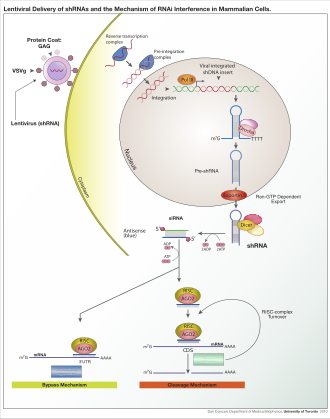
RNA interference (RNAi) is a biological process in which RNA molecules are involved in sequence-specific suppression of gene expression by double-stranded RNA, through translational or transcriptional repression. Historically, RNAi was known by other names, including co-suppression, post-transcriptional gene silencing (PTGS), and quelling. The detailed study of each of these seemingly different processes elucidated that the identity of these phenomena were all actually RNAi. Andrew Fire and Craig C. Mello shared the 2006 Nobel Prize in Physiology or Medicine for their work on RNAi in the nematode worm Caenorhabditis elegans, which they published in 1998. Since the discovery of RNAi and its regulatory potentials, it has become evident that RNAi has immense potential in suppression of desired genes. RNAi is now known as precise, efficient, stable and better than antisense therapy for gene suppression. [1] Antisense RNA produced intracellularly by an expression vector may be developed and find utility as novel therapeutic agents. [2]
Two types of small ribonucleic acid (RNA) molecules, microRNA (miRNA) and small interfering RNA ( siRNA), are central to components to the RNAi pathway. Once mRNA is degraded, post-transcriptional silencing occurs as protein translation is prevented. Transcription can be inhibited via the pre-transcriptional silencing mechanism of RNAi, through which an enzyme complex catalyzes DNA methylation at genomic positions complementary to complexed siRNA or miRNA. RNAi has an important role in defending cells against parasitic nucleotide sequences (e.g., viruses or transposons) and also influences development of organisms.
The RNAi pathway is a naturally occurring process found in many eukaryotes and animal cells. It is initiated by the enzyme Dicer, which cleaves long double-stranded RNA (dsRNA) molecules into short double-stranded fragments of approximately 21 to 23 nucleotide siRNAs. Each siRNA is unwound into two single-stranded RNAs (ssRNAs), the passenger (sense) strand and the guide (antisense) strand. The passenger strand is then cleaved by the protein Argonaute 2 (Ago2). The passenger strand is degraded and the guide strand is incorporated into the RNA-induced silencing complex (RISC). The RISC assembly then binds and degrades the target mRNA. Specifically, this is accomplished when the guide strand pairs with a complementary sequence in a mRNA molecule and induces cleavage by Ago2, a catalytic component of the RISC. In some organisms, this process spreads systemically, despite the initially limited molar concentrations of siRNA. [3]
RNAi is a valuable research tool, both in cell culture and in living organisms, because synthetic dsRNA introduced into cells can selectively and robustly induce suppression of specific genes of interest. RNAi may be used for large-scale screens that systematically shut down each gene (and the subsequent proteins it codes for) in the cell, which can help to identify the components necessary for a particular cellular process or an event such as cell division. The pathway is also used as a practical tool for food, medicine and insecticides. [4]
Cellular mechanism

RNAi is an RNA-dependent gene silencing process that is controlled by RISC and is initiated by short double-stranded RNA molecules in a cell's cytoplasm, where they interact with the catalytic RISC component Argonaute. [6] When the dsRNA is exogenous (coming from infection by a virus with an RNA genome or laboratory manipulations), the RNA is imported directly into the cytoplasm and cleaved to short fragments by Dicer. The initiating dsRNA can also be endogenous (originating in the cell), as in pre-microRNAs expressed from RNA-coding genes in the genome. The primary transcripts from such genes are first processed to form the characteristic stem-loop structure of pre-miRNA in the nucleus, then exported to the cytoplasm. Thus, the two dsRNA pathways, exogenous and endogenous, converge at the RISC. [7]
Exogenous dsRNA initiates RNAi by activating the ribonuclease protein Dicer, [8] which binds and cleaves dsRNAs in plants, or short hairpin RNAs (shRNAs) in humans, to produce double-stranded fragments of 20–25 base pairs with a 2-nucleotide overhang at the 3′ end. [9] Bioinformatics studies on the genomes of multiple organisms suggest this length maximizes target-gene specificity and minimizes non-specific effects. [10] These short double-stranded fragments are called siRNAs. These siRNAs are then separated into single strands and integrated into an active RISC, by RISC-Loading Complex (RLC). RLC includes Dicer-2 and R2D2, and is crucial to unite Ago2 and RISC. [11] TATA-binding protein-associated factor 11 (TAF11) assembles the RLC by facilitating Dcr-2-R2D2 tetramerization, which increases the binding affinity to siRNA by 10-fold. Association with TAF11 would convert the R2-D2-Initiator (RDI) complex into the RLC. [12] R2D2 carries tandem double-stranded RNA-binding domains to recognize the thermodynamically stable terminus of siRNA duplexes, whereas Dicer-2 the other less stable extremity. Loading is asymmetric: the MID domain of Ago2 recognizes the thermodynamically stable end of the siRNA. Therefore, the "passenger" (sense) strand whose 5′ end is discarded by MID is ejected, while the saved "guide" (antisense) strand cooperates with AGO to form the RISC. [11]
After integration into the RISC, siRNAs base-pair to their target mRNA and cleave it, thereby preventing it from being used as a translation template. [13] Differently from siRNA, a miRNA-loaded RISC complex scans cytoplasmic mRNAs for potential complementarity. Instead of destructive cleavage (by Ago2), miRNAs rather target the 3′ untranslated region (UTR) regions of mRNAs where they typically bind with imperfect complementarity, thus blocking the access of ribosomes for translation. [14]
Exogenous dsRNA is detected and bound by an effector protein, known as RDE-4 in C. elegans and R2D2 in Drosophila, that stimulates Dicer activity. [15] The mechanism producing this length specificity is unknown and this protein only binds long dsRNAs. [15]
In C. elegans this initiation response is amplified through the synthesis of a population of 'secondary' siRNAs during which the Dicer-produced initiating or 'primary' siRNAs are used as templates. [16] These 'secondary' siRNAs are structurally distinct from Dicer-produced siRNAs and appear to be produced by an RNA-dependent RNA polymerase (RdRP). [17] [18]
MicroRNA

MicroRNAs (miRNAs) are genomically encoded non-coding RNAs that help regulate gene expression, particularly during development. [19] The phenomenon of RNAi, broadly defined, includes the endogenously induced gene silencing effects of miRNAs as well as silencing triggered by foreign dsRNA. Mature miRNAs are structurally similar to siRNAs produced from exogenous dsRNA, but before reaching maturity, miRNAs must first undergo extensive post-transcriptional modification. A miRNA is expressed from a much longer RNA-coding gene as a primary transcript known as a pri-miRNA which is processed, in the cell nucleus, to a 70-nucleotide stem-loop structure called a pre-miRNA by the microprocessor complex. This complex consists of an RNase III enzyme called Drosha and a dsRNA-binding protein DGCR8. The dsRNA portion of this pre-miRNA is bound and cleaved by Dicer to produce the mature miRNA molecule that can be integrated into the RISC complex; thus, miRNA and siRNA share the same downstream cellular machinery. [20] First, viral encoded miRNA was described in Epstein–Barr virus (EBV). [21] Thereafter, an increasing number of microRNAs have been described in viruses. VIRmiRNA is a comprehensive catalogue covering viral microRNA, their targets and anti-viral miRNAs [22] (see also VIRmiRNA resource: http://crdd.osdd.net/servers/virmirna/).
siRNAs derived from long dsRNA precursors differ from miRNAs in that miRNAs, especially those in animals, typically have incomplete base pairing to a target and inhibit the translation of many different mRNAs with similar sequences. In contrast, siRNAs typically base-pair perfectly and induce mRNA cleavage only in a single, specific target. [23] In Drosophila and C. elegans, miRNA and siRNA are processed by distinct Argonaute proteins and Dicer enzymes. [24] [25]
Three prime untranslated regions and microRNAs
Three prime untranslated regions (3′UTRs) of mRNAs often contain regulatory sequences that post-transcriptionally cause RNAi. Such 3′-UTRs often contain both binding sites for miRNAs as well as for regulatory proteins. By binding to specific sites within the 3′-UTR, miRNAs can decrease gene expression of various mRNAs by either inhibiting translation or directly causing degradation of the transcript. The 3′-UTR also may have silencer regions that bind repressor proteins that inhibit the expression of a mRNA.
The 3′-UTR often contains microRNA response elements (MREs). MREs are sequences to which miRNAs bind. These are prevalent motifs within 3′-UTRs. Among all regulatory motifs within the 3′-UTRs (e.g. including silencer regions), MREs make up about half of the motifs.
As of 2023, the miRBase web site, [26] an archive of miRNA sequences and annotations, listed 28,645 entries in 271 biologic species. Of these, 1,917 miRNAs were in annotated human miRNA loci. miRNAs were predicted to have an average of about four hundred target mRNAs (affecting expression of several hundred genes). [27] Friedman et al. [27] estimate that >45,000 miRNA target sites within human mRNA 3′UTRs are conserved above background levels, and >60% of human protein-coding genes have been under selective pressure to maintain pairing to miRNAs.
Direct experiments show that a single miRNA can reduce the stability of hundreds of unique mRNAs. [28] Other experiments show that a single miRNA may repress the production of hundreds of proteins, but that this repression often is relatively mild (less than 2-fold). [29] [30]
The effects of miRNA dysregulation of gene expression seem to be important in cancer. [31] For instance, in gastrointestinal cancers, nine miRNAs have been identified as epigenetically altered and effective in down regulating DNA repair enzymes. [32]
The effects of miRNA dysregulation of gene expression also seem to be important in neuropsychiatric disorders, such as schizophrenia, bipolar disorder, major depression, Parkinson's disease, Alzheimer's disease and autism spectrum disorders. [33] [34] [35]
RISC activation and catalysis
Exogenous dsRNA is detected and bound by an effector protein, known as RDE-4 in C. elegans and R2D2 in Drosophila, that stimulates Dicer activity. [15] This protein only binds long dsRNAs, but the mechanism producing this length specificity is unknown. [15] This RNA-binding protein then facilitates the transfer of cleaved siRNAs to the RISC complex. [36]
In C. elegans this initiation response is amplified through the synthesis of a population of 'secondary' siRNAs during which the Dicer-produced initiating or 'primary' siRNAs are used as templates. [16] These 'secondary' siRNAs are structurally distinct from Dicer-produced siRNAs and appear to be produced by an RNA-dependent RNA polymerase (RdRP). [17] [18]


The active components of an RNA-induced silencing complex (RISC) are endonucleases called Argonaute proteins, which cleave the target mRNA strand complementary to their bound siRNA. [6] As the fragments produced by Dicer are double-stranded, they could each in theory produce a functional siRNA. However, only one of the two strands, which is known as the guide strand, binds Argonaute and directs gene silencing. The other anti-guide strand or passenger strand is degraded during RISC activation. [37] Although it was first believed that an ATP-dependent helicase separated these two strands, [38] the process proved to be ATP-independent and performed directly by the protein components of RISC. [3] [39] However, an in vitro kinetic analysis of RNAi in the presence and absence of ATP showed that ATP may be required to unwind and remove the cleaved mRNA strand from the RISC complex after catalysis. [40] The guide strand tends to be the one whose 5′ end is less stably paired to its complement, [41] but strand selection is unaffected by the direction in which Dicer cleaves the dsRNA before RISC incorporation. [42] Instead, the R2D2 protein may serve as the differentiating factor by binding the more-stable 5′ end of the passenger strand. [43]
The structural basis for binding of RNA to the Argonaute protein was examined by X-ray crystallography of the binding domain of an RNA-bound Argonaute. Here, the phosphorylated 5′ end of the RNA strand enters a conserved basic surface pocket and makes contacts through a divalent cation (an atom with two positive charges) such as magnesium and by aromatic stacking (a process that allows more than one atom to share an electron by passing it back and forth) between the 5′ nucleotide in the siRNA and a conserved tyrosine residue. This site is thought to form a nucleation site for the binding of the siRNA to its mRNA target. [44] Analysis of the inhibitory effect of mismatches in either the 5’ or 3’ end of the guide strand has demonstrated that the 5’ end of the guide strand is likely responsible for matching and binding the target mRNA, while the 3’ end is responsible for physically arranging target mRNA into a cleavage-favorable RISC region. [40]
It is not understood how the activated RISC complex locates complementary mRNAs within the cell. Although the cleavage process has been proposed to be linked to translation, translation of the mRNA target is not essential for RNAi-mediated degradation. [45] Indeed, RNAi may be more effective against mRNA targets that are not translated. [46] Argonaute proteins are localized to specific regions in the cytoplasm called P-bodies (also cytoplasmic bodies or GW bodies), which are regions with high rates of mRNA decay; [47] miRNA activity is also clustered in P-bodies. [48] Disruption of P-bodies decreases the efficiency of RNAi, suggesting that they are a critical site in the RNAi process. [49]
Transcriptional silencing
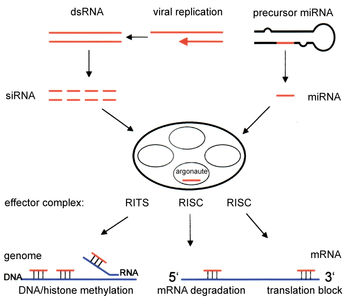
Components of the RNAi pathway are used in many eukaryotes in the maintenance of the organization and structure of their genomes. Modification of histones and associated induction of heterochromatin formation serves to downregulate genes pre- transcriptionally; [51] this process is referred to as RNA-induced transcriptional silencing (RITS), and is carried out by a complex of proteins called the RITS complex. In fission yeast this complex contains Argonaute, a chromodomain protein Chp1, and a protein called Tas3 of unknown function. [52] As a consequence, the induction and spread of heterochromatic regions requires the Argonaute and RdRP proteins. [53] Indeed, deletion of these genes in the fission yeast S. pombe disrupts histone methylation and centromere formation, [54] causing slow or stalled anaphase during cell division. [55] In some cases, similar processes associated with histone modification have been observed to transcriptionally upregulate genes. [56]
The mechanism by which the RITS complex induces heterochromatin formation and organization is not well understood. Most studies have focused on the mating-type region in fission yeast, which may not be representative of activities in other genomic regions/organisms. In maintenance of existing heterochromatin regions, RITS forms a complex with siRNAs complementary to the local genes and stably binds local methylated histones, acting co-transcriptionally to degrade any nascent pre-mRNA transcripts that are initiated by RNA polymerase. The formation of such a heterochromatin region, though not its maintenance, is Dicer-dependent, presumably because Dicer is required to generate the initial complement of siRNAs that target subsequent transcripts. [57] Heterochromatin maintenance has been suggested to function as a self-reinforcing feedback loop, as new siRNAs are formed from the occasional nascent transcripts by RdRP for incorporation into local RITS complexes. [58] The relevance of observations from fission yeast mating-type regions and centromeres to mammals is not clear, as heterochromatin maintenance in mammalian cells may be independent of the components of the RNAi pathway. [59]
Crosstalk with RNA editing
The type of RNA editing that is most prevalent in higher eukaryotes converts adenosine nucleotides into inosine in dsRNAs via the enzyme adenosine deaminase (ADAR). [60] It was originally proposed in 2000 that the RNAi and A→I RNA editing pathways might compete for a common dsRNA substrate. [61] Some pre-miRNAs do undergo A→I RNA editing [62] [63] and this mechanism may regulate the processing and expression of mature miRNAs. [63] Furthermore, at least one mammalian ADAR can sequester siRNAs from RNAi pathway components. [64] Further support for this model comes from studies on ADAR-null C. elegans strains indicating that A→I RNA editing may counteract RNAi silencing of endogenous genes and transgenes. [65]

Variation among organisms
Organisms vary in their ability to take up foreign dsRNA and use it in the RNAi pathway. The effects of RNAi can be both systemic and heritable in plants and C. elegans, although not in Drosophila or mammals. In plants, RNAi is thought to propagate by the transfer of siRNAs between cells through plasmodesmata (channels in the cell walls that enable communication and transport). [38] Heritability comes from methylation of promoters targeted by RNAi; the new methylation pattern is copied in each new generation of the cell. [67] A broad general distinction between plants and animals lies in the targeting of endogenously produced miRNAs; in plants, miRNAs are usually perfectly or nearly perfectly complementary to their target genes and induce direct mRNA cleavage by RISC, while animals' miRNAs tend to be more divergent in sequence and induce translational repression. [66] This translational effect may be produced by inhibiting the interactions of translation initiation factors with the mRNA's polyadenine tail. [68]
Some eukaryotic protozoa such as Leishmania major and Trypanosoma cruzi lack the RNAi pathway entirely. [69] [70] Most or all of the components are also missing in some fungi, most notably the model organism Saccharomyces cerevisiae. [71] The presence of RNAi in other budding yeast species such as Saccharomyces castellii and Candida albicans, further demonstrates that inducing two RNAi-related proteins from S. castellii facilitates RNAi in S. cerevisiae. [72] That certain ascomycetes and basidiomycetes are missing RNAi pathways indicates that proteins required for RNA silencing have been lost independently from many fungal lineages, possibly due to the evolution of a novel pathway with similar function, or to the lack of selective advantage in certain niches. [73]
Related prokaryotic systems
Gene expression in prokaryotes is influenced by an RNA-based system similar in some respects to RNAi. Here, RNA-encoding genes control mRNA abundance or translation by producing a complementary RNA that anneals to an mRNA. However these regulatory RNAs are not generally considered to be analogous to miRNAs because the Dicer enzyme is not involved. [74] It has been suggested that CRISPR interference systems in prokaryotes are analogous to eukaryotic RNAi systems, although none of the protein components are orthologous. [75]
Biological functions
This section needs to be updated. (May 2020) |
Immunity
RNAi is a vital part of the immune response to viruses and other foreign genetic material, especially in plants where it may also prevent the self-propagation of transposons. [76] Plants such as Arabidopsis thaliana express multiple Dicer homologs that are specialized to react differently when the plant is exposed to different viruses. [77] Even before the RNAi pathway was fully understood, it was known that induced gene silencing in plants could spread throughout the plant in a systemic effect and could be transferred from stock to scion plants via grafting. [78] This phenomenon has since been recognized as a feature of the plant immune system which allows the entire plant to respond to a virus after an initial localized encounter. [79] In response, many plant viruses have evolved elaborate mechanisms to suppress the RNAi response. [80] These include viral proteins that bind short double-stranded RNA fragments with single-stranded overhang ends, such as those produced by Dicer. [81] Some plant genomes also express endogenous siRNAs in response to infection by specific types of bacteria. [82] These effects may be part of a generalized response to pathogens that downregulates any metabolic process in the host that aids the infection process. [83]
Although animals generally express fewer variants of the Dicer enzyme than plants, RNAi in some animals produces an antiviral response. In both juvenile and adult Drosophila, RNAi is important in antiviral innate immunity and is active against pathogens such as Drosophila X virus. [84] [85] A similar role in immunity may operate in C. elegans, as Argonaute proteins are upregulated in response to viruses and worms that overexpress components of the RNAi pathway are resistant to viral infection. [86] [87]
The role of RNAi in mammalian innate immunity is poorly understood, and relatively little data is available. However, the existence of viruses that encode genes able to suppress the RNAi response in mammalian cells may be evidence in favour of an RNAi-dependent mammalian immune response, [88] [89] although this hypothesis has been challenged as poorly substantiated. [90] Evidence for the existence of a functional antiviral RNAi pathway in mammalian cells has been presented. [91] [92]
Other functions for RNAi in mammalian viruses also exist, such as miRNAs expressed by the herpes virus that may act as heterochromatin organization triggers to mediate viral latency. [93]
Downregulation of genes
Endogenously expressed miRNAs, including both intronic and intergenic miRNAs, are most important in translational repression [66] and in the regulation of development, especially on the timing of morphogenesis and the maintenance of undifferentiated or incompletely differentiated cell types such as stem cells. [94] The role of endogenously expressed miRNA in downregulating gene expression was first described in C. elegans in 1993. [95] In plants this function was discovered when the "JAW microRNA" of Arabidopsis was shown to be involved in the regulation of several genes that control plant shape. [96] In plants, the majority of genes regulated by miRNAs are transcription factors; [97] thus miRNA activity is particularly wide-ranging and regulates entire gene networks during development by modulating the expression of key regulatory genes, including transcription factors as well as F-box proteins. [98] In many organisms, including humans, miRNAs are linked to the formation of tumors and dysregulation of the cell cycle. Here, miRNAs can function as both oncogenes and tumor suppressors. [99]
Evolution
Based on parsimony-based phylogenetic analysis, the most recent common ancestor of all eukaryotes most likely already possessed an early RNAi pathway; the absence of the pathway in certain eukaryotes is thought to be a derived characteristic. [100] This ancestral RNAi system probably contained at least one Dicer-like protein, one Argonaute, one PIWI protein, and an RNA-dependent RNA polymerase that may also have played other cellular roles. A large-scale comparative genomics study likewise indicates that the eukaryotic crown group already possessed these components, which may then have had closer functional associations with generalized RNA degradation systems such as the exosome. [101] This study also suggests that the RNA-binding Argonaute protein family, which is shared among eukaryotes, most archaea, and at least some bacteria (such as Aquifex aeolicus), is homologous to and originally evolved from components of the translation initiation system.
Applications
RNAi pathway for gene knockdown
Gene knockdown is a method used to reduce the expression of an organism’s specific genes. This is accomplished by using the naturally occurring process of RNAi. [6] This gene knockdown technique uses a double-stranded siRNA molecule that is synthesized with a sequence complementary to the gene of interest. The RNAi cascade begins once the Dicer enzyme starts to process siRNA. The end result of the process leads to degradation of mRNA and destroys any instructions needed to build certain proteins. Using this method, researchers are able to decrease (but not completely eliminate) the expression of a targeted gene. Studying the effects of this decrease in expression may show the physiological role or impact of the targeted gene products. [102] [103]
Off-Target Effects of Gene Knockdown
Extensive efforts in computational biology have been directed toward the design of successful dsRNA reagents that maximize gene knockdown but minimize "off-target" effects. Off-target effects arise when an introduced RNA has a base sequence that can pair with and thus reduce the expression of multiple genes. Such problems occur more frequently when the dsRNA contains repetitive sequences. It has been estimated from studying the genomes of humans, C. elegans and S. pombe that about 10% of possible siRNAs have substantial off-target effects. [10] A multitude of software tools have been developed implementing algorithms for the design of general [104] [105] mammal-specific, [106] and virus-specific [107] siRNAs that are automatically checked for possible cross-reactivity.
Depending on the organism and experimental system, the exogenous RNA may be a long strand designed to be cleaved by Dicer, or short RNAs designed to serve as siRNA substrates. In most mammalian cells, shorter RNAs are used because long double-stranded RNA molecules induce the mammalian interferon response, a form of innate immunity that reacts nonspecifically to foreign genetic material. [108] Mouse oocytes and cells from early mouse embryos lack this reaction to exogenous dsRNA and are therefore a common model system for studying mammalian gene-knockdown effects. [109] Specialized laboratory techniques have also been developed to improve the utility of RNAi in mammalian systems by avoiding the direct introduction of siRNA, for example, by stable transfection with a plasmid encoding the appropriate sequence from which siRNAs can be transcribed, [110] or by more elaborate lentiviral vector systems allowing the inducible activation or deactivation of transcription, known as conditional RNAi. [111] [112]
Medications


The technique of knocking down genes using RNAi therapeutics has demonstrated success in randomized controlled clinical studies. These medications are a growing class of siRNA-based drugs that decrease the expression of proteins encoded by certain genes. To date, five RNAi medications have been approved by regulatory authorities in the US and Europe: patisiran (2018), givosiran (2019), lumasiran (2020), inclisiran (2020 in Europe with anticipated US approval in 2021), and vutrisiran (2022). [113] [114] [115] [116]
While all of the current regulatory body approved RNAi therapeutics focus on diseases that originate in the liver, additional medications under investigation target a host of disease areas including cardiovascular diseases, bleeding disorders, alcohol use disorders, cystic fibrosis, gout, carcinoma, and eye disorders.
Patisiran is the first double stranded siRNA-based medication approved in 2018 and developed by Alnylam Pharmaceuticals. Patisiran uses the RNAi cascade to suppress the gene that codes for TTR (transthryetin). Mutations in this gene may cause the misfolding of a protein responsible for hereditary ATTR amyloidosis. To achieve therapeutic response, patirisan is encased by a lipid nanoparticle membrane that facilitates crossover into the cytoplasm. Once inside the cell, the siRNA begins processing by the enzyme Dicer. Patirisan is administered by a healthcare professional through an intravenous infusion with dosing based on body weight. Warnings and precautions include risk of infusion-related reactions and reduced vitamin A levels (serum). [117]
In 2019, the FDA and EMA approved givosiran for the treatment of adults with acute hepatic porphyria (AHP). [118] The FDA also granted givosiran a breakthrough therapy designation, priority review designation, and orphan drug designation for the treatment of acute hepatic porphyria (AHP) in November 2019. [119] By 2020, givosiran received EMA approval. [120] Givosiran is an siRNA that breaks down aminolevulinic acid synthase 1 (ALAS1) mRNA in the liver. Breaking down ALAS1 mRNA prevents toxins (responsible for neurovisceral attacks and AHP disease) such as aminolevulinic acid (ALA) and porphobilinogen (PBG) from accumulating. [121] [122] [123] [124] To facilitate entry into the cytoplasm, givosiran uses GalNAc ligands and enters into liver cells. The medication is administered subcutaneously by a healthcare professional with dosing based on body weight. Warnings and precautions include risk of anaphylactic reactions, hepatic toxicity, renal toxicity and injection site reactions. [125]
Lumasiran was approved as a siRNA-based medication in 2020 for use in both the European Union and the United States. [126] [127] This medication is used for the treatment of primary hyperoxaluria type 1 (PH1) in pediatric and adult populations. The drug is designed to reduce hepatic oxalate production and urinary oxalate levels through RNAi by targeting hydroxyacid oxidase 1 (HAO1) mRNA for breakdown. Lowering HAO1 enzyme levels reduces the oxidation of glycolate to glyoxylate (which is a substrate for oxalate). Lumasiran is administered subcutaneously by a healthcare professional with dosing based on body weight. [128] Data from randomized controlled clinical trials indicate that the most common adverse reaction that was reported was injection site reactions. These reactions were mild and were present in 38 percent of patients treated with lumasiran. [129]
In 2022, the FDA and EMA approved vutrisiran for the treatment of adults with hereditary transthyretin mediated amyloidosis with polyneuropathy stage 1 or 2. [130] [131] Vutrisiran is designed to break down the mRNA that codes for transthyretin.
Other investigational drugs using RNAi that are being developed by pharmaceutical companies such as Arrowhead Pharmaceuticals, Dicerna, Alnylam Pharmaceuticals, Amgen, and Sylentis. These medications cover a variety of targets via RNAi and diseases.
Investigational RNAi therapeutics in development:
| Drug | Target | Delivery System | Disease | Phase | Status | Company | Identifier |
| ALN–VSP02 | KSP and VEGF | LNP | Solid tumours | I | Completed | Alnylam Pharmaceuticals | NCT01158079 |
| siRNA–EphA2–DOPC | EphA2 | LNP | Advanced cancers | I | Recruiting | MD Anderson Cancer Center | NCT01591356 |
| Atu027 | PKN3 | LNP | Solid tumours | I | Completed | Silence Therapeutics | NCT00938574 |
| TKM–080301 | PLK1 | LNP | Cancer | I | Recruiting | Tekmira Pharmaceutical | NCT01262235 |
| TKM–100201 | VP24, VP35, Zaire Ebola L-polymerase | LNP | Ebola-virus infection | I | Recruiting | Tekmira Pharmaceutical | NCT01518881 |
| ALN–RSV01 | RSV nucleocapsid | Naked siRNA | Respiratory syncytial virus infections | II | Completed | Alnylam Pharmaceuticals | NCT00658086 |
| PRO-040201 | ApoB | LNP | Hypercholesterolaemia | I | Terminated | Tekmira Pharmaceutical | NCT00927459 |
| ALN–PCS02 | PCSK9 | LNP | Hypercholesterolaemia | I | Completed | Alnylam Pharmaceuticals | NCT01437059 |
| ALN–TTR02 | TTR | LNP | Transthyretin-mediated amyloidosis | II | Recruiting | Alnylam Pharmaceuticals | NCT01617967 |
| CALAA-01 | RRM2 | Cyclodextrin NP | Solid tumours | I | Active | Calando Pharmaceuticals | NCT00689065 |
| TD101 | K6a (N171K mutation) | Naked siRNA | Pachyonychia congenita | I | Completed | Pachyonychia Congenita Project | NCT00716014 |
| AGN211745 | VEGFR1 | Naked siRNA | Age-related macular degeneration, choroidal neovascularization | II | Terminated | Allergan | NCT00395057 |
| QPI-1007 | CASP2 | Naked siRNA | Optic atrophy, non-arteritic anterior ischaemic optic neuropathy | I | Completed | Quark Pharmaceuticals | NCT01064505 |
| I5NP | p53 | Naked siRNA | Kidney injury, acute renal failure | I | Completed | Quark Pharmaceuticals | NCT00554359 |
| Delayed graft function, complications of kidney transplant | I, II | Recruiting | Quark Pharmaceuticals | NCT00802347 | |||
| PF-655 (PF-04523655) | RTP801 (Proprietary target) | Naked siRNA | Choroidal neovascularization, diabetic retinopathy, diabetic macular oedema | II | Active | Quark Pharmaceuticals | NCT01445899 |
| siG12D LODER | KRAS | LODER polymer | Pancreatic cancer | II | Recruiting | Silenseed | NCT01676259 |
| Bevasiranib | VEGF | Naked siRNA | Diabetic macular oedema, macular degeneration | II | Completed | Opko Health | NCT00306904 |
| SYL1001 | TRPV1 | Naked siRNA | Ocular pain, dry-eye syndrome | I, II | Recruiting | Sylentis | NCT01776658 |
| SYL040012 | ADRB2 | Naked siRNA | Ocular hypertension, open-angle glaucoma | II | Recruiting | Sylentis | NCT01739244 |
| CEQ508 | CTNNB1 | Escherichia coli-carrying shRNA | Familial adenomatous polyposis | I, II | Recruiting | Marina Biotech | Unknown |
| RXi-109 | CTGF | Self-delivering RNAi compound | Cicatrix scar prevention | I | Recruiting | RXi Pharmaceuticals | NCT01780077 |
| ALN–TTRsc | TTR | siRNA–GalNAc conjugate | Transthyretin-mediated amyloidosis | I | Recruiting | Alnylam Pharmaceuticals | NCT01814839 |
| ARC-520 | Conserved regions of HBV | DPC | HBV | I | Recruiting | Arrowhead Research | NCT01872065 |
Legal categorization and legal issues in a near future
Currently, both miRNA and SiRNA are currently chemically synthesized and so, are legally categorized inside EU and in USA as "simple" medicinal products. But as bioengineered siRNA (BERAs) are in development, these would be classified as biological medicinal products, at least in EU. The development of the BERAs technology raises the question of the categorization of drugs having the same mechanism of action but being produced chemically or biologically. This lack of consistency should be addressed. [132]
Delivery mechanisms
To achieve the clinical potential of RNAi, siRNA must be efficiently transported to the cells of target tissues. However, there are various barriers that must be fixed before it can be used clinically. For example, "naked" siRNA is susceptible to several obstacles that reduce its therapeutic efficacy. [133] Additionally, once siRNA has entered the bloodstream, naked RNA can be degraded by serum nucleases and can stimulate the innate immune system. [133] Due to its size and highly polyanionic (containing negative charges at several sites) nature, unmodified siRNA molecules cannot readily enter the cells through the cell membrane. Therefore, artificial or nanoparticle encapsulated siRNA must be used. If siRNA is transferred across the cell membrane, unintended toxicities can occur if therapeutic doses are not optimized, and siRNAs can exhibit off-target effects (e.g. unintended downregulation of genes with partial sequence complementarity). [134] Even after entering the cells, repeated dosing is required since their effects are diluted at each cell division. In response to these potential issues and barriers, two approaches help facilitate siRNA delivery to target cells: lipid nanoparticles and conjugates. [135]
Lipid nanoparticles
Lipid nanoparticles (LNPs) are based on liposome-like structures that are typically made of an aqueous center surrounded by a lipid shell. [136] A subset of liposomal structures used for delivery drugs to tissues rest in large unilamellar vesicles (LUVs) which may be 100 nm in size. LNP delivery mechanisms have become an increasing source of encasing nucleic acids and may include plasmids, CRISPR and mRNA. [137]
The first approved use of lipid nanoparticles as a drug delivery mechanism began in 2018 with the siRNA drug patisiran, developed by Alnylam Pharmaceuticals. Dicerna Pharmaceuticals, Persomics, Sanofi and Sirna Therapeutics also worked to bring RNAi therapies to market. [138] [139]
Other recent applications include two FDA approved COVID-19 vaccines: mRNA-1273, developed by Moderna. and BNT162b, developed by a collaboration between Pfizer and BioNtech. [140] These two vaccines use lipid nanoparticles to deliver antigen mRNA. Encapsulating the mRNA molecule in lipid nanoparticles was a critical breakthrough for producing viable mRNA vaccines, solving a number of key technical barriers in delivering the mRNA molecule into the host cell as distributed through apolipoprotein E (apoE) in the low-density lipoprotein receptor (LDLR). In December 2020, Novartis announced that positive results from phase III efficacy studies deemed inclisiran was a treatment for heterozygous familial hypercholesterolemia (HeFH) and atherosclerotic cardiovascular disease (ASCVD). [141]
Conjugates
In addition to LNPs, RNAi therapeutics have targeted delivery through siRNA conjugates (e.g., GalNAc, carbohydrates, peptides, aptamers, antibodies). [142] Therapeutics using siRNA conjugates have been developed for rare or genetic diseases such as acute hepatic porphyria (AHP), hemophilia, primary hyperoxaluria (PH) and hereditary ATTR amyloidosis as well as other cardiometabolic diseases such as hypertension and non-alcoholic steatohepatitis (NASH). [143]
Biotechnology
RNAi has been used for a variety of other applications including food, crops and insecticides. The use of the RNAi pathway has developed numerous products such as foods like Arctic apples, nicotine-free tobacco, decaffeinated coffee, nutrient fortified vegetation and hypoallergenic crops. [144] [145] [146] The emerging use of RNAi has the potential to develop many other products for future use.
Viral infection
Antiviral treatment is one of the earliest proposed RNAi-based medical applications, and two different types have been developed. The first type is to target viral RNAs. Many studies have shown that targeting viral RNAs can suppress the replication of numerous viruses, including HIV, [147] HPV, [148] hepatitis A, [149] hepatitis B, [150] influenza virus, [151] [152] [153] [154] respiratory syncytial virus (RSV), [154] SARS coronavirus (SARS-CoV), [154] adenovirus [154] and measles virus. [155] The other strategy is to block the initial viral entries by targeting the host cell genes. [156] For example, suppression of chemokine receptors ( CXCR4 and CCR5) on host cells can prevent HIV viral entry. [157]
Cancer
While traditional chemotherapy can effectively kill cancer cells, lack of specificity for discriminating normal cells and cancer cells in these treatments usually cause severe side effects. Numerous studies have demonstrated that RNAi can provide a more specific approach to inhibit tumor growth by targeting cancer-related genes (i.e., oncogene). [158] It has also been proposed that RNAi can enhance the sensitivity of cancer cells to chemotherapeutic agents, providing a combinatorial therapeutic approach with chemotherapy. [159] Another potential RNAi-based treatment is to inhibit cell invasion and migration. [160]
Compared with chemotherapy or other anti-cancer drugs, there are a lot of advantages of siRNA drug. [161] SiRNA acts on the post-transcriptional stage of gene expression, so it does not modify or change DNA in a deleterious effect. [161] SiRNA can also be used to produce a specific response in a certain type of way, such as by downgrading suppression of gene expression. [161] In a single cancer cell, siRNA can cause dramatic suppression of gene expression with just several copies. [161] This happens by silencing cancer-promoting genes with RNAi, as well as targeting an mRNA sequence. [161]
RNAi drugs treat cancer by silencing certain cancer promoting genes. [161] This is done by complementing the cancer genes with the RNAi, such as keeping the mRNA sequences in accordance with the RNAi drug. [161] Ideally, RNAi is should be injected and/or chemically modified so the RNAi can reach cancer cells more efficiently. [161] RNAi uptake and regulation is controlled by the kidneys. [161]
Neurological diseases
RNAi strategies also show potential for treating neurodegenerative diseases. Studies in cells and in mouse have shown that specifically targeting Amyloid beta-producing genes (e.g. BACE1 and APP) by RNAi can significantly reduced the amount of Aβ peptide which is correlated with the cause of Alzheimer's disease. [162] [163] [164] In addition, this silencing-based approaches also provide promising results in treatment of Parkinson's disease and Polyglutamine disease. [165] [166] [167]
Stimulation of immune response
The human immune system is divided into two separate branches: the innate immune system and the adaptive immune system. [168] The innate immune system is the first defense against infection and responds to pathogens in a generic fashion. [168] On the other hand, the adaptive immune system, a system that was evolved later than the innate, is composed mainly of highly specialized B and T cells that are trained to react to specific portions of pathogenic molecules. [168]
The challenge between old pathogens and new has helped create a system of guarded cells and particles that are called safe framework. [168] This framework has given humans an army of systems that search out and destroy invader particles, such as pathogens, microscopic organisms, parasites, and infections. [168] The mammalian safe framework has developed to incorporate siRNA as a tool to indicate viral contamination, which has allowed siRNA is create an intense innate immune response. [168]
siRNA is controlled by the innate immune system, which can be divided into the acute inflammatory responses and antiviral responses. [168] The inflammatory response is created with signals from small signaling molecules, or cytokines. [168] These include interleukin-1 (IL-1), interleukin-6 (IL-6), interleukin-12 (IL-12) and tumor necrosis factor α (TNF-α). [168] The innate immune system generates inflammation and antiviral responses, which cause the release pattern recognition receptors (PRRs). [168] These receptors help in labeling which pathogens are viruses, fungi, or bacteria. [168] Moreover, the importance of siRNA and the innate immune system is to include more PRRs to help recognize different RNA structures. [168] This makes it more likely for the siRNA to cause an immunostimulant response in the event of the pathogen. [168]
Food
RNAi has been used to genetically engineer plants to produce lower levels of natural plant toxins. Such techniques take advantage of the stable and heritable RNAi phenotype in plant stocks. Cotton seeds are rich in dietary protein but naturally contain the toxic terpenoid product gossypol, making them unsuitable for human consumption. RNAi has been used to produce cotton stocks whose seeds contain reduced levels of delta-cadinene synthase, a key enzyme in gossypol production, without affecting the enzyme's production in other parts of the plant, where gossypol is itself important in preventing damage from plant pests. [169]
Development efforts have successfully reduced the levels of allergens in tomato plants [170] and fortification of plants such as tomatoes with dietary antioxidants. [171] RNAi silencing of alpha-amylase have also been used to decrease Aspergillus flavus fungal growth in maize which would have otherwise contaminated the kernels with dangerous aflatoxins. [172] Silencing lachrymatory factor synthase in onions have produced tearless onions and RNAi has been used in BP1 genes in rapeseeds to improve photosynthesis. [173] SBEIIa and SBEIIb genes in wheat have been targeted in wheat in order to produce higher levels of amylose in order to improve bowel function, [174] and Travella et al. 2006 employed RNAi for functional genomics an investigation of hexaploid bread races, while virus-induced gene silencing (VIGS, a subtype of RNAi) was used by Scofield et al. 2005 to investigate the mechanism of resistance provided by Lr21 against wheat leaf rust in hexaploid wheat. [175]
Insecticides
RNAi is under development as an insecticide, employing multiple approaches, including genetic engineering and topical application. [4] Cells in the midgut of some insects take up the dsRNA molecules in the process referred to as environmental RNAi. [176] In some insects the effect is systemic as the signal spreads throughout the insect's body (referred to as systemic RNAi). [177]
Animals exposed to RNAi at doses millions of times higher than anticipated human exposure levels show no adverse effects. [178] RNAi has varying effects in different species of Lepidoptera (butterflies and moths). [179]
Drosophila spp., Bombyx mori, Locusta spp., Spodoptera spp., Tribolium castaneum, Nilaparvata lugens, Helicoverpa armigera, and Apis mellifera are models that have been widely used to learn how RNAi works within particular taxa of insects. Musca domestica has two Ago2 genes and Glossina morsitans three, as found by Lewis et al. 2016 and Hain et al. 2010. [180] [181] In the case of the miRNA pathway Diuraphis noxia has two Ago1s, M. domestica two Dcr1s, Acyrthosiphon pisum two each of Ago1 and Loqs and Dcr1 and four of Pasha. While in the piRNA, G. morsitans and A. pisum have two or three Ago3s each. [181] This has led to identification of future insecticide development targets, and the modes of action and reasons for insecticide resistance of other insecticides. [181]
Transgenic plants
Transgenic crops have been made to express dsRNA, carefully chosen to silence crucial genes in target pests. These dsRNAs are designed to affect only insects that express specific gene sequences. As a proof of principle, in 2009 a study showed RNAs that could kill any one of four fruit fly species while not harming the other three. [4]
Topical
Alternatively dsRNA can be supplied without genetic engineering. One approach is to add them to irrigation water. The molecules are absorbed into the plants' vascular system and poison insects feeding on them. Another approach involves spraying dsRNA like a conventional pesticide. This would allow faster adaptation to resistance. Such approaches would require low cost sources of dsRNAs that do not currently exist. [4]
Functional genomics
Approaches to the design of genome-wide RNAi libraries can require more sophistication than the design of a single siRNA for a defined set of experimental conditions. Artificial neural networks are frequently used to design siRNA libraries [182] and to predict their likely efficiency at gene knockdown. [183] Mass genomic screening is widely seen as a promising method for genome annotation and has triggered the development of high-throughput screening methods based on microarrays. [184] [185]
Genome-scale screening
Genome-scale RNAi research relies on high-throughput screening (HTS) technology. RNAi HTS technology allows genome-wide loss-of-function screening and is broadly used in the identification of genes associated with specific phenotypes. This technology has been hailed as a potential second genomics wave, following the first genomics wave of gene expression microarray and single nucleotide polymorphism discovery platforms. [186] One major advantage of genome-scale RNAi screening is its ability to simultaneously interrogate thousands of genes. With the ability to generate a large amount of data per experiment, genome-scale RNAi screening has led to an explosion of data generation rates. Exploiting such large data sets is a fundamental challenge, requiring suitable statistics/bioinformatics methods. The basic process of cell-based RNAi screening includes the choice of an RNAi library, robust and stable cell types, transfection with RNAi agents, treatment/incubation, signal detection, analysis and identification of important genes or therapeutical targets. [187]
History
RNAi discovery

The process of RNAi was referred to as "co-suppression" and "quelling" when observed prior to the knowledge of an RNA-related mechanism. The discovery of RNAi was preceded first by observations of transcriptional inhibition by antisense RNA expressed in transgenic plants, [189] and more directly by reports of unexpected outcomes in experiments performed by plant scientists in the United States and the Netherlands in the early 1990s. [190] In an attempt to alter flower colors in petunias, researchers introduced additional copies of a gene encoding chalcone synthase, a key enzyme for flower pigmentation into petunia plants of normally pink or violet flower color. The overexpressed gene was expected to result in darker flowers, but instead caused some flowers to have less visible purple pigment, sometimes in variegated patterns, indicating that the activity of chalcone synthase had been substantially decreased or became suppressed in a context-specific manner. This would later be explained as the result of the transgene being inserted adjacent to promoters in the opposite direction in various positions throughout the genomes of some transformants, thus leading to expression of antisense transcripts and gene silencing when these promoters are active.[ citation needed] Another early observation of RNAi came from a study of the fungus Neurospora crassa, [191] although it was not immediately recognized as related. Further investigation of the phenomenon in plants indicated that the downregulation was due to post-transcriptional inhibition of gene expression via an increased rate of mRNA degradation. [192] This phenomenon was called co-suppression of gene expression, but the molecular mechanism remained unknown. [193]
Not long after, plant virologists working on improving plant resistance to viral diseases observed a similar unexpected phenomenon. While it was known that plants expressing virus-specific proteins showed enhanced tolerance or resistance to viral infection, it was not expected that plants carrying only short, non-coding regions of viral RNA sequences would show similar levels of protection. Researchers believed that viral RNA produced by transgenes could also inhibit viral replication. [194] The reverse experiment, in which short sequences of plant genes were introduced into viruses, showed that the targeted gene was suppressed in an infected plant. [195] This phenomenon was labeled "virus-induced gene silencing" (VIGS), [175] and the set of such phenomena were collectively called post transcriptional gene silencing. [196]
After these initial observations in plants, laboratories searched for this phenomenon in other organisms. [197] [198] The first instance of RNA silencing in animals was documented in 1996, when Guo and Kemphues observed that, by introducing sense and antisense RNA to par-1 mRNA in Caenorhabditis elegans caused degradation of the par-1 message. [199] It was thought that this degradation was triggered by single-stranded RNA (ssRNA), but two years later, in 1998, Fire and Mello discovered that this ability to silence the par-1 gene expression was actually triggered by double-stranded RNA (dsRNA). [199] Craig C. Mello and Andrew Fire's 1998 Nature paper reported a potent gene silencing effect after injecting double stranded RNA into C. elegans. [200] In investigating the regulation of muscle protein production, they observed that neither mRNA nor antisense RNA injections had an effect on protein production, but double-stranded RNA successfully silenced the targeted gene. As a result of this work, they coined the term RNAi. This discovery represented the first identification of the causative agent for the phenomenon. Fire and Mello were awarded the 2006 Nobel Prize in Physiology or Medicine. [6] [201]
RNAi therapeutics
Just after Fire and Mello's ground-breaking discovery, Elbashir et al. discovered, by using synthetically made small interfering RNA (siRNA), it was possible to target the silencing of specific sequences in a gene, rather than silencing the entire gene. [202] Only a year later, McCaffrey and colleagues demonstrated that this sequence-specific silencing had therapeutic applications by targeting a sequence from the Hepatitis C virus in transgenic mice. [203] Since then, multiple researchers have been attempting to expand the therapeutic applications of RNAi, specifically looking to target genes that cause various types of cancer. [204] [205] By 2006, the first applications to reach clinical trials were in the treatment of macular degeneration and respiratory syncytial virus. [206] Four years later the first-in-human Phase I clinical trial was started, using a nanoparticle delivery system to target solid tumors. [207]
The FDA approved the first siRNA-based drug ( patisiran) in 2018. Givosiran and lumasiran later won FDA approval for the treatment of AHP and PH1 in 2019 and 2020, respectively. [113] Inclisiran received EMA approval in 2020 for the treatment of high cholesterol and is currently under review by the FDA. [208]
See also
References
- ^ Saurabh S, Vidyarthi AS, Prasad D (March 2014). "RNA interference: concept to reality in crop improvement". Planta. 239 (3): 543–64. Bibcode: 2014Plant.239..543S. doi: 10.1007/s00425-013-2019-5. PMID 24402564.
- ^ Weiss B, Davidkova G, Zhou LW (March 1999). "Antisense RNA gene therapy for studying and modulating biological processes". Cellular and Molecular Life Sciences. 55 (3): 334–58. doi: 10.1007/s000180050296. PMID 10228554. S2CID 9448271.
- ^ a b Matranga C, Tomari Y, Shin C, Bartel DP, Zamore PD (November 2005). "Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes". Cell. 123 (4): 607–20. doi: 10.1016/j.cell.2005.08.044. PMID 16271386.
- ^ a b c d Kupferschmidt K (August 2013). "A lethal dose of RNA". Science. 341 (6147): 732–3. Bibcode: 2013Sci...341..732K. doi: 10.1126/science.341.6147.732. PMID 23950525.
- ^ Macrae IJ, Zhou K, Li F, Repic A, Brooks AN, Cande WZ, et al. (January 2006). "Structural basis for double-stranded RNA processing by Dicer". Science. 311 (5758): 195–8. Bibcode: 2006Sci...311..195M. doi: 10.1126/science.1121638. PMID 16410517. S2CID 23785494.
- ^ a b c d Daneholt B. "Advanced Information: RNA interference". The Nobel Prize in Physiology or Medicine 2006. Archived from the original on 20 January 2007. Retrieved 25 January 2007.
- ^ Bagasra O, Prilliman KR (August 2004). "RNA interference: the molecular immune system". Journal of Molecular Histology. 35 (6): 545–53. CiteSeerX 10.1.1.456.1701. doi: 10.1007/s10735-004-2192-8. PMID 15614608. S2CID 2966105.
-
^ Bernstein E, Caudy AA, Hammond SM, Hannon GJ (January 2001). "Role for a bidentate ribonuclease in the initiation step of RNA interference". Nature. 409 (6818): 363–6.
Bibcode:
2001Natur.409..363B.
doi:
10.1038/35053110.
PMID
11201747.
S2CID
4371481.

-
^ Siomi H, Siomi MC (January 2009). "On the road to reading the RNA-interference code". Nature. 457 (7228): 396–404.
Bibcode:
2009Natur.457..396S.
doi:
10.1038/nature07754.
PMID
19158785.
S2CID
205215974.
Zamore PD, Tuschl T, Sharp PA, Bartel DP (March 2000). "RNAi: double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals". Cell. 101 (1): 25–33. doi: 10.1016/S0092-8674(00)80620-0. PMID 10778853.
Vermeulen A, Behlen L, Reynolds A, Wolfson A, Marshall WS, Karpilow J, et al. (May 2005). "The contributions of dsRNA structure to Dicer specificity and efficiency". RNA. 11 (5): 674–82. doi: 10.1261/rna.7272305. PMC 1370754. PMID 15811921.
Castanotto D, Rossi JJ (January 2009). "The promises and pitfalls of RNA-interference-based therapeutics". Nature. 457 (7228): 426–33. Bibcode: 2009Natur.457..426C. doi: 10.1038/nature07758. PMC 2702667. PMID 19158789. - ^ a b Qiu S, Adema CM, Lane T (2005). "A computational study of off-target effects of RNA interference". Nucleic Acids Research. 33 (6): 1834–47. doi: 10.1093/nar/gki324. PMC 1072799. PMID 15800213.
- ^ a b Nakanishi K (September 2016). "Anatomy of RISC: how do small RNAs and chaperones activate Argonaute proteins?". Wiley Interdisciplinary Reviews: RNA. 7 (5): 637–60. doi: 10.1002/wrna.1356. PMC 5084781. PMID 27184117.
- ^ Liang C, Wang Y, Murota Y, Liu X, Smith D, Siomi MC, et al. (September 2015). "TAF11 Assembles the RISC Loading Complex to Enhance RNAi Efficiency". Molecular Cell. 59 (5): 807–18. doi: 10.1016/j.molcel.2015.07.006. PMC 4560963. PMID 26257286.
- ^ Ahlquist P (May 2002). "RNA-dependent RNA polymerases, viruses, and RNA silencing". Science. 296 (5571): 1270–3. Bibcode: 2002Sci...296.1270A. doi: 10.1126/science.1069132. PMID 12016304. S2CID 42526536.
- ^ Roberts TC (2015). "The microRNA Machinery". MicroRNA: Basic Science. Advances in Experimental Medicine and Biology. Vol. 887. pp. 15–30. doi: 10.1007/978-3-319-22380-3_2. ISBN 978-3-319-22379-7. PMID 26662984.
- ^ a b c d Parker G, Eckert D, Bass B (2006). "RDE-4 preferentially binds long dsRNA and its dimerization is necessary for cleavage of dsRNA to siRNA". RNA. 12 (5): 807–18. doi: 10.1261/rna.2338706. PMC 1440910. PMID 16603715.
- ^ a b Baulcombe DC (January 2007). "Molecular biology. Amplified silencing". Science. 315 (5809): 199–200. doi: 10.1126/science.1138030. PMID 17218517. S2CID 46285020.
- ^ a b Pak J, Fire A (January 2007). "Distinct populations of primary and secondary effectors during RNAi in C. elegans". Science. 315 (5809): 241–4. Bibcode: 2007Sci...315..241P. doi: 10.1126/science.1132839. PMID 17124291. S2CID 46620298.
- ^ a b Sijen T, Steiner FA, Thijssen KL, Plasterk RH (January 2007). "Secondary siRNAs result from unprimed RNA synthesis and form a distinct class". Science. 315 (5809): 244–7. Bibcode: 2007Sci...315..244S. doi: 10.1126/science.1136699. PMID 17158288. S2CID 9483460.
-
^ Wang QL, Li ZH (May 2007).
"The functions of microRNAs in plants". Frontiers in Bioscience. 12: 3975–82.
doi:
10.2741/2364.
PMC
2851543.
PMID
17485351.
S2CID
23014413.
Zhao Y, Srivastava D (April 2007). "A developmental view of microRNA function". Trends in Biochemical Sciences. 32 (4): 189–97. doi: 10.1016/j.tibs.2007.02.006. PMID 17350266. - ^ Gregory RI, Chendrimada TP, Shiekhattar R (2006). "MicroRNA biogenesis: isolation and characterization of the microprocessor complex". MicroRNA Protocols. Methods in Molecular Biology. Vol. 342. pp. 33–47. doi: 10.1385/1-59745-123-1:33. ISBN 978-1-59745-123-9. PMID 16957365.
- ^ Pfeffer S, Zavolan M, Grässer FA, Chien M, Russo JJ, Ju J, et al. (April 2004). "Identification of virus-encoded microRNAs". Science. 304 (5671): 734–6. Bibcode: 2004Sci...304..734P. doi: 10.1126/science.1096781. PMID 15118162. S2CID 25287167.
- ^ Qureshi A, Thakur N, Monga I, Thakur A, Kumar M (1 January 2014). "VIRmiRNA: a comprehensive resource for experimentally validated viral miRNAs and their targets". Database. 2014: bau103. doi: 10.1093/database/bau103. PMC 4224276. PMID 25380780.
- ^ Pillai RS, Bhattacharyya SN, Filipowicz W (2007). "Repression of protein synthesis by miRNAs: how many mechanisms?". Trends Cell Biol. 17 (3): 118–26. doi: 10.1016/j.tcb.2006.12.007. PMID 17197185.
- ^ Okamura K, Ishizuka A, Siomi H, Siomi M (2004). "Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways". Genes Dev. 18 (14): 1655–66. doi: 10.1101/gad.1210204. PMC 478188. PMID 15231716.
- ^ Lee Y, Nakahara K, Pham J, Kim K, He Z, Sontheimer E, et al. (2004). "Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways". Cell. 117 (1): 69–81. doi: 10.1016/S0092-8674(04)00261-2. PMID 15066283.
- ^ miRBase.org
- ^ a b Friedman RC, Farh KK, Burge CB, Bartel DP (2009). "Most mammalian mRNAs are conserved targets of microRNAs". Genome Res. 19 (1): 92–105. doi: 10.1101/gr.082701.108. PMC 2612969. PMID 18955434.
- ^ Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, et al. (February 2005). "Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs". Nature. 433 (7027): 769–73. Bibcode: 2005Natur.433..769L. doi: 10.1038/nature03315. PMID 15685193. S2CID 4430576.
- ^ Selbach M, Schwanhäusser B, Thierfelder N, Fang Z, Khanin R, Rajewsky N (September 2008). "Widespread changes in protein synthesis induced by microRNAs". Nature. 455 (7209): 58–63. Bibcode: 2008Natur.455...58S. doi: 10.1038/nature07228. PMID 18668040. S2CID 4429008.
- ^ Baek D, Villén J, Shin C, Camargo FD, Gygi SP, Bartel DP (September 2008). "The impact of microRNAs on protein output". Nature. 455 (7209): 64–71. Bibcode: 2008Natur.455...64B. doi: 10.1038/nature07242. PMC 2745094. PMID 18668037.
- ^ Palmero EI, de Campos SG, Campos M, de Souza NC, Guerreiro ID, Carvalho AL, et al. (July 2011). "Mechanisms and role of microRNA deregulation in cancer onset and progression". Genetics and Molecular Biology. 34 (3): 363–70. doi: 10.1590/S1415-47572011000300001. PMC 3168173. PMID 21931505.
- ^ Bernstein C, Bernstein H (May 2015). "Epigenetic reduction of DNA repair in progression to gastrointestinal cancer". World Journal of Gastrointestinal Oncology. 7 (5): 30–46. doi: 10.4251/wjgo.v7.i5.30. PMC 4434036. PMID 25987950.
- ^ Maffioletti E, Tardito D, Gennarelli M, Bocchio-Chiavetto L (2014). "Micro spies from the brain to the periphery: new clues from studies on microRNAs in neuropsychiatric disorders". Frontiers in Cellular Neuroscience. 8: 75. doi: 10.3389/fncel.2014.00075. PMC 3949217. PMID 24653674.
- ^ Mellios N, Sur M (2012). "The Emerging Role of microRNAs in Schizophrenia and Autism Spectrum Disorders". Frontiers in Psychiatry. 3: 39. doi: 10.3389/fpsyt.2012.00039. PMC 3336189. PMID 22539927.
- ^ Geaghan M, Cairns MJ (August 2015). "MicroRNA and Posttranscriptional Dysregulation in Psychiatry". Biological Psychiatry. 78 (4): 231–9. doi: 10.1016/j.biopsych.2014.12.009. hdl: 1959.13/1335073. PMID 25636176.
- ^ Liu Q, Rand TA, Kalidas S, Du F, Kim HE, Smith DP, et al. (September 2003). "R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway". Science. 301 (5641): 1921–5. Bibcode: 2003Sci...301.1921L. doi: 10.1126/science.1088710. PMID 14512631. S2CID 41436233.
- ^ Gregory RI, Chendrimada TP, Cooch N, Shiekhattar R (November 2005). "Human RISC couples microRNA biogenesis and posttranscriptional gene silencing". Cell. 123 (4): 631–40. doi: 10.1016/j.cell.2005.10.022. PMID 16271387.
- ^ a b Lodish H, Berk A, Matsudaira P, Kaiser CA, Krieger M, Scott MP, et al. (2004). Molecular Cell Biology (5th ed.). WH Freeman: New York, NY. ISBN 978-0-7167-4366-8.
- ^ Leuschner PJ, Ameres SL, Kueng S, Martinez J (March 2006). "Cleavage of the siRNA passenger strand during RISC assembly in human cells". EMBO Reports. 7 (3): 314–20. doi: 10.1038/sj.embor.7400637. PMC 1456892. PMID 16439995.
- ^ a b Haley B, Zamore PD (July 2004). "Kinetic analysis of the RNAi enzyme complex". Nature Structural & Molecular Biology. 11 (7): 599–606. doi: 10.1038/nsmb780. PMID 15170178. S2CID 12400060.
- ^ Schwarz DS, Hutvágner G, Du T, Xu Z, Aronin N, Zamore PD (October 2003). "Asymmetry in the assembly of the RNAi enzyme complex". Cell. 115 (2): 199–208. doi: 10.1016/S0092-8674(03)00759-1. PMID 14567917.
- ^ Preall JB, He Z, Gorra JM, Sontheimer EJ (March 2006). "Short interfering RNA strand selection is independent of dsRNA processing polarity during RNAi in Drosophila". Current Biology. 16 (5): 530–5. Bibcode: 2006CBio...16..530P. doi: 10.1016/j.cub.2006.01.061. PMID 16527750.
- ^ Tomari Y, Matranga C, Haley B, Martinez N, Zamore PD (November 2004). "A protein sensor for siRNA asymmetry". Science. 306 (5700): 1377–80. Bibcode: 2004Sci...306.1377T. doi: 10.1126/science.1102755. PMID 15550672. S2CID 31558409.
- ^ Ma JB, Yuan YR, Meister G, Pei Y, Tuschl T, Patel DJ (March 2005). "Structural basis for 5'-end-specific recognition of guide RNA by the A. fulgidus Piwi protein". Nature. 434 (7033): 666–70. Bibcode: 2005Natur.434..666M. doi: 10.1038/nature03514. PMC 4694588. PMID 15800629.
- ^ Sen G, Wehrman T, Blau H (2005). "mRNA translation is not a prerequisite for small interfering RNA-mediated mRNA cleavage". Differentiation. 73 (6): 287–93. doi: 10.1111/j.1432-0436.2005.00029.x. PMID 16138829. S2CID 41117614.
- ^ Gu S, Rossi J (2005). "Uncoupling of RNAi from active translation in mammalian cells". RNA. 11 (1): 38–44. doi: 10.1261/rna.7158605. PMC 1370689. PMID 15574516.
- ^ Sen G, Blau H (2005). "Argonaute 2/RISC resides in sites of mammalian mRNA decay known as cytoplasmic bodies". Nat Cell Biol. 7 (6): 633–6. doi: 10.1038/ncb1265. PMID 15908945. S2CID 6085169.
- ^ Lian S, Jakymiw A, Eystathioy T, Hamel J, Fritzler M, Chan E (2006). "GW bodies, microRNAs and the cell cycle". Cell Cycle. 5 (3): 242–5. doi: 10.4161/cc.5.3.2410. PMID 16418578.
- ^ Jakymiw A, Lian S, Eystathioy T, Li S, Satoh M, Hamel J, et al. (2005). "Disruption of P bodies impairs mammalian RNA interference". Nat Cell Biol. 7 (12): 1267–74. doi: 10.1038/ncb1334. PMID 16284622. S2CID 36630239.
-
^ Hammond S, Bernstein E, Beach D,
Hannon G (2000). "An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells".
Nature. 404 (6775): 293–6.
Bibcode:
2000Natur.404..293H.
doi:
10.1038/35005107.
PMID
10749213.
S2CID
9091863.

- ^ Holmquist GP, Ashley T (2006). "Chromosome organization and chromatin modification: influence on genome function and evolution". Cytogenetic and Genome Research. 114 (2): 96–125. doi: 10.1159/000093326. PMID 16825762. S2CID 29910065.
- ^ Verdel A, Jia S, Gerber S, Sugiyama T, Gygi S, Grewal SI, et al. (January 2004). "RNAi-mediated targeting of heterochromatin by the RITS complex". Science. 303 (5658): 672–6. Bibcode: 2004Sci...303..672V. doi: 10.1126/science.1093686. PMC 3244756. PMID 14704433.
- ^ Irvine DV, Zaratiegui M, Tolia NH, Goto DB, Chitwood DH, Vaughn MW, et al. (August 2006). "Argonaute slicing is required for heterochromatic silencing and spreading". Science. 313 (5790): 1134–7. Bibcode: 2006Sci...313.1134I. doi: 10.1126/science.1128813. PMID 16931764. S2CID 42997104.
- ^ Volpe TA, Kidner C, Hall IM, Teng G, Grewal SI, Martienssen RA (September 2002). "Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi". Science. 297 (5588): 1833–7. Bibcode: 2002Sci...297.1833V. doi: 10.1126/science.1074973. PMID 12193640. S2CID 2613813.
- ^ Volpe T, Schramke V, Hamilton GL, White SA, Teng G, Martienssen RA, et al. (2003). "RNA interference is required for normal centromere function in fission yeast". Chromosome Research. 11 (2): 137–46. doi: 10.1023/A:1022815931524. PMID 12733640. S2CID 23813417.
- ^ Li LC, Okino ST, Zhao H, Pookot D, Place RF, Urakami S, et al. (November 2006). "Small dsRNAs induce transcriptional activation in human cells". Proceedings of the National Academy of Sciences of the United States of America. 103 (46): 17337–42. Bibcode: 2006PNAS..10317337L. doi: 10.1073/pnas.0607015103. PMC 1859931. PMID 17085592.
- ^ Noma K, Sugiyama T, Cam H, Verdel A, Zofall M, Jia S, et al. (November 2004). "RITS acts in cis to promote RNA interference-mediated transcriptional and post-transcriptional silencing". Nature Genetics. 36 (11): 1174–80. doi: 10.1038/ng1452. PMID 15475954.
- ^ Sugiyama T, Cam H, Verdel A, Moazed D, Grewal SI (January 2005). "RNA-dependent RNA polymerase is an essential component of a self-enforcing loop coupling heterochromatin assembly to siRNA production". Proceedings of the National Academy of Sciences of the United States of America. 102 (1): 152–7. Bibcode: 2005PNAS..102..152S. doi: 10.1073/pnas.0407641102. PMC 544066. PMID 15615848.
- ^ Wang F, Koyama N, Nishida H, Haraguchi T, Reith W, Tsukamoto T (June 2006). "The assembly and maintenance of heterochromatin initiated by transgene repeats are independent of the RNA interference pathway in mammalian cells". Molecular and Cellular Biology. 26 (11): 4028–40. doi: 10.1128/MCB.02189-05. PMC 1489094. PMID 16705157.
- ^ Bass BL (2002). "RNA editing by adenosine deaminases that act on RNA". Annual Review of Biochemistry. 71: 817–46. doi: 10.1146/annurev.biochem.71.110601.135501. PMC 1823043. PMID 12045112.
- ^ Bass BL (April 2000). "Double-stranded RNA as a template for gene silencing". Cell. 101 (3): 235–8. doi: 10.1016/S0092-8674(02)71133-1. PMID 10847677.
- ^ Luciano DJ, Mirsky H, Vendetti NJ, Maas S (August 2004). "RNA editing of a miRNA precursor". RNA. 10 (8): 1174–7. doi: 10.1261/rna.7350304. PMC 1370607. PMID 15272117.
- ^ a b Yang W, Chendrimada TP, Wang Q, Higuchi M, Seeburg PH, Shiekhattar R, et al. (January 2006). "Modulation of microRNA processing and expression through RNA editing by ADAR deaminases". Nature Structural & Molecular Biology. 13 (1): 13–21. doi: 10.1038/nsmb1041. PMC 2950615. PMID 16369484.
- ^ Yang W, Wang Q, Howell KL, Lee JT, Cho DS, Murray JM, et al. (February 2005). "ADAR1 RNA deaminase limits short interfering RNA efficacy in mammalian cells". The Journal of Biological Chemistry. 280 (5): 3946–53. doi: 10.1074/jbc.M407876200. PMC 2947832. PMID 15556947.
- ^ Nishikura K (December 2006). "Editor meets silencer: crosstalk between RNA editing and RNA interference". Nature Reviews Molecular Cell Biology. 7 (12): 919–31. doi: 10.1038/nrm2061. PMC 2953463. PMID 17139332.
- ^ a b c Saumet A, Lecellier CH (2006). "Anti-viral RNA silencing: do we look like plants ?". Retrovirology. 3 (1): 3. doi: 10.1186/1742-4690-3-3. PMC 1363733. PMID 16409629.
- ^ Jones L, Ratcliff F, Baulcombe DC (May 2001). "RNA-directed transcriptional gene silencing in plants can be inherited independently of the RNA trigger and requires Met1 for maintenance". Current Biology. 11 (10): 747–57. Bibcode: 2001CBio...11..747J. doi: 10.1016/S0960-9822(01)00226-3. PMID 11378384. S2CID 16789197.
- ^ Humphreys DT, Westman BJ, Martin DI, Preiss T (November 2005). "MicroRNAs control translation initiation by inhibiting eukaryotic initiation factor 4E/cap and poly(A) tail function". Proceedings of the National Academy of Sciences of the United States of America. 102 (47): 16961–6. Bibcode: 2005PNAS..10216961H. doi: 10.1073/pnas.0506482102. PMC 1287990. PMID 16287976.
- ^ DaRocha WD, Otsu K, Teixeira SM, Donelson JE (February 2004). "Tests of cytoplasmic RNA interference (RNAi) and construction of a tetracycline-inducible T7 promoter system in Trypanosoma cruzi". Molecular and Biochemical Parasitology. 133 (2): 175–86. doi: 10.1016/j.molbiopara.2003.10.005. PMID 14698430.
- ^ Robinson KA, Beverley SM (May 2003). "Improvements in transfection efficiency and tests of RNA interference (RNAi) approaches in the protozoan parasite Leishmania". Molecular and Biochemical Parasitology. 128 (2): 217–28. doi: 10.1016/S0166-6851(03)00079-3. PMID 12742588.
- ^ Aravind L, Watanabe H, Lipman DJ, Koonin EV (October 2000). "Lineage-specific loss and divergence of functionally linked genes in eukaryotes". Proceedings of the National Academy of Sciences of the United States of America. 97 (21): 11319–24. Bibcode: 2000PNAS...9711319A. doi: 10.1073/pnas.200346997. PMC 17198. PMID 11016957.
- ^ Drinnenberg IA, Weinberg DE, Xie KT, Mower JP, Wolfe KH, Fink GR, et al. (October 2009). "RNAi in budding yeast". Science. 326 (5952): 544–550. Bibcode: 2009Sci...326..544D. doi: 10.1126/science.1176945. PMC 3786161. PMID 19745116.
- ^ Nakayashiki H, Kadotani N, Mayama S (July 2006). "Evolution and diversification of RNA silencing proteins in fungi" (PDF). Journal of Molecular Evolution. 63 (1): 127–35. Bibcode: 2006JMolE..63..127N. doi: 10.1007/s00239-005-0257-2. PMID 16786437. S2CID 22639035. Archived from the original (PDF) on 6 August 2020. Retrieved 4 December 2019.
- ^ Morita T, Mochizuki Y, Aiba H (March 2006). "Translational repression is sufficient for gene silencing by bacterial small noncoding RNAs in the absence of mRNA destruction". Proceedings of the National Academy of Sciences of the United States of America. 103 (13): 4858–63. Bibcode: 2006PNAS..103.4858M. doi: 10.1073/pnas.0509638103. PMC 1458760. PMID 16549791.
- ^ Makarova KS, Grishin NV, Shabalina SA, Wolf YI, Koonin EV (March 2006). "A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action". Biology Direct. 1: 7. doi: 10.1186/1745-6150-1-7. PMC 1462988. PMID 16545108.
- ^ Stram Y, Kuzntzova L (June 2006). "Inhibition of viruses by RNA interference". Virus Genes. 32 (3): 299–306. doi: 10.1007/s11262-005-6914-0. PMC 7088519. PMID 16732482.
- ^ Blevins T, Rajeswaran R, Shivaprasad PV, Beknazariants D, Si-Ammour A, Park HS, et al. (2006). "Four plant Dicers mediate viral small RNA biogenesis and DNA virus induced silencing". Nucleic Acids Research. 34 (21): 6233–46. doi: 10.1093/nar/gkl886. PMC 1669714. PMID 17090584.
- ^ Palauqui JC, Elmayan T, Pollien JM, Vaucheret H (August 1997). "Systemic acquired silencing: transgene-specific post-transcriptional silencing is transmitted by grafting from silenced stocks to non-silenced scions". The EMBO Journal. 16 (15): 4738–45. doi: 10.1093/emboj/16.15.4738. PMC 1170100. PMID 9303318.
- ^ Voinnet O (August 2001). "RNA silencing as a plant immune system against viruses". Trends in Genetics. 17 (8): 449–59. doi: 10.1016/S0168-9525(01)02367-8. PMID 11485817.
- ^ Lucy AP, Guo HS, Li WX, Ding SW (April 2000). "Suppression of post-transcriptional gene silencing by a plant viral protein localized in the nucleus". The EMBO Journal. 19 (7): 1672–80. doi: 10.1093/emboj/19.7.1672. PMC 310235. PMID 10747034.
- ^ Mérai Z, Kerényi Z, Kertész S, Magna M, Lakatos L, Silhavy D (June 2006). "Double-stranded RNA binding may be a general plant RNA viral strategy to suppress RNA silencing". Journal of Virology. 80 (12): 5747–56. doi: 10.1128/JVI.01963-05. PMC 1472586. PMID 16731914.
- ^ Katiyar-Agarwal S, Morgan R, Dahlbeck D, Borsani O, Villegas A, Zhu JK, et al. (November 2006). "A pathogen-inducible endogenous siRNA in plant immunity". Proceedings of the National Academy of Sciences of the United States of America. 103 (47): 18002–7. Bibcode: 2006PNAS..10318002K. doi: 10.1073/pnas.0608258103. PMC 1693862. PMID 17071740.
- ^ Fritz JH, Girardin SE, Philpott DJ (June 2006). "Innate immune defense through RNA interference". Science's STKE. 2006 (339): pe27. doi: 10.1126/stke.3392006pe27. PMID 16772641. S2CID 33972766.
- ^ Zambon RA, Vakharia VN, Wu LP (May 2006). "RNAi is an antiviral immune response against a dsRNA virus in Drosophila melanogaster". Cellular Microbiology. 8 (5): 880–9. doi: 10.1111/j.1462-5822.2006.00688.x. PMID 16611236. S2CID 32439482.
- ^ Wang XH, Aliyari R, Li WX, Li HW, Kim K, Carthew R, et al. (April 2006). "RNA interference directs innate immunity against viruses in adult Drosophila". Science. 312 (5772): 452–4. Bibcode: 2006Sci...312..452W. doi: 10.1126/science.1125694. PMC 1509097. PMID 16556799.
- ^ Lu R, Maduro M, Li F, Li HW, Broitman-Maduro G, Li WX, et al. (August 2005). "Animal virus replication and RNAi-mediated antiviral silencing in Caenorhabditis elegans". Nature. 436 (7053): 1040–1043. Bibcode: 2005Natur.436.1040L. doi: 10.1038/nature03870. PMC 1388260. PMID 16107851.
- ^ Wilkins C, Dishongh R, Moore SC, Whitt MA, Chow M, Machaca K (August 2005). "RNA interference is an antiviral defence mechanism in Caenorhabditis elegans". Nature. 436 (7053): 1044–7. Bibcode: 2005Natur.436.1044W. doi: 10.1038/nature03957. PMID 16107852. S2CID 4431035.
- ^ Berkhout B, Haasnoot J (May 2006). "The interplay between virus infection and the cellular RNA interference machinery". FEBS Letters. 580 (12): 2896–902. doi: 10.1016/j.febslet.2006.02.070. PMC 7094296. PMID 16563388.
- ^ Schütz S, Sarnow P (January 2006). "Interaction of viruses with the mammalian RNA interference pathway". Virology. 344 (1): 151–7. doi: 10.1016/j.virol.2005.09.034. PMID 16364746.
- ^ Cullen BR (June 2006). "Is RNA interference involved in intrinsic antiviral immunity in mammals?". Nature Immunology. 7 (6): 563–7. doi: 10.1038/ni1352. PMID 16715068. S2CID 23467688.
- ^ Maillard PV, Ciaudo C, Marchais A, Li Y, Jay F, Ding SW, et al. (October 2013). "Antiviral RNA interference in mammalian cells". Science. 342 (6155): 235–8. Bibcode: 2013Sci...342..235M. doi: 10.1126/science.1241930. PMC 3853215. PMID 24115438.
- ^ Li Y, Lu J, Han Y, Fan X, Ding SW (October 2013). "RNA interference functions as an antiviral immunity mechanism in mammals". Science. 342 (6155): 231–4. Bibcode: 2013Sci...342..231L. doi: 10.1126/science.1241911. PMC 3875315. PMID 24115437.
- ^ Li HW, Ding SW (October 2005). "Antiviral silencing in animals". FEBS Letters. 579 (26): 5965–73. doi: 10.1016/j.febslet.2005.08.034. PMC 1350842. PMID 16154568.
- ^ Carrington JC, Ambros V (July 2003). "Role of microRNAs in plant and animal development". Science. 301 (5631): 336–8. Bibcode: 2003Sci...301..336C. doi: 10.1126/science.1085242. PMID 12869753. S2CID 43395657.
- ^ Lee RC, Feinbaum RL, Ambros V (December 1993). "The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14". Cell. 75 (5): 843–54. doi: 10.1016/0092-8674(93)90529-Y. PMID 8252621.
- ^ Palatnik JF, Allen E, Wu X, Schommer C, Schwab R, Carrington JC, et al. (September 2003). "Control of leaf morphogenesis by microRNAs". Nature. 425 (6955): 257–63. Bibcode: 2003Natur.425..257P. doi: 10.1038/nature01958. PMID 12931144. S2CID 992057.
- ^ Zhang B, Pan X, Cobb GP, Anderson TA (January 2006). "Plant microRNA: a small regulatory molecule with big impact". Developmental Biology. 289 (1): 3–16. doi: 10.1016/j.ydbio.2005.10.036. PMID 16325172.
- ^ Jones-Rhoades MW, Bartel DP, Bartel B (2006). "MicroRNAS and their regulatory roles in plants". Annual Review of Plant Biology. 57: 19–53. doi: 10.1146/annurev.arplant.57.032905.105218. PMID 16669754. S2CID 13010154.
- ^ Zhang B, Pan X, Cobb GP, Anderson TA (February 2007). "microRNAs as oncogenes and tumor suppressors". Developmental Biology. 302 (1): 1–12. doi: 10.1016/j.ydbio.2006.08.028. PMID 16989803.
- ^ Cerutti H, Casas-Mollano JA (August 2006). "On the origin and functions of RNA-mediated silencing: from protists to man". Current Genetics. 50 (2): 81–99. doi: 10.1007/s00294-006-0078-x. PMC 2583075. PMID 16691418.
- ^ Anantharaman V, Koonin EV, Aravind L (April 2002). "Comparative genomics and evolution of proteins involved in RNA metabolism". Nucleic Acids Research. 30 (7): 1427–64. doi: 10.1093/nar/30.7.1427. PMC 101826. PMID 11917006.
- ^ Voorhoeve PM, Agami R (January 2003). "Knockdown stands up". Trends in Biotechnology. 21 (1): 2–4. doi: 10.1016/S0167-7799(02)00002-1. PMID 12480342.
- ^ Munkácsy G, Sztupinszki Z, Herman P, Bán B, Pénzváltó Z, Szarvas N, et al. (September 2016). "Validation of RNAi Silencing Efficiency Using Gene Array Data shows 18.5% Failure Rate across 429 Independent Experiments". Molecular Therapy: Nucleic Acids. 5 (9): e366. doi: 10.1038/mtna.2016.66. PMC 5056990. PMID 27673562.
- ^ Naito Y, Yamada T, Matsumiya T, Ui-Tei K, Saigo K, Morishita S (July 2005). "dsCheck: highly sensitive off-target search software for double-stranded RNA-mediated RNA interference". Nucleic Acids Research. 33 (Web Server issue): W589–91. doi: 10.1093/nar/gki419. PMC 1160180. PMID 15980542.
- ^ Henschel A, Buchholz F, Habermann B (July 2004). "DEQOR: a web-based tool for the design and quality control of siRNAs". Nucleic Acids Research. 32 (Web Server issue): W113–20. doi: 10.1093/nar/gkh408. PMC 441546. PMID 15215362.
- ^ Naito Y, Yamada T, Ui-Tei K, Morishita S, Saigo K (July 2004). "siDirect: highly effective, target-specific siRNA design software for mammalian RNA interference". Nucleic Acids Research. 32 (Web Server issue): W124–9. doi: 10.1093/nar/gkh442. PMC 441580. PMID 15215364.
- ^ Naito Y, Ui-Tei K, Nishikawa T, Takebe Y, Saigo K (July 2006). "siVirus: web-based antiviral siRNA design software for highly divergent viral sequences". Nucleic Acids Research. 34 (Web Server issue): W448–50. doi: 10.1093/nar/gkl214. PMC 1538817. PMID 16845046.
- ^ Reynolds A, Anderson EM, Vermeulen A, Fedorov Y, Robinson K, Leake D, et al. (June 2006). "Induction of the interferon response by siRNA is cell type- and duplex length-dependent". RNA. 12 (6): 988–93. doi: 10.1261/rna.2340906. PMC 1464853. PMID 16611941.
- ^ Stein P, Zeng F, Pan H, Schultz RM (October 2005). "Absence of non-specific effects of RNA interference triggered by long double-stranded RNA in mouse oocytes". Developmental Biology. 286 (2): 464–71. doi: 10.1016/j.ydbio.2005.08.015. PMID 16154556.
- ^ Brummelkamp TR, Bernards R, Agami R (April 2002). "A system for stable expression of short interfering RNAs in mammalian cells". Science. 296 (5567): 550–3. Bibcode: 2002Sci...296..550B. doi: 10.1126/science.1068999. hdl: 1874/15573. PMID 11910072. S2CID 18460980.
- ^ Tiscornia G, Tergaonkar V, Galimi F, Verma IM (May 2004). "CRE recombinase-inducible RNA interference mediated by lentiviral vectors". Proceedings of the National Academy of Sciences of the United States of America. 101 (19): 7347–51. Bibcode: 2004PNAS..101.7347T. doi: 10.1073/pnas.0402107101. PMC 409921. PMID 15123829.
- ^ Ventura A, Meissner A, Dillon CP, McManus M, Sharp PA, Van Parijs L, et al. (July 2004). "Cre-lox-regulated conditional RNA interference from transgenes". Proceedings of the National Academy of Sciences of the United States of America. 101 (28): 10380–5. Bibcode: 2004PNAS..10110380V. doi: 10.1073/pnas.0403954101. PMC 478580. PMID 15240889.
- ^ a b Adams D, Gonzalez-Duarte A, O'Riordan WD, Yang CC, Ueda M, Kristen AV, et al. (5 July 2018). "Patisiran, an RNAi Therapeutic, for Hereditary Transthyretin Amyloidosis". New England Journal of Medicine. 379 (1): 11–21. doi: 10.1056/NEJMoa1716153. hdl: 2445/138257. PMID 29972753. S2CID 205102839.
- ^ Balwani M, Sardh E, Ventura P, Peiró PA, Rees DC, Stölzel U, et al. (11 June 2020). "Phase 3 Trial of RNAi Therapeutic Givosiran for Acute Intermittent Porphyria". New England Journal of Medicine. 382 (24): 2289–2301. doi: 10.1056/NEJMoa1913147. ISSN 0028-4793. PMID 32521132.
- ^ Garrelfs SF, Frishberg Y, Hulton SA, Koren MJ, O'Riordan WD, Cochat P, et al. (1 April 2021). "Lumasiran, an RNAi Therapeutic for Primary Hyperoxaluria Type 1". The New England Journal of Medicine. 384 (13): 1216–1226. doi: 10.1056/NEJMoa2021712. ISSN 1533-4406. PMID 33789010.
- ^ Adams D, Tournev IL, Taylor MS, Coelho T, Planté-Bordeneuve V, Berk JL, et al. (March 2023). "Efficacy and safety of vutrisiran for patients with hereditary transthyretin-mediated amyloidosis with polyneuropathy: a randomized clinical trial". Amyloid. 30 (1): 18–26. doi: 10.1080/13506129.2022.2091985. hdl: 1959.4/unsworks_80922. ISSN 1744-2818. PMID 35875890.
- ^ "LABEL:ONPATTRO- patisiran injection, lipid complex". Daily Med. 10 May 2021.
- ^ "Drug Approval Package: GIVLAARI (givosiran)Injection". FDA.
- ^ Gonzalez-Aseguinolaza G (11 June 2020). "Givosiran — Running RNA Interference to Fight Porphyria Attacks". New England Journal of Medicine. 382 (28–4793): 2366–2367. doi: 10.1056/NEJMe2010986. PMID 32521139. S2CID 219592948.
- ^ "Givlaari". European Medicines Agency.
- ^ Puy H, Gouya L, Deybach JC (2010). "Porphyrias". Lancet. 375 (9718): 924–937. doi: 10.1016/S0140-6736(09)61925-5. PMID 20226990. S2CID 208791867.
- ^ Simon A, Pompilus F, Querbes W, Wei A, Strzok S, Penz C, et al. (2018). "Patient Perspective on Acute Intermittent Porphyria with Frequent Attacks: A Disease with Intermittent and Chronic Manifestations". Patient. 11 (5): 527–537. doi: 10.1007/s40271-018-0319-3. PMC 6132435. PMID 29915990.
- ^ Pischik E, Kauppinen R (2015). "An update of clinical management of acute intermittent porphyria". Application of Clinical Genetics. 8: 201–214. doi: 10.2147/TACG.S48605. PMC 4562648. PMID 26366103.
- ^ Bissell DM, Lai JC, Meister RK, Blanc PD (2015). "Role of delta-aminolevulinic acid in the symptoms of acute porphyria". The American Journal of Medicine. 128 (3): 313–317. doi: 10.1016/j.amjmed.2014.10.026. PMC 4339446. PMID 25446301.
- ^ "LABEL:GIVLAARI- givosiran sodium injection, solution". Daily Med. 5 April 2022.
- ^ "Drug Approval Package: OXLUMO". FDA.
- ^ "Oxlumo". European Medicines Agency.
- ^ "LABEL: OXLUMO- lumasiran injection, solution". Daily Med. 2 December 2021.
- ^ Garrelfs SF, Frishberg Y, Hulton SA, Koren MJ, O'Riordan WD, Cochat P, et al. (1 April 2021). "Lumasiran, an RNAi Therapeutic for Primary Hyperoxaluria Type 1". New England Journal of Medicine. 384 (13): 1216–1226. doi: 10.1056/NEJMoa2021712. PMID 33789010. S2CID 232482623.
- ^ Research Cf (27 November 2023). "Drug Trial Snapshots: AMVUTTRA". FDA.
- ^ "Amvuttra | European Medicines Agency". www.ema.europa.eu. Retrieved 11 January 2024.
- ^ Guerriaud M, Kohli E (2022). "RNA-based drugs and regulation: Toward a necessary evolution of the definitions issued from the European union legislation". Frontiers in Medicine. 9. doi: 10.3389/fmed.2022.1012497. ISSN 2296-858X. PMC 9618588. PMID 36325384.
- ^ a b Kanasty R, Dorkin JR, Vegas A, Anderson D (November 2013). "Delivery materials for siRNA therapeutics". Nature Materials. 12 (11): 967–77. Bibcode: 2013NatMa..12..967K. doi: 10.1038/nmat3765. PMID 24150415.
- ^ Wittrup A, Lieberman J (September 2015). "Knocking down disease: a progress report on siRNA therapeutics". Nature Reviews Genetics. 16 (9): 543–52. doi: 10.1038/nrg3978. PMC 4756474. PMID 26281785.
- ^ De-Souza EA, Camara H, Salgueiro WG, Moro RP, Knittel TL, Tonon G, et al. (May 2019). "RNA interference may result in unexpected phenotypes in Caenorhabditis elegans". Nucleic Acids Research. 47 (8): 3957–3969. doi: 10.1093/nar/gkz154. PMC 6486631. PMID 30838421.
- ^ Li W, Szoka FC (24 March 2007). "Lipid-based nanoparticles for nucleic acid delivery". Pharm Res. 24 (3): 438–449. doi: 10.1007/s11095-006-9180-5. PMID 17252188. S2CID 9995555.
-
^ Leung AK, Tam YY, Cullis PR (2014). "Lipid Nanoparticles for Short Interfering RNA Delivery". Nonviral Vectors for Gene Therapy - Lipid- and Polymer-based Gene Transfer. Advances in Genetics. Vol. 88. pp. 71–110.
doi:
10.1016/B978-0-12-800148-6.00004-3.
ISBN
978-0-12-800148-6.
PMC
7149983.
PMID
25409604.
{{ cite book}}:|journal=ignored ( help) - ^ John LaMattina (15 April 2014). "Big Pharma's Turn On RNAi Shows That New Technologies Don't Guarantee R&D Success". Forbes.
- ^ Eric Bender (1 September 2014). "The Second Coming of RNAi". The Scientist.
- ^ Baden LR (2021). "Efficacy and safety of the mRNA-1273 SARS-CoV-2 vaccine". New England Journal of Medicine. 384 (5): 403–416. doi: 10.1056/NEJMoa2035389. PMC 7787219. PMID 33378609.
- ^ Fitzgerald K, White S, Borodovsky A, Bettencourt BR, Strahs A, Clausen V (January 2017). "A Highly Durable RNAi Therapeutic Inhibitor of PCSK9". New England Journal of Medicine. 376 (1): 41–51. doi: 10.1056/NEJMoa1609243. PMC 5778873. PMID 27959715.
- ^ Kruspe S, Giangrande P (2017). "Aptamer-siRNA Chimeras: Discovery, Progress, and Future Prospects". Biomedicines. 5 (4): 45. doi: 10.3390/biomedicines5030045. PMC 5618303. PMID 28792479.
- ^ Balwani M, Sardh E, Ventura P, Peiró PA, Rees DC, Stölzel U, et al. (11 June 2020). "Phase 3 Trial of RNAi Therapeutic Givosiran for Acute Intermittent Porphyria". New England Journal of Medicine. 382 (24): 2289–2301. doi: 10.1056/NEJMoa1913147. PMID 32521132. S2CID 219586624.
- ^ Andersson M, Melander M, Pojmark P, Larsson H, Bülow L, Hofvander P (2006). "Targeted gene suppression by RNA interference: an efficient method for production of high-amylose potato lines". Journal of Biotechnology. 123 (2): 137–148. doi: 10.1016/j.jbiotec.2005.11.001. PMID 16466822.
- ^ Frizzi A, Huang S (2010). "Tapping RNA silencing pathways for plant biotechnology". Plant Biotechnology Journal. 8 (6): 655–677. doi: 10.1111/j.1467-7652.2010.00505.x. PMID 20331529.
- ^ Parrott W, Chassy B, Ligon J, Meyer L, Patrick J, Zhou J, et al. (2010). "Application of food and feed safety assessment principles to evaluate transgenic approaches to gene modulation in crops". British Industrial Biological Research Association. 48 (7): 1773–1790. doi: 10.1016/j.fct.2010.04.017. PMID 20399824.
- ^ Berkhout B (April 2004). "RNA interference as an antiviral approach: targeting HIV-1". Current Opinion in Molecular Therapeutics. 6 (2): 141–5. PMID 15195925.
- ^ Jiang M, Milner J (September 2002). "Selective silencing of viral gene expression in HPV-positive human cervical carcinoma cells treated with siRNA, a primer of RNA interference". Oncogene. 21 (39): 6041–8. doi: 10.1038/sj.onc.1205878. PMID 12203116.
- ^ Kusov Y, Kanda T, Palmenberg A, Sgro JY, Gauss-Müller V (June 2006). "Silencing of hepatitis A virus infection by small interfering RNAs". Journal of Virology. 80 (11): 5599–610. doi: 10.1128/jvi.01773-05. PMC 1472172. PMID 16699041.
- ^ Jia F, Zhang YZ, Liu CM (October 2006). "A retrovirus-based system to stably silence hepatitis B virus genes by RNA interference". Biotechnology Letters. 28 (20): 1679–85. doi: 10.1007/s10529-006-9138-z. PMID 16900331. S2CID 34511611.
- ^ Li YC, Kong LH, Cheng BZ, Li KS (December 2005). "Construction of influenza virus siRNA expression vectors and their inhibitory effects on multiplication of influenza virus". Avian Diseases. 49 (4): 562–73. doi: 10.1637/7365-041205R2.1. PMID 16405000. S2CID 86214047.
- ^ Khanna M, Saxena L, Rajput R, Kumar B, Prasad R (2015). "Gene silencing: a therapeutic approach to combat influenza virus infections". Future Microbiology. 10 (1): 131–40. doi: 10.2217/fmb.14.94. PMID 25598342.
- ^ Rajput R, Khanna M, Kumar P, Kumar B, Sharma S, Gupta N, et al. (December 2012). "Small interfering RNA targeting the nonstructural gene 1 transcript inhibits influenza A virus replication in experimental mice". Nucleic Acid Therapeutics. 22 (6): 414–22. doi: 10.1089/nat.2012.0359. PMID 23062009.
- ^ a b c d Asha K, Kumar P, Sanicas M, Meseko CA, Khanna M, Kumar B (December 2018). "Advancements in Nucleic Acid Based Therapeutics against Respiratory Viral Infections". Journal of Clinical Medicine. 8 (1): 6. doi: 10.3390/jcm8010006. PMC 6351902. PMID 30577479.
- ^ Hu L, Wang Z, Hu C, Liu X, Yao L, Li W, et al. (2005). "Inhibition of Measles virus multiplication in cell culture by RNA interference". Acta Virologica. 49 (4): 227–34. PMID 16402679.
- ^ Qureshi A, Thakur N, Monga I, Thakur A, Kumar M (2014). "VIRmiRNA: a comprehensive resource for experimentally validated viral miRNAs and their targets". Database. 2014. doi: 10.1093/database/bau103. PMC 4224276. PMID 25380780.
- ^ Crowe S (2003). "Suppression of chemokine receptor expression by RNA interference allows for inhibition of HIV-1 replication, by Martínez et al". AIDS. 17 (Suppl 4): S103–5. doi: 10.1097/00002030-200317004-00014. PMID 15080188.
- ^ Fuchs U, Damm-Welk C, Borkhardt A (August 2004). "Silencing of disease-related genes by small interfering RNAs". Current Molecular Medicine. 4 (5): 507–17. doi: 10.2174/1566524043360492. PMID 15267222.
- ^ Cioca DP, Aoki Y, Kiyosawa K (February 2003). "RNA interference is a functional pathway with therapeutic potential in human myeloid leukemia cell lines". Cancer Gene Therapy. 10 (2): 125–33. doi: 10.1038/sj.cgt.7700544. PMID 12536201.
- ^ Lapteva N, Yang AG, Sanders DE, Strube RW, Chen SY (January 2005). "CXCR4 knockdown by small interfering RNA abrogates breast tumor growth in vivo". Cancer Gene Therapy. 12 (1): 84–9. doi: 10.1038/sj.cgt.7700770. PMID 15472715.
- ^ a b c d e f g h i Xu CF, Wang J (1 February 2015). "Delivery systems for siRNA drug development in cancer therapy". Asian Journal of Pharmaceutical Sciences. 10 (1): 1–12. doi: 10.1016/j.ajps.2014.08.011.
- ^ Singer O, Marr RA, Rockenstein E, Crews L, Coufal NG, Gage FH, et al. (October 2005). "Targeting BACE1 with siRNAs ameliorates Alzheimer disease neuropathology in a transgenic model". Nature Neuroscience. 8 (10): 1343–9. doi: 10.1038/nn1531. PMID 16136043. S2CID 6978101.
- ^ Rodríguez-Lebrón E, Gouvion CM, Moore SA, Davidson BL, Paulson HL (September 2009). "Allele-specific RNAi mitigates phenotypic progression in a transgenic model of Alzheimer's disease". Molecular Therapy. 17 (9): 1563–73. doi: 10.1038/mt.2009.123. PMC 2835271. PMID 19532137.
- ^ Piedrahita D, Hernández I, López-Tobón A, Fedorov D, Obara B, Manjunath BS, et al. (October 2010). "Silencing of CDK5 reduces neurofibrillary tangles in transgenic alzheimer's mice". The Journal of Neuroscience. 30 (42): 13966–76. doi: 10.1523/jneurosci.3637-10.2010. PMC 3003593. PMID 20962218.
- ^ Raoul C, Barker SD, Aebischer P (March 2006). "Viral-based modelling and correction of neurodegenerative diseases by RNA interference". Gene Therapy. 13 (6): 487–95. doi: 10.1038/sj.gt.3302690. PMID 16319945.
- ^ Harper SQ, Staber PD, He X, Eliason SL, Martins IH, Mao Q, et al. (April 2005). "RNA interference improves motor and neuropathological abnormalities in a Huntington's disease mouse model". Proceedings of the National Academy of Sciences of the United States of America. 102 (16): 5820–5. Bibcode: 2005PNAS..102.5820H. doi: 10.1073/pnas.0501507102. PMC 556303. PMID 15811941.
- ^ Boudreau RL, Rodríguez-Lebrón E, Davidson BL (April 2011). "RNAi medicine for the brain: progresses and challenges". Human Molecular Genetics. 20 (R1): R21–7. doi: 10.1093/hmg/ddr137. PMC 3095054. PMID 21459775.
- ^ a b c d e f g h i j k l m Whitehead KA, Dahlman JE, Langer RS, Anderson DG (2011). "Silencing or stimulation? siRNA delivery and the immune system". Annual Review of Chemical and Biomolecular Engineering. 2: 77–96. doi: 10.1146/annurev-chembioeng-061010-114133. PMID 22432611. S2CID 28803811.
- ^ Sunilkumar G, Campbell LM, Puckhaber L, Stipanovic RD, Rathore KS (November 2006). "Engineering cottonseed for use in human nutrition by tissue-specific reduction of toxic gossypol". Proceedings of the National Academy of Sciences of the United States of America. 103 (48): 18054–9. doi: 10.1073/pnas.0605389103. PMC 1838705. PMID 17110445.
- ^ Le LQ, Lorenz Y, Scheurer S, Fötisch K, Enrique E, Bartra J, et al. (March 2006). "Design of tomato fruits with reduced allergenicity by dsRNAi-mediated inhibition of ns-LTP (Lyc e 3) expression". Plant Biotechnology Journal. 4 (2): 231–42. doi: 10.1111/j.1467-7652.2005.00175.x. PMID 17177799.
- ^ Niggeweg R, Michael AJ, Martin C (June 2004). "Engineering plants with increased levels of the antioxidant chlorogenic acid". Nature Biotechnology. 22 (6): 746–54. doi: 10.1038/nbt966. PMID 15107863. S2CID 21588259.
- ^ Gilbert MK, Majumdar R, Rajasekaran K, Chen ZY, Wei Q, Sickler CM, et al. (June 2018). "RNA interference-based silencing of the alpha-amylase (amy1) gene in Aspergillus flavus decreases fungal growth and aflatoxin production in maize kernels". Planta. 247 (6): 1465–1473. Bibcode: 2018Plant.247.1465G. doi: 10.1007/s00425-018-2875-0. PMID 29541880. S2CID 3918937.
- ^ Katoch R, Thakur N (March 2013). "RNA interference: a promising technique for the improvement of traditional crops". International Journal of Food Sciences and Nutrition. 64 (2): 248–59. doi: 10.3109/09637486.2012.713918. PMID 22861122. S2CID 45212581.
- ^ Katoch R, Thakur N (March 2013). "Advances in RNA interference technology and its impact on nutritional improvement, disease and insect control in plants". Applied Biochemistry and Biotechnology. 169 (5): 1579–605. doi: 10.1007/s12010-012-0046-5. PMID 23322250. S2CID 23733295.
- ^ a b Downie RC, Lin M, Corsi B, Ficke A, Lillemo M, Oliver RP, et al. (27 July 2021). "Septoria Nodorum Blotch of Wheat: Disease Management and Resistance Breeding in the Face of Shifting Disease Dynamics and a Changing Environment". Phytopathology. 111 (6). American Phytopathological Society: PHYTO–07–20–028. doi: 10.1094/phyto-07-20-0280-rvw. hdl: 20.500.11937/83208. ISSN 0031-949X. PMID 33245254. S2CID 227181536.
- ^ Ivashuta S, Zhang Y, Wiggins BE, Ramaseshadri P, Segers GC, Johnson S, et al. (May 2015). "Environmental RNAi in herbivorous insects". RNA. 21 (5): 840–50. doi: 10.1261/rna.048116.114. PMC 4408792. PMID 25802407.
- ^ Miller SC, Miyata K, Brown SJ, Tomoyasu Y (2012). "Dissecting systemic RNA interference in the red flour beetle Tribolium castaneum: parameters affecting the efficiency of RNAi". PLOS ONE. 7 (10): e47431. Bibcode: 2012PLoSO...747431M. doi: 10.1371/journal.pone.0047431. PMC 3484993. PMID 23133513.
- ^ Petrick JS, Friedrich GE, Carleton SM, Kessenich CR, Silvanovich A, Zhang Y, et al. (November 2016). "Corn rootworm-active RNA DvSnf7: Repeat dose oral toxicology assessment in support of human and mammalian safety". Regulatory Toxicology and Pharmacology. 81: 57–68. doi: 10.1016/j.yrtph.2016.07.009. PMID 27436086.
- ^ Terenius O, Papanicolaou A, Garbutt JS, Eleftherianos I, Huvenne H, Kanginakudru S, et al. (February 2011). "RNA interference in Lepidoptera: an overview of successful and unsuccessful studies and implications for experimental design". Journal of Insect Physiology. 57 (2): 231–45. doi: 10.1016/j.jinsphys.2010.11.006. hdl: 1854/LU-1101411. PMID 21078327.
- ^ Mongelli V, Saleh MC (29 September 2016). "Bugs Are Not to Be Silenced: Small RNA Pathways and Antiviral Responses in Insects" (PDF). Annual Review of Virology. 3 (1). Annual Reviews: 573–589. doi: 10.1146/annurev-virology-110615-042447. ISSN 2327-056X. PMID 27741406. S2CID 38499958.
- ^ a b c Zhu KY, Palli SR (7 January 2020). "Mechanisms, Applications, and Challenges of Insect RNA Interference". Annual Review of Entomology. 65 (1). Annual Reviews: 293–311. doi: 10.1146/annurev-ento-011019-025224. ISSN 0066-4170. PMC 9939233. PMID 31610134. S2CID 204702574.
- ^ Huesken D, Lange J, Mickanin C, Weiler J, Asselbergs F, Warner J, et al. (August 2005). "Design of a genome-wide siRNA library using an artificial neural network". Nature Biotechnology. 23 (8): 995–1001. doi: 10.1038/nbt1118. PMID 16025102. S2CID 11030533.
- ^ Ge G, Wong GW, Luo B (October 2005). "Prediction of siRNA knockdown efficiency using artificial neural network models". Biochemical and Biophysical Research Communications. 336 (2): 723–8. doi: 10.1016/j.bbrc.2005.08.147. PMID 16153609.
- ^ Janitz M, Vanhecke D, Lehrach H (2006). "High-Throughput RNA Interference in Functional Genomics". RNA Towards Medicine. Handbook of Experimental Pharmacology. Vol. 173. pp. 97–104. doi: 10.1007/3-540-27262-3_5. ISBN 978-3-540-27261-8. PMID 16594612.
- ^ Vanhecke D, Janitz M (February 2005). "Functional genomics using high-throughput RNA interference". Drug Discovery Today. 10 (3): 205–12. doi: 10.1016/S1359-6446(04)03352-5. hdl: 11858/00-001M-0000-0010-86E7-8. PMID 15708535. S2CID 9934291.
- ^ Matson RS (2005). Applying genomic and proteomic microarray technology in drug discovery. CRC Press. p. 6. ISBN 978-0-8493-1469-8.
- ^ Zhang XHD (2011). Optimal High-Throughput Screening: Practical Experimental Design and Data Analysis for Genome-scale RNAi Research. Cambridge University Press. pp. ix–xiii. ISBN 978-0-521-73444-8.
- ^ Matzke MA, Matzke AJ (2004). "Planting the Seeds of a New Paradigm". PLOS Biol. 2 (5): e133. doi: 10.1371/journal.pbio.0020133. PMC 406394. PMID 15138502.
- ^ Ecker JR, Davis RW (August 1986). "Inhibition of gene expression in plant cells by expression of antisense RNA". Proceedings of the National Academy of Sciences of the United States of America. 83 (15): 5372–6. Bibcode: 1986PNAS...83.5372E. doi: 10.1073/pnas.83.15.5372. PMC 386288. PMID 16593734.
- ^ Napoli C, Lemieux C, Jorgensen R (April 1990). "Introduction of a Chimeric Chalcone Synthase Gene into Petunia Results in Reversible Co-Suppression of Homologous Genes in trans". The Plant Cell. 2 (4): 279–289. doi: 10.1105/tpc.2.4.279. PMC 159885. PMID 12354959.
- ^ Romano N, Macino G (November 1992). "Quelling: transient inactivation of gene expression in Neurospora crassa by transformation with homologous sequences". Molecular Microbiology. 6 (22): 3343–53. doi: 10.1111/j.1365-2958.1992.tb02202.x. PMID 1484489. S2CID 31234985.
- ^ Van Blokland R, Van der Geest N, Mol JN, Kooter JM (1994). "Transgene-mediated suppression of chalcone synthase expression in Petunia hybrida results from an increase in RNA turnover". Plant J. 6 (6): 861–77. doi: 10.1046/j.1365-313X.1994.6060861.x.
- ^ Mol JN, van der Krol AR (1991). Antisense nucleic acids and proteins: fundamentals and applications. M. Dekker. pp. 4, 136. ISBN 978-0-8247-8516-1.
- ^ Covey S, Al-Kaff N, Lángara A, Turner D (1997). "Plants combat infection by gene silencing". Nature. 385 (6619): 781–2. Bibcode: 1997Natur.385..781C. doi: 10.1038/385781a0. S2CID 43229760.
- ^ Kumagai MH, Donson J, della-Cioppa G, Harvey D, Hanley K, Grill LK (February 1995). "Cytoplasmic inhibition of carotenoid biosynthesis with virus-derived RNA". Proceedings of the National Academy of Sciences of the United States of America. 92 (5): 1679–83. Bibcode: 1995PNAS...92.1679K. doi: 10.1073/pnas.92.5.1679. PMC 42583. PMID 7878039.
- ^ Ratcliff F, Harrison BD, Baulcombe DC (June 1997). "A similarity between viral defense and gene silencing in plants". Science. 276 (5318): 1558–60. doi: 10.1126/science.276.5318.1558. PMID 18610513.
- ^ Guo S, Kemphues KJ (May 1995). "par-1, a gene required for establishing polarity in C. elegans embryos, encodes a putative Ser/Thr kinase that is asymmetrically distributed". Cell. 81 (4): 611–20. doi: 10.1016/0092-8674(95)90082-9. PMID 7758115.
- ^ Pal-Bhadra M, Bhadra U, Birchler JA (August 1997). "Cosuppression in Drosophila: gene silencing of Alcohol dehydrogenase by white-Adh transgenes is Polycomb dependent". Cell. 90 (3): 479–90. doi: 10.1016/S0092-8674(00)80508-5. PMID 9267028.
- ^ a b Sen GL, Blau HM (July 2006). "A brief history of RNAi: the silence of the genes". FASEB Journal. 20 (9): 1293–9. doi: 10.1096/fj.06-6014rev. PMID 16816104. S2CID 12917676.
- ^ Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (February 1998). "Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans". Nature. 391 (6669): 806–11. Bibcode: 1998Natur.391..806F. doi: 10.1038/35888. PMID 9486653. S2CID 4355692.
- ^ Daneholt B (2 October 2006). "The Nobel Prize in Physiology or Medicine 2006". nobelprize.org. Retrieved 30 October 2017.
- ^ Elbashir S, Harborth J, Lendeckel W, et al. (2001). "Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells". Nature. 411 (6836): 494–498. Bibcode: 2001Natur.411..494E. doi: 10.1038/35078107. PMID 11373684. S2CID 710341.
-
^ McCaffrey AP, Meuse L, Pham TT, Conklin DS,
Hannon GJ, Kay MA (July 2002). "RNA interference in adult mice". Nature. 418 (6893): 38–9.
Bibcode:
2002Natur.418...38M.
doi:
10.1038/418038a.
PMID
12097900.
S2CID
4361399.

- ^ Devi GR (September 2006). "siRNA-based approaches in cancer therapy". Cancer Gene Therapy. 13 (9): 819–29. doi: 10.1038/sj.cgt.7700931. PMID 16424918.
- ^ Wall NR, Shi Y (October 2003). "Small RNA: can RNA interference be exploited for therapy?". Lancet. 362 (9393): 1401–3. doi: 10.1016/s0140-6736(03)14637-5. PMID 14585643. S2CID 25034627.
- ^ Sah D (2006). "Therapeutic potential of RNA interference for neurological disorders". Life Sci. 79 (19): 1773–80. doi: 10.1016/j.lfs.2006.06.011. PMID 16815477.
- ^ Davis ME, Zuckerman JE, Choi CH, Seligson D, Tolcher A, Alabi CA, et al. (April 2010). "Evidence of RNAi in humans from systemically administered siRNA via targeted nanoparticles". Nature. 464 (7291): 1067–70. Bibcode: 2010Natur.464.1067D. doi: 10.1038/nature08956. PMC 2855406. PMID 20305636.
- ^ "Novartis receives EU approval for Leqvio(inclisiran), a first-in-class siRNA to lower cholesterol with two doses a year". Novartis. 11 December 2020.
External links
- Overview of the RNAi process, from Cambridge University's The Naked Scientists
- Animation of the RNAi process, from Nature
- NOVA scienceNOW explains RNAi – A 15-minute video of the Nova broadcast that aired on PBS, 26 July 2005
- Silencing Genomes Archived 10 August 2019 at the Wayback Machine RNA interference (RNAi) experiments and bioinformatics in C. elegans for education. From the Dolan DNA Learning Center of Cold Spring Harbor Laboratory.
- RNAi screens in C. elegans in a 96-well liquid format and their application to the systematic identification of genetic interactions (a protocol)
- 2 American ‘Worm People’ Win Nobel for RNA Work, from NY Times
- Molecular Therapy web focus: "The development of RNAi as a therapeutic strategy", a collection of free articles about RNAi as a therapeutic strategy.
- GenomeRNAi: a database of phenotypes from RNA interference screening experiments in Drosophila melanogaster and Homo sapiens
- RNAi tools Archived 19 June 2018 at the Wayback Machine Pre-designed and custom RNA Interference tools