
| Part of a series on |
| Genetics |
|---|
 |
Ribonucleic acid (RNA) is a polymeric molecule that is essential for most biological functions, either by performing the function itself ( non-coding RNA) or by forming a template for the production of proteins ( messenger RNA). RNA and deoxyribonucleic acid (DNA) are nucleic acids. The nucleic acids constitute one of the four major macromolecules essential for all known forms of life. RNA is assembled as a chain of nucleotides. Cellular organisms use messenger RNA (mRNA) to convey genetic information (using the nitrogenous bases of guanine, uracil, adenine, and cytosine, denoted by the letters G, U, A, and C) that directs synthesis of specific proteins. Many viruses encode their genetic information using an RNA genome.
Some RNA molecules play an active role within cells by catalyzing biological reactions, controlling gene expression, or sensing and communicating responses to cellular signals. One of these active processes is protein synthesis, a universal function in which RNA molecules direct the synthesis of proteins on ribosomes. This process uses transfer RNA (tRNA) molecules to deliver amino acids to the ribosome, where ribosomal RNA (rRNA) then links amino acids together to form coded proteins.
It has become widely accepted in science [1] that early in the history of life on Earth, prior to the evolution of DNA and possibly of protein-based enzymes as well, an " RNA world" existed in which RNA served as both living organisms' storage method for genetic information—a role fulfilled today by DNA, except in the case of RNA viruses—and potentially performed catalytic functions in cells—a function performed today by protein enzymes, with the notable and important exception of the ribosome, which is a ribozyme.
Comparison with DNA
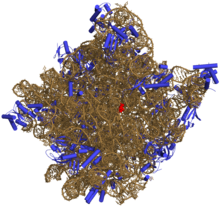
Like DNA, most biologically active RNAs, including mRNA, tRNA, rRNA, snRNAs, and other non-coding RNAs, contain self-complementary sequences that allow parts of the RNA to fold [6] and pair with itself to form double helices. Analysis of these RNAs has revealed that they are highly structured. Unlike DNA, their structures do not consist of long double helices, but rather collections of short helices packed together into structures akin to proteins.
In this fashion, RNAs can achieve chemical catalysis (like enzymes). [7] For instance, determination of the structure of the ribosome—an RNA-protein complex that catalyzes peptide bond formation—revealed that its active site is composed entirely of RNA. [8]
Structure
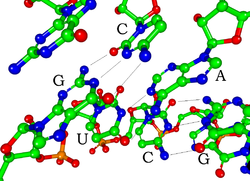
Each nucleotide in RNA contains a ribose sugar, with carbons numbered 1' through 5'. A base is attached to the 1' position, in general, adenine (A), cytosine (C), guanine (G), or uracil (U). Adenine and guanine are purines, and cytosine and uracil are pyrimidines. A phosphate group is attached to the 3' position of one ribose and the 5' position of the next. The phosphate groups have a negative charge each, making RNA a charged molecule (polyanion). The bases form hydrogen bonds between cytosine and guanine, between adenine and uracil and between guanine and uracil. [9] However, other interactions are possible, such as a group of adenine bases binding to each other in a bulge, [10] or the GNRA tetraloop that has a guanine–adenine base-pair. [9]
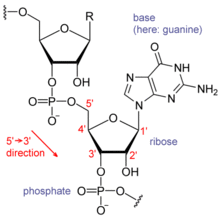
An important structural component of RNA that distinguishes it from DNA is the presence of a hydroxyl group at the 2' position of the ribose sugar. The presence of this functional group causes the helix to mostly take the A-form geometry, [11] although in single strand dinucleotide contexts, RNA can rarely also adopt the B-form most commonly observed in DNA. [12] The A-form geometry results in a very deep and narrow major groove and a shallow and wide minor groove. [13] A second consequence of the presence of the 2'-hydroxyl group is that in conformationally flexible regions of an RNA molecule (that is, not involved in formation of a double helix), it can chemically attack the adjacent phosphodiester bond to cleave the backbone. [14]
RNA is transcribed with only four bases (adenine, cytosine, guanine and uracil), [15] but these bases and attached sugars can be modified in numerous ways as the RNAs mature. Pseudouridine (Ψ), in which the linkage between uracil and ribose is changed from a C–N bond to a C–C bond, and ribothymidine (T) are found in various places (the most notable ones being in the TΨC loop of tRNA). [16] Another notable modified base is hypoxanthine, a deaminated adenine base whose nucleoside is called inosine (I). Inosine plays a key role in the wobble hypothesis of the genetic code. [17]
There are more than 100 other naturally occurring modified nucleosides. [18] The greatest structural diversity of modifications can be found in tRNA, [19] while pseudouridine and nucleosides with 2'-O-methylribose often present in rRNA are the most common. [20] The specific roles of many of these modifications in RNA are not fully understood. However, it is notable that, in ribosomal RNA, many of the post-transcriptional modifications occur in highly functional regions, such as the peptidyl transferase center [21] and the subunit interface, implying that they are important for normal function. [22]
The functional form of single-stranded RNA molecules, just like proteins, frequently requires a specific tertiary structure. The scaffold for this structure is provided by secondary structural elements that are hydrogen bonds within the molecule. This leads to several recognizable "domains" of secondary structure like hairpin loops, bulges, and internal loops. [23] In order to create, i.e., design, a RNA for any given secondary structure, two or three bases would not be enough, but four bases are enough. [24] This is likely why nature has "chosen" a four base alphabet: fewer than four would not allow the creation of all structures, while more than four bases are not necessary to do so. Since RNA is charged, metal ions such as Mg2+ are needed to stabilise many secondary and tertiary structures. [25]
The naturally occurring enantiomer of RNA is D-RNA composed of D-ribonucleotides. All chirality centers are located in the D-ribose. By the use of L-ribose or rather L-ribonucleotides, L-RNA can be synthesized. L-RNA is much more stable against degradation by RNase. [26]
Like other structured biopolymers such as proteins, one can define topology of a folded RNA molecule. This is often done based on arrangement of intra-chain contacts within a folded RNA, termed as circuit topology.
Synthesis
Synthesis of RNA is usually catalyzed by an enzyme— RNA polymerase—using DNA as a template, a process known as transcription. Initiation of transcription begins with the binding of the enzyme to a promoter sequence in the DNA (usually found "upstream" of a gene). The DNA double helix is unwound by the helicase activity of the enzyme. The enzyme then progresses along the template strand in the 3’ to 5’ direction, synthesizing a complementary RNA molecule with elongation occurring in the 5’ to 3’ direction. The DNA sequence also dictates where termination of RNA synthesis will occur. [27]
Primary transcript RNAs are often modified by enzymes after transcription. For example, a poly(A) tail and a 5' cap are added to eukaryotic pre-mRNA and introns are removed by the spliceosome.
There are also a number of RNA-dependent RNA polymerases that use RNA as their template for synthesis of a new strand of RNA. For instance, a number of RNA viruses (such as poliovirus) use this type of enzyme to replicate their genetic material. [28] Also, RNA-dependent RNA polymerase is part of the RNA interference pathway in many organisms. [29]
Types of RNA
Overview

Messenger RNA (mRNA) is the RNA that carries information from DNA to the ribosome, the sites of protein synthesis ( translation) in the cell. The mRNA is a copy of DNA. The coding sequence of the mRNA determines the amino acid sequence in the protein that is produced. [30] However, many RNAs do not code for protein (about 97% of the transcriptional output is non-protein-coding in eukaryotes [31] [32] [33] [34]).
These so-called non-coding RNAs ("ncRNA") can be encoded by their own genes (RNA genes), but can also derive from mRNA introns. [35] The most prominent examples of non-coding RNAs are transfer RNA (tRNA) and ribosomal RNA (rRNA), both of which are involved in the process of translation. [5] There are also non-coding RNAs involved in gene regulation, RNA processing and other roles. Certain RNAs are able to catalyse chemical reactions such as cutting and ligating other RNA molecules, [36] and the catalysis of peptide bond formation in the ribosome; [8] these are known as ribozymes.
In length
According to the length of RNA chain, RNA includes small RNA and long RNA. [37] Usually, small RNAs are shorter than 200 nt in length, and long RNAs are greater than 200 nt long. [38] Long RNAs, also called large RNAs, mainly include long non-coding RNA (lncRNA) and mRNA. Small RNAs mainly include 5.8S ribosomal RNA (rRNA), 5S rRNA, transfer RNA (tRNA), microRNA (miRNA), small interfering RNA (siRNA), small nucleolar RNA (snoRNAs), Piwi-interacting RNA (piRNA), tRNA-derived small RNA (tsRNA) [39] and small rDNA-derived RNA (srRNA). [40] There are certain exceptions as in the case of the 5S rRNA of the members of the genus Halococcus ( Archaea), which have an insertion, thus increasing its size. [41] [42] [43]
In translation
Messenger RNA (mRNA) carries information about a protein sequence to the ribosomes, the protein synthesis factories in the cell. It is coded so that every three nucleotides (a codon) corresponds to one amino acid. In eukaryotic cells, once precursor mRNA (pre-mRNA) has been transcribed from DNA, it is processed to mature mRNA. This removes its introns—non-coding sections of the pre-mRNA. The mRNA is then exported from the nucleus to the cytoplasm, where it is bound to ribosomes and translated into its corresponding protein form with the help of tRNA. In prokaryotic cells, which do not have nucleus and cytoplasm compartments, mRNA can bind to ribosomes while it is being transcribed from DNA. After a certain amount of time, the message degrades into its component nucleotides with the assistance of ribonucleases. [30]
Transfer RNA (tRNA) is a small RNA chain of about 80 nucleotides that transfers a specific amino acid to a growing polypeptide chain at the ribosomal site of protein synthesis during translation. It has sites for amino acid attachment and an anticodon region for codon recognition that binds to a specific sequence on the messenger RNA chain through hydrogen bonding. [35]

Ribosomal RNA (rRNA) is the catalytic component of the ribosomes. The rRNA is the component of the ribosome that hosts translation. Eukaryotic ribosomes contain four different rRNA molecules: 18S, 5.8S, 28S and 5S rRNA. Three of the rRNA molecules are synthesized in the nucleolus, and one is synthesized elsewhere. In the cytoplasm, ribosomal RNA and protein combine to form a nucleoprotein called a ribosome. The ribosome binds mRNA and carries out protein synthesis. Several ribosomes may be attached to a single mRNA at any time. [30] Nearly all the RNA found in a typical eukaryotic cell is rRNA.
Transfer-messenger RNA (tmRNA) is found in many bacteria and plastids. It tags proteins encoded by mRNAs that lack stop codons for degradation and prevents the ribosome from stalling. [44]
Regulatory RNA
The earliest known regulators of gene expression were proteins known as repressors and activators – regulators with specific short binding sites within enhancer regions near the genes to be regulated. [45] Later studies have shown that RNAs also regulate genes. There are several kinds of RNA-dependent processes in eukaryotes regulating the expression of genes at various points, such as RNAi repressing genes post-transcriptionally, long non-coding RNAs shutting down blocks of chromatin epigenetically, and enhancer RNAs inducing increased gene expression. [46] Bacteria and archaea have also been shown to use regulatory RNA systems such as bacterial small RNAs and CRISPR. [47] Fire and Mello were awarded the 2006 Nobel Prize in Physiology or Medicine for discovering microRNAs (miRNAs), specific short RNA molecules that can base-pair with mRNAs. [48]
RNA interference by miRNAs
Post-transcriptional expression levels of many genes can be controlled by RNA interference, in which miRNAs, specific short RNA molecules, pair with mRNA regions and target them for degradation. [49] This antisense-based process involves steps that first process the RNA so that it can base-pair with a region of its target mRNAs. Once the base pairing occurs, other proteins direct the mRNA to be destroyed by nucleases. [46]
Long non-coding RNAs
Next to be linked to regulation were Xist and other long noncoding RNAs associated with X chromosome inactivation. Their roles, at first mysterious, were shown by Jeannie T. Lee and others to be the silencing of blocks of chromatin via recruitment of Polycomb complex so that messenger RNA could not be transcribed from them. [50] Additional lncRNAs, currently defined as RNAs of more than 200 base pairs that do not appear to have coding potential, [51] have been found associated with regulation of stem cell pluripotency and cell division. [51]
Enhancer RNAs
The third major group of regulatory RNAs is called enhancer RNAs. [51] It is not clear at present whether they are a unique category of RNAs of various lengths or constitute a distinct subset of lncRNAs. In any case, they are transcribed from enhancers, which are known regulatory sites in the DNA near genes they regulate. [51] [52] They up-regulate the transcription of the gene(s) under control of the enhancer from which they are transcribed. [51] [53]
Regulatory RNA in prokaryotes
At first, regulatory RNA was thought to be a eukaryotic phenomenon, a part of the explanation for why so much more transcription in higher organisms was seen than had been predicted. But as soon as researchers began to look for possible RNA regulators in bacteria, they turned up there as well, termed as small RNA (sRNA). [54] [47] Currently, the ubiquitous nature of systems of RNA regulation of genes has been discussed as support for the RNA World theory. [46] [55] There are indications that the enterobacterial sRNAs are involved in various cellular processes and seem to have significant role in stress responses such as membrane stress, starvation stress, phosphosugar stress and DNA damage. Also, it has been suggested that sRNAs have been evolved to have important role in stress responses because of their kinetic properties that allow for rapid response and stabilisation of the physiological state. [2] Bacterial small RNAs generally act via antisense pairing with mRNA to down-regulate its translation, either by affecting stability or affecting cis-binding ability. [46] Riboswitches have also been discovered. They are cis-acting regulatory RNA sequences acting allosterically. They change shape when they bind metabolites so that they gain or lose the ability to bind chromatin to regulate expression of genes. [56] [57]
Archaea also have systems of regulatory RNA. [58] The CRISPR system, recently being used to edit DNA in situ, acts via regulatory RNAs in archaea and bacteria to provide protection against virus invaders. [46] [59]
In RNA processing

Many RNAs are involved in modifying other RNAs. Introns are spliced out of pre-mRNA by spliceosomes, which contain several small nuclear RNAs (snRNA), [5] or the introns can be ribozymes that are spliced by themselves. [60] RNA can also be altered by having its nucleotides modified to nucleotides other than A, C, G and U. In eukaryotes, modifications of RNA nucleotides are in general directed by small nucleolar RNAs (snoRNA; 60–300 nt), [35] found in the nucleolus and cajal bodies. snoRNAs associate with enzymes and guide them to a spot on an RNA by basepairing to that RNA. These enzymes then perform the nucleotide modification. rRNAs and tRNAs are extensively modified, but snRNAs and mRNAs can also be the target of base modification. [61] [62] RNA can also be methylated. [63] [64]
RNA genomes
Like DNA, RNA can carry genetic information. RNA viruses have genomes composed of RNA that encodes a number of proteins. The viral genome is replicated by some of those proteins, while other proteins protect the genome as the virus particle moves to a new host cell. Viroids are another group of pathogens, but they consist only of RNA, do not encode any protein and are replicated by a host plant cell's polymerase. [65]
In reverse transcription
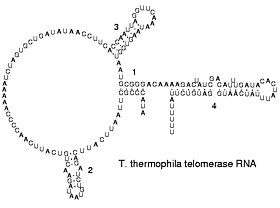
Reverse transcribing viruses replicate their genomes by reverse transcribing DNA copies from their RNA; these DNA copies are then transcribed to new RNA. Retrotransposons also spread by copying DNA and RNA from one another, [66] and telomerase contains an RNA that is used as template for building the ends of eukaryotic chromosomes. [67]
Double-stranded RNA

Double-stranded RNA (dsRNA) is RNA with two complementary strands, similar to the DNA found in all cells, but with the replacement of thymine by uracil and the adding of one oxygen atom. dsRNA forms the genetic material of some viruses ( double-stranded RNA viruses). Double-stranded RNA, such as viral RNA or siRNA, can trigger RNA interference in eukaryotes, as well as interferon response in vertebrates. [68] [69] [70] [71] In eukaryotes, double-stranded RNA (dsRNA) plays a role in the activation of the innate immune system against viral infections. [72]
Circular RNA
In the late 1970s, it was shown that there is a single stranded covalently closed, i.e. circular form of RNA expressed throughout the animal and plant kingdom (see circRNA). [73] circRNAs are thought to arise via a "back-splice" reaction where the spliceosome joins a upstream 3' acceptor to a downstream 5' donor splice site. So far the function of circRNAs is largely unknown, although for few examples a microRNA sponging activity has been demonstrated.
Key discoveries in RNA biology

Research on RNA has led to many important biological discoveries and numerous Nobel Prizes. Nucleic acids were discovered in 1868 by Friedrich Miescher, who called the material 'nuclein' since it was found in the nucleus. [74] It was later discovered that prokaryotic cells, which do not have a nucleus, also contain nucleic acids. The role of RNA in protein synthesis was suspected already in 1939. [75] Severo Ochoa won the 1959 Nobel Prize in Medicine (shared with Arthur Kornberg) after he discovered an enzyme that can synthesize RNA in the laboratory. [76] However, the enzyme discovered by Ochoa ( polynucleotide phosphorylase) was later shown to be responsible for RNA degradation, not RNA synthesis. In 1956 Alex Rich and David Davies hybridized two separate strands of RNA to form the first crystal of RNA whose structure could be determined by X-ray crystallography. [77]
The sequence of the 77 nucleotides of a yeast tRNA was found by Robert W. Holley in 1965, [78] winning Holley the 1968 Nobel Prize in Medicine (shared with Har Gobind Khorana and Marshall Nirenberg).
In the early 1970s, retroviruses and reverse transcriptase were discovered, showing for the first time that enzymes could copy RNA into DNA (the opposite of the usual route for transmission of genetic information). For this work, David Baltimore, Renato Dulbecco and Howard Temin were awarded a Nobel Prize in 1975. In 1976, Walter Fiers and his team determined the first complete nucleotide sequence of an RNA virus genome, that of bacteriophage MS2. [79]
In 1977, introns and RNA splicing were discovered in both mammalian viruses and in cellular genes, resulting in a 1993 Nobel to Philip Sharp and Richard Roberts. Catalytic RNA molecules ( ribozymes) were discovered in the early 1980s, leading to a 1989 Nobel award to Thomas Cech and Sidney Altman. In 1990, it was found in Petunia that introduced genes can silence similar genes of the plant's own, now known to be a result of RNA interference. [80] [81]
At about the same time, 22 nt long RNAs, now called microRNAs, were found to have a role in the development of C. elegans. [82] Studies on RNA interference gleaned a Nobel Prize for Andrew Fire and Craig Mello in 2006, and another Nobel was awarded for studies on the transcription of RNA to Roger Kornberg in the same year. The discovery of gene regulatory RNAs has led to attempts to develop drugs made of RNA, such as siRNA, to silence genes. [83] Adding to the Nobel prizes awarded for research on RNA in 2009 it was awarded for the elucidation of the atomic structure of the ribosome to Venki Ramakrishnan, Thomas A. Steitz, and Ada Yonath.
Relevance for prebiotic chemistry and abiogenesis
In 1968, Carl Woese hypothesized that RNA might be catalytic and suggested that the earliest forms of life (self-replicating molecules) could have relied on RNA both to carry genetic information and to catalyze biochemical reactions—an RNA world. [84] [85] In May 2022, scientists reported that they discovered RNA forms spontaneously on prebiotic basalt lava glass which is presumed to have been abundantly available on the early Earth. [86] [87]
In March 2015, DNA and RNA nucleobases, including uracil, cytosine and thymine, were reportedly formed in the laboratory under outer space conditions, using starter chemicals, such as pyrimidine, an organic compound commonly found in meteorites. Pyrimidine, like polycyclic aromatic hydrocarbons (PAHs), is one of the most carbon-rich compounds found in the Universe and may have been formed in red giants or in interstellar dust and gas clouds. [88] In July 2022, astronomers reported the discovery of massive amounts of prebiotic molecules, including possible RNA precursors, in the Galactic Center of the Milky Way Galaxy. [89] [90]
Medical applications
RNA, initially deemed unsuitable for therapeutic use due to its short half-life, has been proven to possess numerous therapeutic properties through advancements in stabilization chemistry. RNA molecules have potential therapeutic applications due to their ability to fold into complex conformations and binding proteins, nucleic acids, small molecules, and form catalytic centers. [91] RNA-based vaccines are thought to be a quicker way to obtain immunological resistance than the traditional approach of vaccines that rely on a killed or altered version of the pathogen, because it can take months or even years to grow and study a pathogen in order to determine which molecular parts to extract, inactivate, and use in a vaccine. Small molecules with conventional therapeutic properties can target RNA and DNA structures, thereby treating novel diseases. However, research on small molecules targeting RNA and approved drugs for human illness therapy is scarce. Ribavirin, branaplam, and ataluren are currently available medications that stabilize double-stranded RNA structures and control splicing in a variety of disorders. [92] [93]
Protein-coding mRNAs have emerged as new therapeutic candidates, with RNA replacement being particularly beneficial for brief but torrent-like protein expression. [94] In vitro transcribed mRNAs (IVT-mRNA) have been used to deliver proteins for bone regeneration, pluripotency, and heart function in animal models. [95] [96] [97] [98] [99] SiRNAs, short RNA molecules, play a crucial role in innate defense against viruses and chromatin structure. They can be artificially introduced to silence specific genes, making them valuable for gene function studies, therapeutic target validation, and drug development. [94]
See also
References
-
^ Copley SD, Smith E, Markowitz HJ (December 2007). "The origin of the RNA world: co-evolution of genes and metabolism". Bioorganic Chemistry. 35 (6): 430–443.
doi:
10.1016/j.bioorg.2007.08.001.
PMID
17897696.
The proposal that life on Earth arose from an RNA World is widely accepted.
- ^ a b "RNA: The Versatile Molecule". University of Utah. 2015.
- ^ "Nucleotides and Nucleic Acids" (PDF). University of California, Los Angeles. Archived from the original (PDF) on 2015-09-23. Retrieved 2015-08-26.
- ^ Shukla RN (2014). Analysis of Chromosomes. Agrotech Press. ISBN 978-93-84568-17-7.[ permanent dead link]
- ^ a b c Berg JM, Tymoczko JL, Stryer L (2002). Biochemistry (5th ed.). WH Freeman and Company. pp. 118–19, 781–808. ISBN 978-0-7167-4684-3. OCLC 179705944.
- ^ Tinoco I, Bustamante C (October 1999). "How RNA folds". Journal of Molecular Biology. 293 (2): 271–81. doi: 10.1006/jmbi.1999.3001. PMID 10550208.
- ^ Higgs PG (August 2000). "RNA secondary structure: physical and computational aspects". Quarterly Reviews of Biophysics. 33 (3): 199–253. doi: 10.1017/S0033583500003620. PMID 11191843. S2CID 37230785.
- ^ a b Nissen P, Hansen J, Ban N, Moore PB, Steitz TA (August 2000). "The structural basis of ribosome activity in peptide bond synthesis". Science. 289 (5481): 920–30. Bibcode: 2000Sci...289..920N. doi: 10.1126/science.289.5481.920. PMID 10937990.
- ^ a b Lee JC, Gutell RR (December 2004). "Diversity of base-pair conformations and their occurrence in rRNA structure and RNA structural motifs". Journal of Molecular Biology. 344 (5): 1225–49. doi: 10.1016/j.jmb.2004.09.072. PMID 15561141.
- ^ Barciszewski J, Frederic B, Clark C (1999). RNA biochemistry and biotechnology. Springer. pp. 73–87. ISBN 978-0-7923-5862-6. OCLC 52403776.
- ^ Salazar M, Fedoroff OY, Miller JM, Ribeiro NS, Reid BR (April 1993). "The DNA strand in DNA.RNA hybrid duplexes is neither B-form nor A-form in solution". Biochemistry. 32 (16): 4207–15. doi: 10.1021/bi00067a007. PMID 7682844.
- ^ Sedova A, Banavali NK (February 2016). "RNA approaches the B-form in stacked single strand dinucleotide contexts". Biopolymers. 105 (2): 65–82. doi: 10.1002/bip.22750. PMID 26443416. S2CID 35949700.
- ^ Hermann T, Patel DJ (March 2000). "RNA bulges as architectural and recognition motifs". Structure. 8 (3): R47–54. doi: 10.1016/S0969-2126(00)00110-6. PMID 10745015.
- ^ Mikkola S, Stenman E, Nurmi K, Yousefi-Salakdeh E, Strömberg R, Lönnberg H (1999). "The mechanism of the metal ion promoted cleavage of RNA phosphodiester bonds involves a general acid catalysis by the metal aquo ion on the departure of the leaving group". Journal of the Chemical Society, Perkin Transactions 2 (8): 1619–26. doi: 10.1039/a903691a.
- ^ Jankowski JA, Polak JM (1996). Clinical gene analysis and manipulation: Tools, techniques and troubleshooting. Cambridge University Press. p. 14. ISBN 978-0-521-47896-0. OCLC 33838261.
- ^ Yu Q, Morrow CD (May 2001). "Identification of critical elements in the tRNA acceptor stem and T(Psi)C loop necessary for human immunodeficiency virus type 1 infectivity". Journal of Virology. 75 (10): 4902–6. doi: 10.1128/JVI.75.10.4902-4906.2001. PMC 114245. PMID 11312362.
- ^ Elliott MS, Trewyn RW (February 1984). "Inosine biosynthesis in transfer RNA by an enzymatic insertion of hypoxanthine". The Journal of Biological Chemistry. 259 (4): 2407–10. doi: 10.1016/S0021-9258(17)43367-9. PMID 6365911.
- ^ Cantara WA, Crain PF, Rozenski J, McCloskey JA, Harris KA, Zhang X, Vendeix FA, Fabris D, Agris PF (January 2011). "The RNA Modification Database, RNAMDB: 2011 update". Nucleic Acids Research. 39 (Database issue): D195-201. doi: 10.1093/nar/gkq1028. PMC 3013656. PMID 21071406.
- ^ Söll D, RajBhandary U (1995). TRNA: Structure, biosynthesis, and function. ASM Press. p. 165. ISBN 978-1-55581-073-3. OCLC 183036381.
- ^ Kiss T (July 2001). "Small nucleolar RNA-guided post-transcriptional modification of cellular RNAs". The EMBO Journal. 20 (14): 3617–22. doi: 10.1093/emboj/20.14.3617. PMC 125535. PMID 11447102.
- ^ Tirumalai MR, Rivas M, Tran Q, Fox GE (November 2021). "The Peptidyl Transferase Center: a Window to the Past". Microbiol Mol Biol Rev. 85 (4): e0010421. doi: 10.1128/MMBR.00104-21. PMC 8579967. PMID 34756086.
- ^ King TH, Liu B, McCully RR, Fournier MJ (February 2003). "Ribosome structure and activity are altered in cells lacking snoRNPs that form pseudouridines in the peptidyl transferase center". Molecular Cell. 11 (2): 425–35. doi: 10.1016/S1097-2765(03)00040-6. PMID 12620230.
- ^ Mathews DH, Disney MD, Childs JL, Schroeder SJ, Zuker M, Turner DH (May 2004). "Incorporating chemical modification constraints into a dynamic programming algorithm for prediction of RNA secondary structure". Proceedings of the National Academy of Sciences of the United States of America. 101 (19): 7287–92. Bibcode: 2004PNAS..101.7287M. doi: 10.1073/pnas.0401799101. PMC 409911. PMID 15123812.
- ^ Burghardt B, Hartmann AK (February 2007). "RNA secondary structure design". Physical Review E. 75 (2): 021920. arXiv: physics/0609135. Bibcode: 2007PhRvE..75b1920B. doi: 10.1103/PhysRevE.75.021920. PMID 17358380. S2CID 17574854.
- ^ Tan ZJ, Chen SJ (July 2008). "Salt dependence of nucleic acid hairpin stability". Biophysical Journal. 95 (2): 738–52. Bibcode: 2008BpJ....95..738T. doi: 10.1529/biophysj.108.131524. PMC 2440479. PMID 18424500.
- ^ Vater A, Klussmann S (January 2015). "Turning mirror-image oligonucleotides into drugs: the evolution of Spiegelmer(®) therapeutics". Drug Discovery Today. 20 (1): 147–55. doi: 10.1016/j.drudis.2014.09.004. PMID 25236655.
- ^ Nudler E, Gottesman ME (August 2002). "Transcription termination and anti-termination in E. coli". Genes to Cells. 7 (8): 755–68. doi: 10.1046/j.1365-2443.2002.00563.x. PMID 12167155. S2CID 23191624.
- ^ Hansen JL, Long AM, Schultz SC (August 1997). "Structure of the RNA-dependent RNA polymerase of poliovirus". Structure. 5 (8): 1109–22. doi: 10.1016/S0969-2126(97)00261-X. PMID 9309225.
- ^ Ahlquist P (May 2002). "RNA-dependent RNA polymerases, viruses, and RNA silencing". Science. 296 (5571): 1270–73. Bibcode: 2002Sci...296.1270A. doi: 10.1126/science.1069132. PMID 12016304. S2CID 42526536.
- ^ a b c Cooper GC, Hausman RE (2004). The Cell: A Molecular Approach (3rd ed.). Sinauer. pp. 261–76, 297, 339–44. ISBN 978-0-87893-214-6. OCLC 174924833.
- ^ Mattick JS, Gagen MJ (September 2001). "The evolution of controlled multitasked gene networks: the role of introns and other noncoding RNAs in the development of complex organisms". Molecular Biology and Evolution. 18 (9): 1611–30. doi: 10.1093/oxfordjournals.molbev.a003951. PMID 11504843.
- ^ Mattick JS (November 2001). "Non-coding RNAs: the architects of eukaryotic complexity". EMBO Reports. 2 (11): 986–91. doi: 10.1093/embo-reports/kve230. PMC 1084129. PMID 11713189.
- ^ Mattick JS (October 2003). "Challenging the dogma: the hidden layer of non-protein-coding RNAs in complex organisms" (PDF). BioEssays. 25 (10): 930–39. CiteSeerX 10.1.1.476.7561. doi: 10.1002/bies.10332. PMID 14505360. Archived from the original (PDF) on 2009-03-06.
- ^ Mattick JS (October 2004). "The hidden genetic program of complex organisms". Scientific American. 291 (4): 60–67. Bibcode: 2004SciAm.291d..60M. doi: 10.1038/scientificamerican1004-60. PMID 15487671.[ dead link]
- ^ a b c Wirta W (2006). Mining the transcriptome – methods and applications. Stockholm: School of Biotechnology, Royal Institute of Technology. ISBN 978-91-7178-436-0. OCLC 185406288.
- ^ Rossi JJ (July 2004). "Ribozyme diagnostics comes of age". Chemistry & Biology. 11 (7): 894–95. doi: 10.1016/j.chembiol.2004.07.002. PMID 15271347.
- ^ Storz G (May 2002). "An expanding universe of noncoding RNAs". Science. 296 (5571): 1260–63. Bibcode: 2002Sci...296.1260S. doi: 10.1126/science.1072249. PMID 12016301. S2CID 35295924.
- ^ Fatica A, Bozzoni I (January 2014). "Long non-coding RNAs: new players in cell differentiation and development". Nature Reviews Genetics. 15 (1): 7–21. doi: 10.1038/nrg3606. PMID 24296535. S2CID 12295847.[ permanent dead link]
- ^ Chen Q, Yan M, Cao Z, Li X, Zhang Y, Shi J, et al. (January 2016). "Sperm tsRNAs contribute to intergenerational inheritance of an acquired metabolic disorder" (PDF). Science. 351 (6271): 397–400. Bibcode: 2016Sci...351..397C. doi: 10.1126/science.aad7977. PMID 26721680. S2CID 21738301.
- ^ Wei H, Zhou B, Zhang F, Tu Y, Hu Y, Zhang B, Zhai Q (2013). "Profiling and identification of small rDNA-derived RNAs and their potential biological functions". PLOS ONE. 8 (2): e56842. Bibcode: 2013PLoSO...856842W. doi: 10.1371/journal.pone.0056842. PMC 3572043. PMID 23418607.
- ^ Luehrsen KR, Nicholson DE, Eubanks DC, Fox GE (1981). "An archaebacterial 5S rRNA contains a long insertion sequence". Nature. 293 (5835): 755–756. Bibcode: 1981Natur.293..755L. doi: 10.1038/293755a0. PMID 6169998. S2CID 4341755.
- ^ Stan-Lotter H, McGenity TJ, Legat A, Denner EB, Glaser K, Stetter KO, Wanner G (1999). "Very similar strains of Halococcus salifodinae are found in geographically separated permo-triassic salt deposits". Microbiology. 145 (Pt 12): 3565–3574. doi: 10.1099/00221287-145-12-3565. PMID 10627054.
- ^ Tirumalai MR, Kaelber JT, Park DR, Tran Q, Fox GE (August 2020). "Cryo-Electron Microscopy Visualization of a Large Insertion in the 5S ribosomal RNA of the Extremely Halophilic Archaeon Halococcus morrhuae". FEBS Open Bio. 10 (10): 1938–1946. doi: 10.1002/2211-5463.12962. PMC 7530397. PMID 32865340.
- ^ Gueneau de Novoa P, Williams KP (January 2004). "The tmRNA website: reductive evolution of tmRNA in plastids and other endosymbionts". Nucleic Acids Research. 32 (Database issue): D104–08. doi: 10.1093/nar/gkh102. PMC 308836. PMID 14681369.
- ^ Jacob F, Monod J (1961). "Genetic Regulatory Mechanisms in the Synthesis of Proteins". Journal of Molecular Biology. 3 (3): 318–56. doi: 10.1016/s0022-2836(61)80072-7. PMID 13718526. S2CID 19804795.
- ^ a b c d e Morris K, Mattick J (2014). "The rise of regulatory RNA". Nature Reviews Genetics. 15 (6): 423–37. doi: 10.1038/nrg3722. PMC 4314111. PMID 24776770.
- ^ a b Gottesman S (2005). "Micros for microbes: non-coding regulatory RNAs in bacteria". Trends in Genetics. 21 (7): 399–404. doi: 10.1016/j.tig.2005.05.008. PMID 15913835.
- ^ "The Nobel Prize in Physiology or Medicine 2006". Nobelprize.org. Nobel Media AB 2014. Web. 6 Aug 2018. http://www.nobelprize.org/nobel_prizes/medicine/laureates/2006
- ^ Fire, et al. (1998). "Potent and Specific Genetic Interference by double stranded RNA in Ceanorhabditis elegans". Nature. 391 (6669): 806–11. Bibcode: 1998Natur.391..806F. doi: 10.1038/35888. PMID 9486653. S2CID 4355692.
- ^ Zhao J, Sun BK, Erwin JA, Song JJ, Lee JT (2008). "Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome". Science. 322 (5902): 750–56. Bibcode: 2008Sci...322..750Z. doi: 10.1126/science.1163045. PMC 2748911. PMID 18974356.
- ^ a b c d e Rinn JL, Chang HY (2012). "Genome regulation by long noncoding RNAs". Annu. Rev. Biochem. 81: 1–25. doi: 10.1146/annurev-biochem-051410-092902. PMC 3858397. PMID 22663078.
- ^ Taft RJ, Kaplan CD, Simons C, Mattick JS (2009). "Evolution, biogenesis and function of promoter- associated RNAs". Cell Cycle. 8 (15): 2332–38. doi: 10.4161/cc.8.15.9154. PMID 19597344.
- ^ Orom UA, Derrien T, Beringer M, Gumireddy K, Gardini A, et al. (2010). "'Long noncoding RNAs with enhancer-like function in human cells". Cell. 143 (1): 46–58. doi: 10.1016/j.cell.2010.09.001. PMC 4108080. PMID 20887892.
- ^ EGH Wagner, P Romby. (2015). "Small RNAs in bacteria and archaea: who they are, what they do, and how they do it". Advances in genetics (Vol. 90, pp. 133–208).
- ^ J.W. Nelson, R.R. Breaker (2017) "The lost language of the RNA World."Sci. Signal.10, eaam8812 1–11.
- ^ Winklef WC (2005). "Riboswitches and the role of noncoding RNAs in bacterial metabolic control". Curr. Opin. Chem. Biol. 9 (6): 594–602. doi: 10.1016/j.cbpa.2005.09.016. PMID 16226486.
- ^ Tucker BJ, Breaker RR (2005). "Riboswitches as versatile gene control elements". Curr. Opin. Struct. Biol. 15 (3): 342–48. doi: 10.1016/j.sbi.2005.05.003. PMID 15919195.
- ^ Mojica FJ, Diez-Villasenor C, Soria E, Juez G (2000). "" "Biological significance of a family of regularly spaced repeats in the genomes of archaea, bacteria and mitochondria". Mol. Microbiol. 36 (1): 244–46. doi: 10.1046/j.1365-2958.2000.01838.x. PMID 10760181. S2CID 22216574.
- ^ Brouns S, Jore MM, Lundgren M, Westra E, Slijkhuis R, Snijders A, Dickman M, Makarova K, Koonin E, Der Oost JV (2008). "Small CRISPR RNAs guide antiviral defense in prokaryotes". Science. 321 (5891): 960–64. Bibcode: 2008Sci...321..960B. doi: 10.1126/science.1159689. PMC 5898235. PMID 18703739.
- ^ Steitz TA, Steitz JA (July 1993). "A general two-metal-ion mechanism for catalytic RNA". Proceedings of the National Academy of Sciences of the United States of America. 90 (14): 6498–502. Bibcode: 1993PNAS...90.6498S. doi: 10.1073/pnas.90.14.6498. PMC 46959. PMID 8341661.
- ^ Xie J, Zhang M, Zhou T, Hua X, Tang L, Wu W (January 2007). "Sno/scaRNAbase: a curated database for small nucleolar RNAs and cajal body-specific RNAs". Nucleic Acids Research. 35 (Database issue): D183–87. doi: 10.1093/nar/gkl873. PMC 1669756. PMID 17099227.
- ^ Omer AD, Ziesche S, Decatur WA, Fournier MJ, Dennis PP (May 2003). "RNA-modifying machines in archaea". Molecular Microbiology. 48 (3): 617–29. doi: 10.1046/j.1365-2958.2003.03483.x. PMID 12694609. S2CID 20326977.
- ^ Cavaillé J, Nicoloso M, Bachellerie JP (October 1996). "Targeted ribose methylation of RNA in vivo directed by tailored antisense RNA guides". Nature. 383 (6602): 732–35. Bibcode: 1996Natur.383..732C. doi: 10.1038/383732a0. PMID 8878486. S2CID 4334683.
- ^ Kiss-László Z, Henry Y, Bachellerie JP, Caizergues-Ferrer M, Kiss T (June 1996). "Site-specific ribose methylation of preribosomal RNA: a novel function for small nucleolar RNAs". Cell. 85 (7): 1077–88. doi: 10.1016/S0092-8674(00)81308-2. PMID 8674114. S2CID 10418885.
- ^ Daròs JA, Elena SF, Flores R (June 2006). "Viroids: an Ariadne's thread into the RNA labyrinth". EMBO Reports. 7 (6): 593–98. doi: 10.1038/sj.embor.7400706. PMC 1479586. PMID 16741503.
- ^ Kalendar R, Vicient CM, Peleg O, Anamthawat-Jonsson K, Bolshoy A, Schulman AH (March 2004). "Large retrotransposon derivatives: abundant, conserved but nonautonomous retroelements of barley and related genomes". Genetics. 166 (3): 1437–50. doi: 10.1534/genetics.166.3.1437. PMC 1470764. PMID 15082561.
- ^ Podlevsky JD, Bley CJ, Omana RV, Qi X, Chen JJ (January 2008). "The telomerase database". Nucleic Acids Research. 36 (Database issue): D339–43. doi: 10.1093/nar/gkm700. PMC 2238860. PMID 18073191.
- ^ Blevins T, Rajeswaran R, Shivaprasad PV, Beknazariants D, Si-Ammour A, Park HS, Vazquez F, Robertson D, Meins F, Hohn T, Pooggin MM (2006). "Four plant Dicers mediate viral small RNA biogenesis and DNA virus induced silencing". Nucleic Acids Research. 34 (21): 6233–46. doi: 10.1093/nar/gkl886. PMC 1669714. PMID 17090584.
- ^ Jana S, Chakraborty C, Nandi S, Deb JK (November 2004). "RNA interference: potential therapeutic targets". Applied Microbiology and Biotechnology. 65 (6): 649–57. doi: 10.1007/s00253-004-1732-1. PMID 15372214. S2CID 20963666.
- ^ Virol, J (May 2006). "Double-Stranded RNA Is Produced by Positive-Strand RNA Viruses and DNA Viruses but Not in Detectable Amounts by Negative-Strand RNA Viruses". Journal of Virology. 80 (10): 5059–5064. doi: 10.1128/JVI.80.10.5059-5064.2006. PMC 1472073. PMID 16641297.
- ^ Schultz U, Kaspers B, Staeheli P (May 2004). "The interferon system of non-mammalian vertebrates". Developmental and Comparative Immunology. 28 (5): 499–508. doi: 10.1016/j.dci.2003.09.009. PMID 15062646.
- ^ Whitehead KA, Dahlman JE, Langer RS, Anderson DG (2011). "Silencing or stimulation? siRNA delivery and the immune system". Annual Review of Chemical and Biomolecular Engineering. 2: 77–96. doi: 10.1146/annurev-chembioeng-061010-114133. PMID 22432611.
- ^ Hsu MT, Coca-Prados M (July 1979). "Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells". Nature. 280 (5720): 339–40. Bibcode: 1979Natur.280..339H. doi: 10.1038/280339a0. PMID 460409. S2CID 19968869.
- ^ Dahm R (February 2005). "Friedrich Miescher and the discovery of DNA". Developmental Biology. 278 (2): 274–88. doi: 10.1016/j.ydbio.2004.11.028. PMID 15680349.
- ^ Caspersson T, Schultz J (1939). "Pentose nucleotides in the cytoplasm of growing tissues". Nature. 143 (3623): 602–03. Bibcode: 1939Natur.143..602C. doi: 10.1038/143602c0. S2CID 4140563.
- ^ Ochoa S (1959). "Enzymatic synthesis of ribonucleic acid" (PDF). Nobel Lecture.
- ^ Rich A, Davies D (1956). "A New Two-Stranded Helical Structure: Polyadenylic Acid and Polyuridylic Acid". Journal of the American Chemical Society. 78 (14): 3548–49. doi: 10.1021/ja01595a086.
- ^ Holley RW, et al. (March 1965). "Structure of a ribonucleic acid". Science. 147 (3664): 1462–65. Bibcode: 1965Sci...147.1462H. doi: 10.1126/science.147.3664.1462. PMID 14263761. S2CID 40989800.
- ^ Fiers W, et al. (April 1976). "Complete nucleotide sequence of bacteriophage MS2 RNA: primary and secondary structure of the replicase gene". Nature. 260 (5551): 500–07. Bibcode: 1976Natur.260..500F. doi: 10.1038/260500a0. PMID 1264203. S2CID 4289674.
- ^ Napoli C, Lemieux C, Jorgensen R (April 1990). "Introduction of a Chimeric Chalcone Synthase Gene into Petunia Results in Reversible Co-Suppression of Homologous Genes in trans". The Plant Cell. 2 (4): 279–89. doi: 10.1105/tpc.2.4.279. PMC 159885. PMID 12354959.
- ^ Dafny-Yelin M, Chung SM, Frankman EL, Tzfira T (December 2007). "pSAT RNA interference vectors: a modular series for multiple gene down-regulation in plants". Plant Physiology. 145 (4): 1272–81. doi: 10.1104/pp.107.106062. PMC 2151715. PMID 17766396.
- ^ Ruvkun G (October 2001). "Molecular biology. Glimpses of a tiny RNA world". Science. 294 (5543): 797–99. doi: 10.1126/science.1066315. PMID 11679654. S2CID 83506718.
- ^ Fichou Y, Férec C (December 2006). "The potential of oligonucleotides for therapeutic applications". Trends in Biotechnology. 24 (12): 563–70. doi: 10.1016/j.tibtech.2006.10.003. PMID 17045686.
- ^ Siebert S (2006). "Common sequence structure properties and stable regions in RNA secondary structures" (PDF). Dissertation, Albert-Ludwigs-Universität, Freiburg im Breisgau. p. 1. Archived from the original (PDF) on March 9, 2012.
- ^ Szathmáry E (June 1999). "The origin of the genetic code: amino acids as cofactors in an RNA world". Trends in Genetics. 15 (6): 223–29. doi: 10.1016/S0168-9525(99)01730-8. PMID 10354582.
- ^ Jerome, Craig A.; et al. (19 May 2022). "Catalytic Synthesis of Polyribonucleic Acid on Prebiotic Rock Glasses". Astrobiology. 22 (6): 629–636. Bibcode: 2022AsBio..22..629J. doi: 10.1089/ast.2022.0027. PMC 9233534. PMID 35588195. S2CID 248917871.
- ^ Foundation for Applied Molecular Evolution (3 June 2022). "Scientists announce a breakthrough in determining life's origin on Earth—and maybe Mars". Phys.org. Retrieved 3 June 2022.
- ^ Marlaire R (3 March 2015). "NASA Ames Reproduces the Building Blocks of Life in Laboratory". NASA. Archived from the original on 5 March 2015. Retrieved 5 March 2015.
- ^ Starr, Michelle (8 July 2022). "Loads of Precursors For RNA Have Been Detected in The Center of Our Galaxy". ScienceAlert. Retrieved 9 July 2022.
- ^ Rivilla, Victor M.; et al. (8 July 2022). "Molecular Precursors of the RNA-World in Space: New Nitriles in the G+0.693–0.027 Molecular Cloud". Frontiers in Astronomy and Space Sciences. 9: 876870. arXiv: 2206.01053. Bibcode: 2022FrASS...9.6870R. doi: 10.3389/fspas.2022.876870.
- ^ Cech, Thomas R.; Steitz, Joan A. (March 2014). "The Noncoding RNA Revolution—Trashing Old Rules to Forge New Ones". Cell. 157 (1): 77–94. doi: 10.1016/j.cell.2014.03.008. ISSN 0092-8674. PMID 24679528. S2CID 14852160.
- ^ Palacino, James; Swalley, Susanne E; Song, Cheng; Cheung, Atwood K; Shu, Lei; Zhang, Xiaolu; Van Hoosear, Mailin; Shin, Youngah; Chin, Donovan N; Keller, Caroline Gubser; Beibel, Martin; Renaud, Nicole A; Smith, Thomas M; Salcius, Michael; Shi, Xiaoying (2015-06-01). "SMN2 splice modulators enhance U1–pre-mRNA association and rescue SMA mice". Nature Chemical Biology. 11 (7): 511–517. doi: 10.1038/nchembio.1837. ISSN 1552-4450. PMID 26030728.
- ^ Roy, Bijoyita; Friesen, Westley J.; Tomizawa, Yuki; Leszyk, John D.; Zhuo, Jin; Johnson, Briana; Dakka, Jumana; Trotta, Christopher R.; Xue, Xiaojiao; Mutyam, Venkateshwar; Keeling, Kim M.; Mobley, James A.; Rowe, Steven M.; Bedwell, David M.; Welch, Ellen M. (2016-10-04). "Ataluren stimulates ribosomal selection of near-cognate tRNAs to promote nonsense suppression". Proceedings of the National Academy of Sciences. 113 (44): 12508–12513. Bibcode: 2016PNAS..11312508R. doi: 10.1073/pnas.1605336113. ISSN 0027-8424. PMC 5098639. PMID 27702906.
- ^ a b Qadir, Muhammad Imran; Bukhat, Sherien; Rasul, Sumaira; Manzoor, Hamid; Manzoor, Majid (2019-09-03). "RNA therapeutics: Identification of novel targets leading to drug discovery". Journal of Cellular Biochemistry. 121 (2): 898–929. doi: 10.1002/jcb.29364. ISSN 0730-2312. PMID 31478252. S2CID 201806158.
- ^ Balmayor, Elizabeth R.; Geiger, Johannes P.; Aneja, Manish K.; Berezhanskyy, Taras; Utzinger, Maximilian; Mykhaylyk, Olga; Rudolph, Carsten; Plank, Christian (May 2016). "Chemically modified RNA induces osteogenesis of stem cells and human tissue explants as well as accelerates bone healing in rats". Biomaterials. 87: 131–146. doi: 10.1016/j.biomaterials.2016.02.018. ISSN 0142-9612. PMID 26923361.
- ^ Plews, Jordan R.; Li, JianLiang; Jones, Mark; Moore, Harry D.; Mason, Chris; Andrews, Peter W.; Na, Jie (2010-12-30). "Activation of Pluripotency Genes in Human Fibroblast Cells by a Novel mRNA Based Approach". PLOS ONE. 5 (12): e14397. Bibcode: 2010PLoSO...514397P. doi: 10.1371/journal.pone.0014397. ISSN 1932-6203. PMC 3012685. PMID 21209933.
- ^ Preskey, David; Allison, Thomas F.; Jones, Mark; Mamchaoui, Kamel; Unger, Christian (May 2016). "Synthetically modified mRNA for efficient and fast human iPS cell generation and direct transdifferentiation to myoblasts". Biochemical and Biophysical Research Communications. 473 (3): 743–751. doi: 10.1016/j.bbrc.2015.09.102. ISSN 0006-291X. PMID 26449459.
- ^ Warren, Luigi; Manos, Philip D.; Ahfeldt, Tim; Loh, Yuin-Han; Li, Hu; Lau, Frank; Ebina, Wataru; Mandal, Pankaj K.; Smith, Zachary D.; Meissner, Alexander; Daley, George Q.; Brack, Andrew S.; Collins, James J.; Cowan, Chad; Schlaeger, Thorsten M. (November 2010). "Highly Efficient Reprogramming to Pluripotency and Directed Differentiation of Human Cells with Synthetic Modified mRNA". Cell Stem Cell. 7 (5): 618–630. doi: 10.1016/j.stem.2010.08.012. ISSN 1934-5909. PMC 3656821. PMID 20888316.
- ^ Elangovan, Satheesh; Khorsand, Behnoush; Do, Anh-Vu; Hong, Liu; Dewerth, Alexander; Kormann, Michael; Ross, Ryan D.; Rick Sumner, D.; Allamargot, Chantal; Salem, Aliasger K. (November 2015). "Chemically modified RNA activated matrices enhance bone regeneration". Journal of Controlled Release. 218: 22–28. doi: 10.1016/j.jconrel.2015.09.050. ISSN 0168-3659. PMC 4631704. PMID 26415855.
External links
- RNA World website Link collection (structures, sequences, tools, journals)
- Nucleic Acid Database Images of DNA, RNA, and complexes.
- Anna Marie Pyle's Seminar: RNA Structure, Function, and Recognition