynamics and Binding Free Energy Simulations". International Journal of Molecular Sciences. 22 (17): 9124. doi: 10.3390/ijms22179124. PMC 8430524. PMID 34502033.
- ^ Goetz DH, Choe Y, Hansell E, Chen YT, McDowell M, Jonsson CB, Roush WR, McKerrow J, Craik CS (July 2007). "Substrate specificity profiling and identification of a new class of inhibitor for the major protease of the SARS coronavirus". Biochemistry. 46 (30): 8744–52. doi: 10.1021/bi0621415. PMID 17605471.
- ^ a b Fan K, Wei P, Feng Q, Chen S, Huang C, Ma L, Lai B, Pei J, Liu Y, Chen J, Lai L (January 2004). "Biosynthesis, purification, and substrate specificity of severe acute respiratory syndrome coronavirus 3C-like proteinase". The Journal of Biological Chemistry. 279 (3): 1637–42. doi: 10.1074/jbc.m310875200. PMC 7980035. PMID 14561748.
- ^ Akaji K, Konno H, Onozuka M, Makino A, Saito H, Nosaka K (November 2008). "Evaluation of peptide-aldehyde inhibitors using R188I mutant of SARS 3CL protease as a proteolysis-resistant mutant". Bioorganic & Medicinal Chemistry. 16 (21): 9400–8. doi: 10.1016/j.bmc.2008.09.057. PMC 7126698. PMID 18845442.
-
^ Fehr AR, Perlman S (2015). "Coronaviruses: an overview of their replication and pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.). Coronaviruses. Methods in Molecular Biology. Vol. 1282. Springer. pp. 1–23.
doi:
10.1007/978-1-4939-2438-7_1.
ISBN
978-1-4939-2438-7.
PMC
4369385.
PMID
25720466.
See section: Virion Structure.
- ^ Ryu YB, Park SJ, Kim YM, Lee JY, Seo WD, Chang JS, et al. (March 2010). "SARS-CoV 3CLpro inhibitory effects of quinone-methide triterpenes from Tripterygium regelii". Bioorganic & Medicinal Chemistry Letters. 20 (6): 1873–6. doi: 10.1016/j.bmcl.2010.01.152. ISSN 0960-894X. PMC 7127101. PMID 20167482.
- ^ Chuck CP, Chow HF, Wan DC, Wong KB (2011). "Profiling of substrate specificities of 3C-like proteases from group 1, 2a, 2b, and 3 coronaviruses". PLOS ONE. 6 (11): e27228. Bibcode: 2011PLoSO...627228C. doi: 10.1371/journal.pone.0027228. PMC 3206940. PMID 22073294.
- ^ Vandyck K, Deval J (August 2021). "Considerations for the discovery and development of 3-chymotrypsin-like cysteine protease inhibitors targeting SARS-CoV-2 infection". Curr Opin Virol. 49: 36–40. doi: 10.1016/j.coviro.2021.04.006. PMC 8075814. PMID 34029993.
- ^ "Pfizer begins dosing in Phase II/III trial of antiviral drug for Covid-19". Clinical Trials Arena. 2 September 2021.
- ^ Hilgenfeld R (July 2014). "From SARS to MERS: crystallographic studies on coronaviral proteases enable antiviral drug design". FEBS Journal. 281 (18): 4085–4096. doi: 10.1111/febs.12936. PMC 7163996. PMID 25039866.
- ^ Ullrich S, Nitsche C (July 2020). "The SARS-CoV-2 main protease as a drug target". Bioorganic & Medicinal Chemistry Letters. 30 (17): 127377. doi: 10.1016/j.bmcl.2020.127377. PMC 7331567. PMID 32738988. S2CID 220304661.
- ^ Ocain TD, Rich DH (February 1992). "alpha-Keto amide inhibitors of aminopeptidases". Journal of Medicinal Chemistry. 35 (3): 451–6. doi: 10.1021/jm00081a005. PMID 1738140.
- ^ Anand K, Ziebuhr J, Wadhwani P, Mesters JR, Hilgenfeld R (June 2003). "Coronavirus main proteinase (3CLpro) structure: basis for design of anti-SARS drugs". Science. 300 (5626): 1763–7. Bibcode: 2003Sci...300.1763A. doi: 10.1126/science.1085658. PMID 12746549.
- ^ Pacifico S, Ferretti V, Albanese V, Fantinati A, Gallerani E, Nicoli F, et al. (July 2019). "Synthesis and Biological Activity of Peptide α-Ketoamide Derivatives as Proteasome Inhibitors". ACS Medicinal Chemistry Letters. 10 (7): 1086–1092. doi: 10.1021/acsmedchemlett.9b00233. PMC 6627721. PMID 31312413.
- ^ Kusov Y, Tan J, Alvarez E, Enjuanes L, Hilgenfeld R (October 2015). "A G-quadruplex-binding macrodomain within the "SARS-unique domain" is essential for the activity of the SARS-coronavirus replication-transcription complex". Virology. 484: 313–22. doi: 10.1016/j.virol.2015.06.016. PMC 4567502. PMID 26149721.
- ^ Zhang L, Lin D, Kusov Y, Nian Y, Ma Q, Wang J, et al. (February 2020). "α-Ketoamides as Broad-Spectrum Inhibitors of Coronavirus and Enterovirus Replication: Structure-Based Design, Synthesis, and Activity Assessment". Journal of Medicinal Chemistry. 63 (9): 4562–4578. doi: 10.1021/acs.jmedchem.9b01828. PMC 7098070. PMID 32045235.
- ^ Zhang L, Lin D, Sun X, Curth U, Drosten C, Sauerhering L, et al. (March 2020). "Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors". Science. 368 (6489): 409–412. Bibcode: 2020Sci...368..409Z. doi: 10.1126/science.abb3405. PMC 7164518. PMID 32198291.
- ^ Tian D, Liu Y, Liang C, Xin L, Xie X, Zhang D, Wan M, Li H, Fu X, Liu H, Cao W (May 2021). "An update review of emerging small-molecule therapeutic options for COVID-19". Biomedicine & Pharmacotherapy. 137: 111313. doi: 10.1016/j.biopha.2021.111313. PMC 7857046. PMID 33556871.
- ^ Morse JS, Lalonde T, Xu S, Liu WR (March 2020). "Learning from the Past: Possible Urgent Prevention and Treatment Options for Severe Acute Respiratory Infections Caused by 2019-nCoV". ChemBioChem. 21 (5): 730–738. doi: 10.1002/cbic.202000047. PMC 7162020. PMID 32022370.
- ^ Liu C, Zhou Q, Li Y, Garner LV, Watkins SP, Carter LJ, et al. (March 2020). "Research and Development on Therapeutic Agents and Vaccines for COVID-19 and Related Human Coronavirus Diseases". ACS Central Science. 6 (3): 315–331. doi: 10.1021/acscentsci.0c00272. PMC 7094090. PMID 32226821.
- ^ Ramajayam R, Tan KP, Liang PH (October 2011). "Recent development of 3C and 3CL protease inhibitors for anti-coronavirus and anti-picornavirus drug discovery". Biochemical Society Transactions. 39 (5): 1371–5. doi: 10.1042/BST0391371. PMID 21936817.
- ^ "First-In-Human Study To Evaluate Safety, Tolerability, And Pharmacokinetics Following Single Ascending And Multiple Ascending Doses of PF-07304814 In Hospitalized Participants With COVID-19". Clinical Trials. 24 June 2021. Retrieved 3 July 2021.
- ^ Fact sheet for healthcare providers: Emergency Use Authorization for Paxlovid (PDF) (Technical report). Pfizer. 22 December 2021. LAB-1492-0.8. Archived from the original on 23 December 2021.
- ^ "Pfizer Receives U.S. FDA Emergency Use Authorization for Novel COVID-19 Oral Antiviral Treatment" (Press release). Pfizer. 22 December 2021. Archived from the original on 22 December 2021. Retrieved 22 December 2021 – via Business Wire.
-
^
"FDA Approves First Oral Antiviral for Treatment of COVID-19 in Adults". U.S.
Food and Drug Administration (FDA) (Press release). 26 May 2023. Retrieved 26 May 2023.
 This article incorporates text from this source, which is in the
public domain.
This article incorporates text from this source, which is in the
public domain.
- ^ "Xocova (Ensitrelvir Fumaric Acid) Tablets 125mg Approved in Japan for the Treatment of SARS-CoV-2 Infection, under the Emergency Regulatory Approval System". Shionogi (Press release). 22 November 2022. Retrieved 28 November 2022.
- ^ Lenharo, Mariana (18 October 2023). "New Pill Helps COVID Smell and Taste Loss Fade Quickly". Scientific American. Retrieved 28 October 2023.
- ^ Luttens A, Gullberg H, Abdurakhmanov E, Vo DD, Akaberi D, Talibov VO, et al. (February 2022). "Ultralarge Virtual Screening Identifies SARS-CoV-2 Main Protease Inhibitors with Broad-Spectrum Activity against Coronaviruses". J Am Chem Soc. 144 (7): 2905–2920. doi: 10.1021/jacs.1c08402. ISSN 0002-7863. PMC 8848513. PMID 35142215.
- ^ Kim Y, Lovell S, Tiew KC, Mandadapu SR, Alliston KR, Battaile KP, et al. (November 2012). "Broad-spectrum antivirals against 3C or 3C-like proteases of picornaviruses, noroviruses, and coronaviruses". Journal of Virology. 86 (21): 11754–62. doi: 10.1128/JVI.01348-12. PMC 3486288. PMID 22915796.
- ^ Ziebuhr J, Bayer S, Cowley JA, Gorbalenya AE (January 2003). "The 3C-like proteinase of an invertebrate nidovirus links coronavirus and potyvirus homologs". Journal of Virology. 77 (2): 1415–26. doi: 10.1128/jvi.77.2.1415-1426.2003. PMC 140795. PMID 12502857.
Further reading
- Chuck CP, Chow HF, Wan DC, Wong KB (2 November 2011). "Profiling of substrate specificities of 3C-like proteases from group 1, 2a, 2b, and 3 coronaviruses". PLOS ONE. 6 (11): e27228. Bibcode: 2011PLoSO...627228C. doi: 10.1371/journal.pone.0027228. PMC 3206940. PMID 22073294.
External links
- SARS+coronavirus+main+proteinase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Peptidase C30/C16 in coronavirus, InterPro: IPR013016. The MEROPS C16 one is the "papain-like" PL-PRO.
