Phosphatase and tensin homolog (PTEN) is a phosphatase in humans and is encoded by the PTEN gene. [6] Mutations of this gene are a step in the development of many cancers, specifically glioblastoma, lung cancer, breast cancer, and prostate cancer. Genes corresponding to PTEN ( orthologs) [7] have been identified in most mammals for which complete genome data are available.
PTEN acts as a tumor suppressor gene through the action of its phosphatase protein product. This phosphatase is involved in the regulation of the cell cycle, preventing cells from growing and dividing too rapidly. [8] It is a target of many anticancer drugs.
The protein encoded by this gene is a phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase. It contains a tensin-like domain as well as a catalytic domain similar to that of the dual specificity protein tyrosine phosphatases. Unlike most of the protein tyrosine phosphatases, this protein preferentially dephosphorylates phosphoinositide substrates. It negatively regulates intracellular levels of phosphatidylinositol-3,4,5-trisphosphate in cells and functions as a tumor suppressor by negatively regulating the Akt/PKB signaling pathway. [9]
Function
PTEN protein acts as a phosphatase to dephosphorylate phosphatidylinositol (3,4,5)-trisphosphate (PtdIns (3,4,5)P3 or PIP3). PTEN specifically catalyses the dephosphorylation of the 3` phosphate of the inositol ring in PIP3, resulting in the biphosphate product PIP2 ( PtdIns(4,5)P2). This dephosphorylation is important because it results in inhibition of the Akt signaling pathway, which plays an important role in regulating cellular behaviors such as cell growth, survival, and migration.
PTEN also has weak protein phosphatase activity, but this activity is also crucial for its role as a tumor suppressor. PTEN's protein phosphatase activity may be involved in the regulation of the cell cycle, preventing cells from growing and dividing too rapidly. [8] There have been numerous reported protein substrates for PTEN, including IRS1 [10] and Dishevelled. [11]
Structure
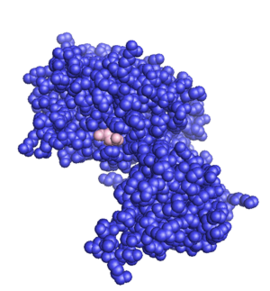
The structure of the core of PTEN (solved by X-ray crystallography, see figure to the upper right [5]) reveals that it consists primarily of a phosphatase domain, and a C2 domain: the phosphatase domain contains the active site, which carries out the enzymatic function of the protein, while the C2 domain binds the phospholipid membrane. Thus PTEN binds the membrane through both its phosphatase and C2 domains, bringing the active site to the membrane-bound PIP3 to dephosphorylate it.
The two domains of PTEN, a protein tyrosine phosphatase domain and a C2 domain, are inherited together as a single unit and thus constitute a superdomain, not only in PTEN but also in various other proteins in fungi, plants and animals, for example, tensin proteins and auxilin. [12]
The active site of PTEN consists of three loops, the TI Loop, the P Loop, and the WPD Loop, all named following the PTPB1 nomenclature. [5] Together they form an unusually deep and wide pocket which allows PTEN to accommodate the bulky phosphatidylinositol 3,4,5-trisphosphate substrate. The dephosphorylation reaction mechanism of PTEN is thought to proceed through a phosphoenzyme intermediate, with the formation of a phosphodiester bond on the active site cysteine, C124.
Not present in the crystal structure of PTEN is a short 10-amino-acid unstructured region N-terminal of the phosphatase domain (from residues 6 to 15), known variously as the PIP2 Binding Domain (PBD) or PIP2 Binding Motif (PBM) [13] [14] [15] This region increases PTEN's affinity for the plasma membrane by binding to Phosphatidylinositol 4,5-bisphosphate, or possibly any anionic lipid.
Also not present in the crystal structure is the intrinsically disordered C-terminal region (CTR) (spanning residues 353–403). The CTR is constitutively phosphorylated at various positions that effect various aspects of PTEN, including its ability to bind to lipid membranes, and also act as either a protein or lipid phosphatase. [16] [17]
Additionally, PTEN can also be expressed as PTEN-L [18] (known as PTEN-Long, or PTEN-α [19]), a leucine initiator alternative start site variant, which adds an additional 173 amino acids to the N-terminus of PTEN. The exact role of this 173-amino acid extension is not yet known, either causing PTEN to be secreted from the cell, or to interact with the mitochondria. The N-terminal extension has been predicted to be largely disordered, [20] although there is evidence that there is some structure in the last twenty amino acids of the extension (most proximal to the start methionine of PTEN). [17]
Clinical significance
Cancer
PTEN is one of the most commonly lost tumor suppressors in human cancer; in fact, up to 70% of men with prostate cancer are estimated to have lost a copy of the PTEN gene at the time of diagnosis. [21] A number of studies have found increased frequency of PTEN loss in tumours which are more highly visible on diagnostic scans such as mpMRI, potentially reflecting increased proliferation and cell density in these tumours. [22]
During tumor development, mutations and deletions of PTEN occur that inactivate its enzymatic activity leading to increased cell proliferation and reduced cell death. Frequent genetic inactivation of PTEN occurs in glioblastoma, endometrial cancer, and prostate cancer; and reduced expression is found in many other tumor types such as lung and breast cancer. Furthermore, PTEN mutation also causes a variety of inherited predispositions to cancer.
Non-cancerous neoplasia
Researchers have identified more than 70 mutations in the PTEN gene in people with Cowden syndrome.[ citation needed] These mutations can be changes in a small number of base pairs or, in some cases, deletions of a large number of base pairs.[ citation needed] Most of these mutations cause the PTEN gene to make a protein that does not function properly or does not work at all. The defective protein is unable to stop cell division or signal abnormal cells to die, which can lead to tumor growth, particularly in the breast, thyroid, or uterus. [23]
Mutations in the PTEN gene cause several other disorders that, like Cowden syndrome, are characterized by the development of non-cancerous tumors called hamartomas. These disorders include Bannayan–Riley–Ruvalcaba syndrome and Proteus-like syndrome. Together, the disorders caused by PTEN mutations are called PTEN hamartoma tumor syndromes, or PHTS. Mutations responsible for these syndromes cause the resulting protein to be non-functional or absent. The defective protein allows the cell to divide in an uncontrolled way and prevents damaged cells from dying, which can lead to the growth of tumors. [23]
Brain function and autism
Defects of the PTEN gene have been cited to be a potential cause of autism spectrum disorders. [24]
When defective, PTEN protein interacts with the protein of a second gene known as Tp53 to dampen energy production in neurons. This severe stress leads to a spike in harmful mitochondrial DNA changes and abnormal levels of energy production in the cerebellum and hippocampus, brain regions critical for social behavior and cognition. When PTEN protein is insufficient, its interaction with p53 triggers deficiencies and defects in other proteins that also have been found in patients with learning disabilities including autism. [24] People with autism and PTEN mutations may have macrocephaly (unusually large heads). [25]
Patients with defective PTEN can develop cerebellar mass lesions called dysplastic gangliocytomas or Lhermitte–Duclos disease. [23]
Cell regeneration
PTEN's strong link to cell growth inhibition is being studied as a possible therapeutic target in tissues that do not traditionally regenerate in mature animals, such as central neurons. PTEN deletion mutants have recently [26] been shown to allow nerve regeneration in mice. [27] [28]
As a drug target
PTEN inhibitors
Bisperoxovanadium compounds may have a neuroprotective effect after CNS injury. [29] PTEN is inhibited by sarcopoterium. [30]
PTEN is one of the targets of the oncomiR, MIRN21. [31]
Cell lines
Cell lines with known PTEN mutations include:
- prostate: LNCaP, PC-3
- kidney: 786-O
- glioblastoma: U87MG [32]
- breast : MB-MDA-468, BT549 [32]
- bladder: J82, UMUC-3
Interactions
PTEN (gene) has been shown to interact with:
See also
References
- ^ a b c ENSG00000284792 GRCh38: Ensembl release 89: ENSG00000171862, ENSG00000284792 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000013663 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b c Lee JO, Yang H, Georgescu MM, Di Cristofano A, Maehama T, Shi Y, et al. (October 1999). "Crystal structure of the PTEN tumor suppressor: implications for its phosphoinositide phosphatase activity and membrane association". Cell. 99 (3): 323–334. doi: 10.1016/S0092-8674(00)81663-3. PMID 10555148. S2CID 5624414.
- ^ Steck PA, Pershouse MA, Jasser SA, Yung WK, Lin H, Ligon AH, et al. (April 1997). "Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers". Nature Genetics. 15 (4): 356–362. doi: 10.1038/ng0497-356. PMID 9090379. S2CID 41286105.
- ^ "OrthoMaM phylogenetic marker: PTEN coding sequence". Archived from the original on 2016-12-27. Retrieved 2009-12-02.
- ^ a b Chu EC, Tarnawski AS (October 2004). "PTEN regulatory functions in tumor suppression and cell biology". Medical Science Monitor. 10 (10): RA235–RA241. PMID 15448614.
- ^ "Entrez Gene: PTEN phosphatase and tensin homolog (mutated in multiple advanced cancers 1)".
- ^ Shi Y, Wang J, Chandarlapaty S, Cross J, Thompson C, Rosen N, Jiang X (June 2014). "PTEN is a protein tyrosine phosphatase for IRS1". Nature Structural & Molecular Biology. 21 (6): 522–527. doi: 10.1038/nsmb.2828. PMC 4167033. PMID 24814346.
- ^ Shnitsar I, Bashkurov M, Masson GR, Ogunjimi AA, Mosessian S, Cabeza EA, et al. (September 2015). "PTEN regulates cilia through Dishevelled". Nature Communications. 6: 8388. Bibcode: 2015NatCo...6.8388S. doi: 10.1038/ncomms9388. PMC 4598566. PMID 26399523.
- ^ Haynie DT, Xue B (May 2015). "Superdomains in the protein structure hierarchy: The case of PTP-C2". Protein Science. 24 (5): 874–882. doi: 10.1002/pro.2664. PMC 4420535. PMID 25694109.
- ^ Campbell RB, Liu F, Ross AH (September 2003). "Allosteric activation of PTEN phosphatase by phosphatidylinositol 4,5-bisphosphate". The Journal of Biological Chemistry. 278 (36): 33617–33620. doi: 10.1074/jbc.C300296200. PMID 12857747.
- ^ Iijima M, Huang YE, Luo HR, Vazquez F, Devreotes PN (April 2004). "Novel mechanism of PTEN regulation by its phosphatidylinositol 4,5-bisphosphate binding motif is critical for chemotaxis". The Journal of Biological Chemistry. 279 (16): 16606–16613. doi: 10.1074/jbc.M312098200. PMID 14764604.
- ^ McConnachie G, Pass I, Walker SM, Downes CP (May 2003). "Interfacial kinetic analysis of the tumour suppressor phosphatase, PTEN: evidence for activation by anionic phospholipids". The Biochemical Journal. 371 (Pt 3): 947–955. doi: 10.1042/BJ20021848. PMC 1223325. PMID 12534371.
- ^ Rahdar M, Inoue T, Meyer T, Zhang J, Vazquez F, Devreotes PN (January 2009). "A phosphorylation-dependent intramolecular interaction regulates the membrane association and activity of the tumor suppressor PTEN". Proceedings of the National Academy of Sciences of the United States of America. 106 (2): 480–485. Bibcode: 2009PNAS..106..480R. doi: 10.1073/pnas.0811212106. PMC 2626728. PMID 19114656.
- ^ a b Masson GR, Perisic O, Burke JE, Williams RL (January 2016). "The intrinsically disordered tails of PTEN and PTEN-L have distinct roles in regulating substrate specificity and membrane activity". The Biochemical Journal. 473 (2): 135–144. doi: 10.1042/BJ20150931. PMC 4700475. PMID 26527737.
- ^ Hopkins BD, Fine B, Steinbach N, Dendy M, Rapp Z, Shaw J, et al. (July 2013). "A secreted PTEN phosphatase that enters cells to alter signaling and survival". Science. 341 (6144): 399–402. Bibcode: 2013Sci...341..399H. doi: 10.1126/science.1234907. PMC 3935617. PMID 23744781.
- ^ Liang H, He S, Yang J, Jia X, Wang P, Chen X, et al. (May 2014). "PTENα, a PTEN isoform translated through alternative initiation, regulates mitochondrial function and energy metabolism". Cell Metabolism. 19 (5): 836–848. doi: 10.1016/j.cmet.2014.03.023. PMC 4097321. PMID 24768297.
- ^ Malaney P, Uversky VN, Davé V (November 2013). "The PTEN Long N-tail is intrinsically disordered: increased viability for PTEN therapy". Molecular BioSystems. 9 (11): 2877–2888. doi: 10.1039/c3mb70267g. PMID 24056727.
- ^ Chen Z, Trotman LC, Shaffer D, Lin HK, Dotan ZA, Niki M, et al. (August 2005). "Crucial role of p53-dependent cellular senescence in suppression of Pten-deficient tumorigenesis". Nature. 436 (7051): 725–730. Bibcode: 2005Natur.436..725C. doi: 10.1038/nature03918. PMC 1939938. PMID 16079851.
- ^ Norris JM, Simpson BS, Parry MA, Allen C, Ball R, Freeman A, et al. (July 2020). "Genetic Landscape of Prostate Cancer Conspicuity on Multiparametric Magnetic Resonance Imaging: A Systematic Review and Bioinformatic Analysis". European Urology Open Science. 20: 37–47. doi: 10.1016/j.euros.2020.06.006. PMC 7497895. PMID 33000006.
- ^ a b c Pilarski R, Eng C (May 2004). "Will the real Cowden syndrome please stand up (again)? Expanding mutational and clinical spectra of the PTEN hamartoma tumour syndrome". Journal of Medical Genetics. 41 (5): 323–326. doi: 10.1136/jmg.2004.018036. PMC 1735782. PMID 15121767.
- ^ a b Napoli E, Ross-Inta C, Wong S, Hung C, Fujisawa Y, Sakaguchi D, et al. (2012). "Mitochondrial dysfunction in Pten haplo-insufficient mice with social deficits and repetitive behavior: interplay between Pten and p53". PLOS ONE. 7 (8): e42504. Bibcode: 2012PLoSO...742504N. doi: 10.1371/journal.pone.0042504. PMC 3416855. PMID 22900024.
- ^ Charney, Dennis S.; Sklar, Pamela B.; Nestler, Eric J.; Buxbaum, Joseph D. (2018). Charney & Nestler's Neurobiology of Mental Illness. Oxford University Press. p. 846. ISBN 9780190681425.
- ^ "Rodent of the Week: Nerves regenerated after spinal cord injury". The Los Angeles Times. August 13, 2010.
- ^ Liu K, Lu Y, Lee JK, Samara R, Willenberg R, Sears-Kraxberger I, et al. (September 2010). "PTEN deletion enhances the regenerative ability of adult corticospinal neurons". Nature Neuroscience. 13 (9): 1075–1081. doi: 10.1038/nn.2603. PMC 2928871. PMID 20694004.
- ^ Leibinger M, Hilla AM, Andreadaki A, Fischer D (2019). "GSK3-CRMP2 signaling mediates axonal regeneration induced by Pten knockout". Communications Biology. 2: 318. doi: 10.1038/s42003-019-0524-1. PMC 6707209. PMID 31453382.
- ^ Walker CL, Walker MJ, Liu NK, Risberg EC, Gao X, Chen J, Xu XM (2012). "Systemic bisperoxovanadium activates Akt/mTOR, reduces autophagy, and enhances recovery following cervical spinal cord injury". PLOS ONE. 7 (1): e30012. Bibcode: 2012PLoSO...730012W. doi: 10.1371/journal.pone.0030012. PMC 3254642. PMID 22253859.
- ^ Rozenberg K, Smirin P, Sampson SR, Rosenzweig T (August 2014). "Insulin-sensitizing and insulin-mimetic activities of Sarcopoterium spinosum extract". Journal of Ethnopharmacology. 155 (1): 362–372. doi: 10.1016/j.jep.2014.05.030. PMID 24882728.
- ^ Xiong B, Cheng Y, Ma L, Zhang C (2013). "MiR-21 regulates biological behavior through the PTEN/PI-3 K/Akt signaling pathway in human colorectal cancer cells". International Journal of Oncology. 42 (1): 219–228. doi: 10.3892/ijo.2012.1707. PMID 23174819.
- ^ a b Li J, Yen C, Liaw D, Podsypanina K, Bose S, Wang SI, et al. (March 1997). "PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer". Science. 275 (5308): 1943–1947. doi: 10.1126/science.275.5308.1943. PMID 9072974. S2CID 23093929.
- ^ a b Miller SJ, Lou DY, Seldin DC, Lane WS, Neel BG (September 2002). "Direct identification of PTEN phosphorylation sites". FEBS Letters. 528 (1–3): 145–153. doi: 10.1016/S0014-5793(02)03274-X. PMID 12297295. S2CID 1093672.
- ^ Wu Y, Dowbenko D, Spencer S, Laura R, Lee J, Gu Q, Lasky LA (July 2000). "Interaction of the tumor suppressor PTEN/MMAC with a PDZ domain of MAGI3, a novel membrane-associated guanylate kinase". The Journal of Biological Chemistry. 275 (28): 21477–21485. doi: 10.1074/jbc.M909741199. PMID 10748157.
- ^ Yu Z, Fotouhi-Ardakani N, Wu L, Maoui M, Wang S, Banville D, Shen SH (October 2002). "PTEN associates with the vault particles in HeLa cells". The Journal of Biological Chemistry. 277 (43): 40247–40252. doi: 10.1074/jbc.M207608200. PMID 12177006.
- ^ Wang X, Shi Y, Wang J, Huang G, Jiang X (September 2008). "Crucial role of the C-terminus of PTEN in antagonizing NEDD4-1-mediated PTEN ubiquitination and degradation". The Biochemical Journal. 414 (2): 221–229. doi: 10.1042/BJ20080674. PMID 18498243.
- ^ Lin HK, Hu YC, Lee DK, Chang C (October 2004). "Regulation of androgen receptor signaling by PTEN (phosphatase and tensin homolog deleted on chromosome 10) tumor suppressor through distinct mechanisms in prostate cancer cells". Molecular Endocrinology. 18 (10): 2409–2423. doi: 10.1210/me.2004-0117. PMID 15205473.
- ^ Freeman DJ, Li AG, Wei G, Li HH, Kertesz N, Lesche R, et al. (February 2003). "PTEN tumor suppressor regulates p53 protein levels and activity through phosphatase-dependent and -independent mechanisms". Cancer Cell. 3 (2): 117–130. doi: 10.1016/S1535-6108(03)00021-7. PMID 12620407.
- ^ Tamura M, Gu J, Danen EH, Takino T, Miyamoto S, Yamada KM (July 1999). "PTEN interactions with focal adhesion kinase and suppression of the extracellular matrix-dependent phosphatidylinositol 3-kinase/Akt cell survival pathway". The Journal of Biological Chemistry. 274 (29): 20693–20703. doi: 10.1074/jbc.274.29.20693. PMID 10400703.
- ^ Haier J, Nicolson GL (February 2002). "PTEN regulates tumor cell adhesion of colon carcinoma cells under dynamic conditions of fluid flow". Oncogene. 21 (9): 1450–1460. doi: 10.1038/sj.onc.1205213. PMID 11857088.
Further reading
- Li J, Yen C, Liaw D, Podsypanina K, Bose S, Wang SI, et al. (March 1997). "PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer". Science. 275 (5308): 1943–1947. doi: 10.1126/science.275.5308.1943. PMID 9072974. S2CID 23093929.
- Simpson L, Parsons R (March 2001). "PTEN: life as a tumor suppressor". Experimental Cell Research. 264 (1): 29–41. doi: 10.1006/excr.2000.5130. PMID 11237521.
- Eng C (September 2003). "PTEN: one gene, many syndromes". Human Mutation. 22 (3): 183–198. doi: 10.1002/humu.10257. PMID 12938083. S2CID 13417857.
- Hamada K, Sasaki T, Koni PA, Natsui M, Kishimoto H, Sasaki J, et al. (September 2005). "The PTEN/PI3K pathway governs normal vascular development and tumor angiogenesis". Genes & Development. 19 (17): 2054–2065. doi: 10.1101/gad.1308805. PMC 1199575. PMID 16107612.
- Leslie NR, Downes CP (August 2004). "PTEN function: how normal cells control it and tumour cells lose it". The Biochemical Journal. 382 (Pt 1): 1–11. doi: 10.1042/BJ20040825. PMC 1133909. PMID 15193142.
- Sansal I, Sellers WR (July 2004). "The biology and clinical relevance of the PTEN tumor suppressor pathway". Journal of Clinical Oncology. 22 (14): 2954–2963. doi: 10.1200/JCO.2004.02.141. PMID 15254063.
- Waite KA, Eng C (April 2002). "Protean PTEN: form and function". American Journal of Human Genetics. 70 (4): 829–844. doi: 10.1086/340026. PMC 379112. PMID 11875759.
- Zhou XP, Waite KA, Pilarski R, Hampel H, Fernandez MJ, Bos C, et al. (August 2003). "Germline PTEN promoter mutations and deletions in Cowden/Bannayan-Riley-Ruvalcaba syndrome result in aberrant PTEN protein and dysregulation of the phosphoinositol-3-kinase/Akt pathway". American Journal of Human Genetics. 73 (2): 404–411. doi: 10.1086/377109. PMC 1180378. PMID 12844284.
- Ji SP, Zhang Y, Van Cleemput J, Jiang W, Liao M, Li L, et al. (March 2006). "Disruption of PTEN coupling with 5-HT2C receptors suppresses behavioral responses induced by drugs of abuse". Nature Medicine. 12 (3): 324–329. doi: 10.1038/nm1349. PMID 16474401. S2CID 22093776.
- Pulido R (May 2015). "PTEN: a yin-yang master regulator protein in health and disease". Methods. 77–78: 3–10. doi: 10.1016/j.ymeth.2015.02.009. PMID 25843297.
- Pulido R (January 2018). "PTEN Inhibition in Human Disease Therapy". Molecules. 23 (2): 285. doi: 10.3390/molecules23020285. PMC 6017825. PMID 29385737.
External links
- GeneReviews/NCBI/NIH/UW entry on PTEN Hamartoma Tumor Syndrome (PHTS)
- PTEN+Protein at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- "PTEN Gene - phosphatase and tensin homolog". GeneCards. The Weizmann Institute of Science. Archived from the original on 2007-10-08. Retrieved 2009-03-12.
- "Gene overview of all published AD-association studies for PTEN". Alzforum: AlzGene. Alzheimer Research Forum. Archived from the original on 2009-02-10. Retrieved 2009-03-12.
- Research shows gene defect's role in autism-like behavior
- Dance Your PhD 2017 : A Story of Tumor Suppression Deepti Mathur. PTEN and cancer explained in dance. A metabolic pathway uses glutamine to create a component of DNA. This pathway is regulated in part by PTEN. Loss of PTEN allows the pathway to go into overdrive, leading to cancer. A drug that interrupts the PTEN pathway preferentially destroys cancer cells.
- PDBe-KB provides an overview of all the structure information available in the PDB for Human Phosphatidylinositol 3,4,5-trisphosphate 3-phosphatase and dual-specificity protein phosphatase PTEN
This article incorporates text from the United States National Library of Medicine, which is in the public domain.





