| Caenorhabditis elegans | |
|---|---|
| |
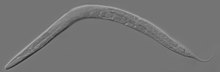
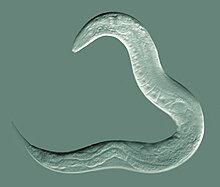
| An adult hermaphrodite C. elegans worm | |
|
Scientific classification
| |
| Domain: | Eukaryota |
| Kingdom: | Animalia |
| Phylum: | Nematoda |
| Class: | Chromadorea |
| Order: | Rhabditida |
| Family: | Rhabditidae |
| Genus: | Caenorhabditis |
| Species: | C. elegans
|
| Binomial name | |
| Caenorhabditis elegans | |
| Subspecies | |
Caenorhabditis elegans ( /ˌsiːnoʊræbˈdaɪtəs ˈɛləɡæns/ [6]) is a free-living transparent nematode about 1 mm in length [7] that lives in temperate soil environments. It is the type species of its genus. [8] The name is a blend of the Greek caeno- (recent), rhabditis (rod-like) [9] and Latin elegans (elegant). In 1900, Maupas initially named it Rhabditides elegans. Osche placed it in the subgenus Caenorhabditis in 1952, and in 1955, Dougherty raised Caenorhabditis to the status of genus. [10]
C. elegans is an unsegmented pseudocoelomate and lacks respiratory or circulatory systems. [11] Most of these nematodes are hermaphrodites and a few are males. [12] Males have specialised tails for mating that include spicules.
In 1963, Sydney Brenner proposed research into C. elegans, primarily in the area of neuronal development. In 1974, he began research into the molecular and developmental biology of C. elegans, which has since been extensively used as a model organism. [13] It was the first multicellular organism to have its whole genome sequenced, and in 2019 [14] it was the first organism to have its connectome (neuronal "wiring diagram") completed. [15] [16] [17]
Anatomy

C. elegans is unsegmented, vermiform, and bilaterally symmetrical. It has a cuticle (a strong outer covering, as an exoskeleton), four main epidermal cords, and a fluid-filled pseudocoelom (body cavity). It also has some of the same organ systems as larger animals. About one in a thousand individuals is male and the rest are hermaphrodites. [18] The basic anatomy of C. elegans includes a mouth, pharynx, intestine, gonad, and collagenous cuticle. Like all nematodes, they have neither a circulatory nor a respiratory system. The four bands of muscles that run the length of the body are connected to a neural system that allows the muscles to move the animal's body only as dorsal bending or ventral bending, but not left or right, except for the head, where the four muscle quadrants are wired independently from one another. When a wave of dorsal/ventral muscle contractions proceeds from the back to the front of the animal, the animal is propelled backwards. When a wave of contractions is initiated at the front and proceeds posteriorly along the body, the animal is propelled forwards. Because of this dorsal/ventral bias in body bends, any normal living, moving individual tends to lie on either its left side or its right side when observed crossing a horizontal surface. A set of ridges on the lateral sides of the body cuticle, the alae, is believed to give the animal added traction during these bending motions.



In relation to lipid metabolism, C. elegans does not have any specialized adipose tissues, a pancreas, a liver, or even blood to deliver nutrients compared to mammals. Neutral lipids are instead stored in the intestine, epidermis, and embryos. The epidermis corresponds to the mammalian adipocytes by being the main triglyceride depot. [19]
The pharynx is a muscular food pump in the head of C. elegans, which is triangular in cross-section. This grinds food and transports it directly to the intestine. A set of "valve cells" connects the pharynx to the intestine, but how this valve operates is not understood. After digestion, the contents of the intestine are released via the rectum, as is the case with all other nematodes. [20] No direct connection exists between the pharynx and the excretory canal, which functions in the release of liquid urine.
Males have a single-lobed gonad, a vas deferens, and a tail specialized for mating, which incorporates spicules. Hermaphrodites have two ovaries, oviducts, and spermatheca, and a single uterus.
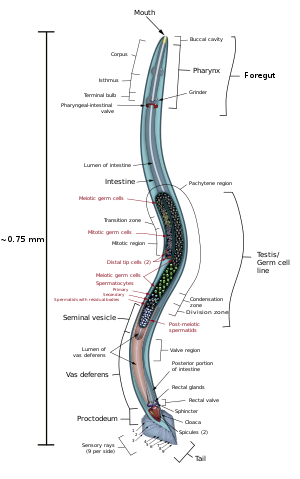
There are 302 neurons in C. elegans, approximately one-third of all the cells in the whole body. [21] Neurons contain dendrites which extend from the cell to receive neurotransmitters, and a process that extends to the nerve ring (the "brain") for a synaptic connection between neurons. [22] C. elegans has motor excitatory and inhibitory neurons, known as cholinergic and gabaergic neurons, which simply act as further regulation for the tiny creature. They have no influence on the nervous system besides regulating neuron impulses. [23]
Gut granules
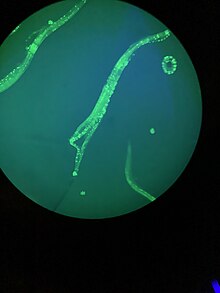
Numerous gut granules are present in the intestine of C. elegans, the functions of which are still not fully known, as are many other aspects of this nematode, despite the many years that it has been studied. These gut granules are found in all of the Rhabditida orders. They are very similar to lysosomes in that they feature an acidic interior and the capacity for endocytosis, but they are considerably larger, reinforcing the view of their being storage organelles. A particular feature of the granules is that when they are observed under ultraviolet light, they react by emitting an intense blue fluorescence. Another phenomenon seen is termed 'death fluorescence'. As the worms die, a dramatic burst of blue fluorescence is emitted. This death fluorescence typically takes place in an anterior to posterior wave that moves along the intestine, and is seen in both young and old worms, whether subjected to lethal injury or peacefully dying of old age.
Many theories have been posited on the functions of the gut granules, with earlier ones being eliminated by later findings. They are thought to store zinc as one of their functions. Recent chemical analysis has identified the blue fluorescent material they contain as a glycosylated form of anthranilic acid (AA). The need for the large amounts of AA the many gut granules contain is questioned. One possibility is that the AA is antibacterial and used in defense against invading pathogens. Another possibility is that the granules provide photoprotection; the bursts of AA fluorescence entail the conversion of damaging UV light to relatively harmless visible light. This is seen a possible link to the melanin–containing melanosomes. [24]
Reproduction
The hermaphroditic worm is considered to be a specialized form of self-fertile female, as its soma is female. The hermaphroditic germline produces male gametes first, and lays eggs through its uterus after internal fertilization. Hermaphrodites produce all their sperm in the L4 stage (150 sperm cells per gonadal arm) and then produce only oocytes. The hermaphroditic gonad acts as an ovotestis with sperm cells being stored in the same area of the gonad as the oocytes until the first oocyte pushes the sperm into the spermatheca (a chamber wherein the oocytes become fertilized by the sperm). [25]
The male can inseminate the hermaphrodite, which will preferentially use male sperm (both types of sperm are stored in the spermatheca).

The sperm of C. elegans is amoeboid, lacking flagella and acrosomes. [27] When self-inseminated, the wild-type worm lays about 300 eggs. When inseminated by a male, the number of progeny can exceed 1,000. Hermaphrodites do not typically mate with other hermaphrodites. At 20 °C, the laboratory strain of C. elegans (N2) has an average lifespan around 2–3 weeks and a generation time of 3 to 4 days.
C. elegans has five pairs of autosomes and one pair of sex chromosomes. Sex in C. elegans is based on an X0 sex-determination system. Hermaphrodites of C. elegans have a matched pair of sex chromosomes (XX); the rare males have only one sex chromosome (X0).
Sex determination
C. elegans are mostly hermaphroditic organisms, producing both sperms and oocytes. [28] Males do occur in the population in a rate of approximately 1 in 200 hermaphrodites, but the two sexes are highly differentiated. [29] [30] Males differ from their hermaphroditic counterparts in that they are smaller and can be identified through the shape of their tail. [30] C.elegans reproduce through a process called androdioecy. This means that they can reproduce in two ways: either through self-fertilization in hermaphrodites or through hermaphrodites breeding with males. Males are produced through non-disjunction of the X chromosomes during meiosis. The worms that reproduce through self-fertilization are at risk for high linkage disequilibrium, which leads to lower genetic diversity in populations and an increase in accumulation of deleterious alleles. [31] In C. elegans, somatic sex determination is attributed to the tra-1 gene. [32] The tra-1 is a gene within the TRA-1 transcription factor sex determination pathway that is regulated post-transcriptionally and works by promoting female development. [32] In hermaphrodites (XX), there are high levels of tra-1 activity, which produces the female reproductive system and inhibits male development. [28] [32] At a certain time in their life cycle, one day before adulthood, hermaphrodites can be identified through the addition of a vulva near their tail. In males (XO), there are low levels of tra-1 activity, resulting in a male reproductive system. [32] Recent research has shown that there are three other genes, fem-1, fem-2, and fem-3, that negatively regulate the TRA-1 pathway and act as the final determiner of sex in C. elegans. [28]
Evolution
The sex determination system in C. elegans is a topic that has been of interest to scientists for years. [29] Since they are used as a model organism, any information discovered about the way their sex determination system might have evolved could further the same evolutionary biology research in other organisms. After almost 30 years of research, scientists have begun to put together the pieces in the evolution of such a system. [29] What they have discovered is that there is a complex pathway involved that has several layers of regulation. [29] The closely related organism Caenorhabditis briggsae has been studied extensively and its whole genome sequence has helped put together the missing pieces in the evolution of C. elegans sex determination. [29] It has been discovered that two genes have assimilated, leading to the proteins XOL-1 and MIX-1 having an effect on sex determination in C. elegans as well. [29] Mutations in the XOL-1 pathway leads to feminization in C. elegans . [33] The mix-1 gene is known to hypoactivate the X chromosome and regulates the morphology of the male tail in C. elegans. [34] Looking at the nematode as a whole, the male and hermaphrodite sex likely evolved from parallel evolution. [29] Parallel evolution is defined as similar traits evolving from an ancestor in similar conditions; simply put, the two species evolve in similar ways over time. An example of this would be marsupial and placental mammals. Scientists have also hypothesized that hermaphrodite asexual reproduction, or "selfing", could have evolved convergently by studying species similar to C. elegans [29] Other studies on the sex determination evolution suggest that genes involving sperm evolve at the faster rate than female genes. [35] However, sperm genes on the X chromosome have reduced evolution rates. Sperm genes have short coding sequences, high codon bias, and disproportionate representation among orphan genes. [35] These characteristics of sperm genes may be the reason for their high rates of evolution and may also suggest how sperm genes evolved out of hermaphrodite worms. Overall, scientists have a general idea of the sex determination pathway in C. elegans, however, the evolution of how this pathway came to be is not yet well defined.
Development

Embryonic development
The fertilized zygote undergoes rotational holoblastic cleavage.
Sperm entry into the oocyte commences formation of an anterior-posterior axis. [36] The sperm microtubule organizing center directs the movement of the sperm pronucleus to the future posterior pole of the embryo, while also inciting the movement of PAR proteins, a group of cytoplasmic determination factors, to their proper respective locations. [37] As a result of the difference in PAR protein distribution, the first cell division is highly asymmetric. [38] C. elegans embryogenesis is among the best understood examples of asymmetric cell division. [39]
All cells of the germline arise from a single primordial germ cell, called the P4 cell, established early in embryogenesis. [40] [41] This primordial cell divides to generate two germline precursors that do not divide further until after hatching. [41]
Axis formation
The resulting daughter cells of the first cell division are called the AB cell (containing PAR-6 and PAR-3) and the P1 cell (containing PAR-1 and PAR-2). A second cell division produces the ABp and ABa cells from the AB cell, and the EMS and P2 cells from the P1 cell. This division establishes the dorsal-ventral axis, with the ABp cell forming the dorsal side and the EMS cell marking the ventral side. [42] Through Wnt signaling, the P2 cell instructs the EMS cell to divide along the anterior-posterior axis. [43] Through Notch signaling, the P2 cell differentially specifies the ABp and ABa cells, which further defines the dorsal-ventral axis. The left-right axis also becomes apparent early in embryogenesis, although it is unclear exactly when specifically the axis is determined. However, most theories of the L-R axis development involve some kind of differences in cells derived from the AB cell. [44]
Gastrulation
Gastrulation occurs after the embryo reaches the 24-cell stage. [45] C. elegans are a species of protostomes, so the blastopore eventually forms the mouth. Involution into the blastopore begins with movement of the endoderm cells and subsequent formation of the gut, followed by the P4 germline precursor, and finally the mesoderm cells, including the cells that eventually form the pharynx. Gastrulation ends when epiboly of the hypoblasts closes the blastopore. [46]
Post-embryonic development


Under environmental conditions favourable for reproduction, hatched larvae develop through four larval stages - L1, L2, L3, and L4 - in just 3 days at 20 °C. When conditions are stressed, as in food insufficiency, excessive population density or high temperature, C. elegans can enter an alternative third larval stage, L2d, called the dauer stage (Dauer is German for permanent). A specific dauer pheromone regulates entry into the dauer state. This pheromone is composed of similar derivatives of the 3,6-dideoxy sugar, ascarylose. Ascarosides, named after the ascarylose base, are involved in many sex-specific and social behaviors. [47] In this way, they constitute a chemical language that C. elegans uses to modulate various phenotypes. Dauer larvae are stress-resistant; they are thin and their mouths are sealed with a characteristic dauer cuticle and cannot take in food. They can remain in this stage for a few months. [48] [49] The stage ends when conditions improve favour further growth of the larva, now moulting into the L4 stage, even though the gonad development is arrested at the L2 stage. [50]
Each stage transition is punctuated by a molt of the worm's transparent cuticle. Transitions through these stages are controlled by genes of the heterochronic pathway, an evolutionarily conserved set of regulatory factors. [51] Many heterochronic genes code for microRNAs, which repress the expression of heterochronic transcription factors and other heterochronic miRNAs. [52] miRNAs were originally discovered in C. elegans. [53] Important developmental events controlled by heterochronic genes include the division and eventual syncitial fusion of the hypodermic seam cells, and their subsequent secretion of the alae in young adults. It is believed that the heterochronic pathway represents an evolutionarily conserved predecessor to circadian clocks. [54]
Some nematodes have a fixed, genetically determined number of cells, a phenomenon known as eutely. The adult C. elegans hermaphrodite has 959 somatic cells and the male has 1033 cells, [55] [56] [57] although it has been suggested that the number of their intestinal cells can increase by one to three in response to gut microbes experienced by mothers. [58] Much of the literature describes the cell number in males as 1031, but the discovery of a pair of left and right MCM neurons increased the number by two in 2015. [57] The number of cells does not change after cell division ceases at the end of the larval period, and subsequent growth is due solely to an increase in the size of individual cells. [59]
Ecology
The different
Caenorhabditis species occupy various nutrient- and bacteria-rich environments. They feed on the bacteria that develop in decaying organic matter (
microbivory). They possess chemosensory receptors which enable the detection of bacteria and bacterial-secreted metabolites (such as iron siderophores), so that they can migrate towards their bacterial prey.
[60] Soil lacks enough organic matter to support self-sustaining populations. C. elegans can survive on a diet of a variety of bacteria, but its wild ecology is largely unknown. Most laboratory strains were taken from artificial environments such as gardens and
compost piles. More recently, C. elegans has been found to thrive in other kinds of organic matter, particularly rotting fruit.
[61]
C. elegans can also ingest pollutants, especially tiny nanoplastics, which could enable the association with antibiotic-resistant bacteria, resulting in the dissemination of nanoplastics and antibiotic-resistant bacteria by C. elegans across the soil.
[62]
C. elegans can also use different species of
yeast, including
Cryptococcus laurentii and
C. kuetzingii, as sole sources of food.
[63] Although a
bacterivore, C. elegans can be killed by a number of pathogenic bacteria, including human pathogens such as
Staphylococcus aureus,
[64]
Pseudomonas aeruginosa,
[65]
Salmonella enterica or
Enterococcus faecalis.
[66] Pathogenic bacteria can also form biofilms, whose sticky exopolymer matrix could impede C. elegans motility
[67] and cloaks bacterial quorum sensing chemoattractants from predator detection.
[68]
Invertebrates such as
millipedes,
insects,
isopods, and
gastropods can transport dauer larvae to various suitable locations. The larvae have also been seen to feed on their hosts when they die.
[69]
Nematodes can survive
desiccation, and in C. elegans, the mechanism for this capability has been demonstrated to be
late embryogenesis abundant proteins.
[70]
C. elegans, as other nematodes, can be eaten by predator nematodes and other omnivores, including some insects.
[71]
The Orsay virus is a virus that affects C. elegans, as well as the Caenorhabditis elegans Cer1 virus [72] and the Caenorhabditis elegans Cer13 virus.
- Interactions with fungi
Wild isolates of Caenorhabditis elegans are regularly found with infections by Microsporidia fungi. One such species, Nematocida parisii, replicates in the intestines of C. elegans. [73]
Arthrobotrys oligospora is the model organism for interactions between fungi and nematodes. [74] It is the most common and widespread nematode capturing fungus.
Use as a model organism
In 1963, Sydney Brenner proposed using C. elegans as a model organism for the investigation primarily of neural development in animals. It is one of the simplest organisms with a nervous system. The neurons do not fire action potentials, and do not express any voltage-gated sodium channels. [75] In the hermaphrodite, this system comprises 302 neurons [76] the pattern of which has been comprehensively mapped, [14] in what is known as a connectome, [77] and shown to be a small-world network. [78]
Research has explored the neural and molecular mechanisms that control several behaviors of C. elegans, including chemotaxis, thermotaxis, mechanotransduction, learning, memory, and mating behaviour. [79] In 2019 the connectome of the male was published using a technique distinct from that used for the hermaphrodite. The same paper used the new technique to redo the hermaphrodite connectome, finding 1,500 new synapses. [80]
It has been used as a model organism to study molecular mechanisms in metabolic diseases. [81] Brenner also chose it as it is easy to grow in bulk populations, and convenient for genetic analysis. [82] It is a multicellular eukaryotic organism, yet simple enough to be studied in great detail. The transparency of C. elegans facilitates the study of cellular differentiation and other developmental processes in the intact organism. The spicules in the male clearly distinguish males from females. Strains are cheap to breed and can be frozen. When subsequently thawed, they remain viable, allowing long-term storage. [13] Maintenance is easy when compared to other multicellular model organisms. A few hundred nematodes can be kept on a single agar plate and suitable growth medium. Brenner described the use of a mutant of E. coli – OP50. OP50 is a uracil-requiring organism and its deficiency in the plate prevents the overgrowth of bacteria which would obscure the worms. [83] The use of OP50 does not demand any major laboratory safety measures, since it is non-pathogenic and easily grown in Luria-Bertani (LB) media overnight. [84]
Cell lineage mapping
The developmental fate of every single somatic cell (959 in the adult hermaphrodite; 1031 in the adult male) has been mapped. [85] [86] These patterns of cell lineage are largely invariant between individuals, whereas in mammals, cell development is more dependent on cellular cues from the embryo.
As mentioned previously, the first cell divisions of early embryogenesis in C. elegans are among the best understood examples of asymmetric cell divisions, and the worm is a very popular model system for studying developmental biology. [39]
Programmed cell death
Programmed cell death ( apoptosis) eliminates many additional cells (131 in the hermaphrodite, most of which would otherwise become neurons); this "apoptotic predictability" has contributed to the elucidation of some apoptotic genes. Cell death-promoting genes and a single cell-death inhibitor have been identified. [87]
RNA interference and gene silencing
RNA interference (RNAi) is a relatively straightforward method of disrupting the function of specific genes. Silencing the function of a gene can sometimes allow a researcher to infer its possible function. The nematode can be soaked in, injected with, [88] or fed with genetically transformed bacteria that express the double-stranded RNA of interest, the sequence of which complements the sequence of the gene that the researcher wishes to disable. [89] RNAi has emerged as a powerful tool in the study of functional genomics. C. elegans has been used to analyse gene functions and claim the promise of future findings in the systematic genetic interactions. [90]
Environmental RNAi uptake is much worse in other species of worms in the genus Caenorhabditis. Although injecting RNA into the body cavity of the animal induces gene silencing in most species, only C. elegans and a few other distantly related nematodes can take up RNA from the bacteria they eat for RNAi. [91] This ability has been mapped down to a single gene, sid-2, which, when inserted as a transgene in other species, allows them to take up RNA for RNAi as C. elegans does. [92]
Cell division and cell cycle
Research into meiosis has been considerably simplified since every germ cell nucleus is at the same given position as it moves down the gonad, so is at the same stage in meiosis. In an early phase of meiosis, the oocytes become extremely resistant to radiation and this resistance depends on expression of genes rad51 and atm that have key roles in recombinational repair. [93] [94] Gene mre-11 also plays a crucial role in recombinational repair of DNA damage during meiosis. [95] A study of the frequency of outcrossing in natural populations showed that selfing is the predominant mode of reproduction in C. elegans, but that infrequent outcrossing events occur at a rate around 1%. [96] Meioses that result in selfing are unlikely to contribute significantly to beneficial genetic variability, but these meioses may provide the adaptive benefit of recombinational repair of DNA damages that arise, especially under stressful conditions. [97]
Drug abuse and addiction
Nicotine dependence can also be studied using C. elegans because it exhibits behavioral responses to nicotine that parallel those of mammals. These responses include acute response, tolerance, withdrawal, and sensitization. [98]
Biological databases
As for most model organisms, scientists that work in the field curate a dedicated online database and the WormBase is that for C. elegans. The WormBase attempts to collate all published information on C. elegans and other related nematodes. Their website has advertised a reward of $4000 for the finder of a new species of closely related nematode. [99] Such a discovery would broaden research opportunities with the worm. [100]
Ageing
C. elegans has been a model organism for research into ageing; for example, the inhibition of an insulin-like growth factor signaling pathway has been shown to increase adult lifespan threefold; [101] [102] while glucose feeding promotes oxidative stress and reduce adult lifespan by a half. [81] Similarly, induced degradation of an insulin/IGF-1 receptor late in life extended life expectancy of worms dramatically. [103] Long-lived mutants of C. elegans were demonstrated to be resistant to oxidative stress and UV light. [104] These long-lived mutants had a higher DNA repair capability than wild-type C. elegans. [104] Knockdown of the nucleotide excision repair gene Xpa-1 increased sensitivity to UV and reduced the life span of the long-lived mutants. These findings indicate that DNA repair capability underlies longevity.
The capacity to repair DNA damage by the process of nucleotide excision repair declines with age. [105]
C. elegans exposed to 5mM lithium chloride (LiCl) showed lengthened life spans. [106] When exposed to 10μM LiCl, reduced mortality was observed, but not with 1μM. [107]
C. elegans has been instrumental in the identification of the functions of genes implicated in Alzheimer's disease, such as presenilin. [108] Moreover, extensive research on C. elegans has identified RNA-binding proteins as essential factors during germline and early embryonic development. [109]
Telomeres, the length of which have been shown to correlate with increased lifespan and delayed onset of senescence in a multitude of organisms, from C. elegans [110] [111] to humans, [112] show an interesting behaviour in C. elegans. While C. elegans maintains its telomeres in a canonical way similar to other eukaryotes, in contrast Drosophila melanogaster is noteworthy in its use of retrotransposons to maintain its telomeres, [113] during knock-out of the catalytic subunit of the telomerase (trt-1) C. elegans can gain the ability of alternative telomere lengthening (ALT). C. elegans was the first eukaryote to gain ALT functionality after knock-out of the canonical telomerase pathway. [114] ALT is also observed in about 10-15% of all clinical cancers. [115] Thus C. elegans is a prime candidate for ALT research. [116] [117] [118] Bayat et al. showed the paradoxical shortening of telomeres during trt-1 over-expression which lead to near sterility while the worms even exhibited a slight increase in lifespan, despite shortened telomeres. [119]
Sleep
C. elegans is notable in animal sleep studies as the most primitive organism to display sleep-like states. In C. elegans, a lethargus phase occurs shortly before each moult. [120] C. elegans has also been demonstrated to sleep after exposure to physical stress, including heat shock, UV radiation, and bacterial toxins. [121]
Sensory biology
While the worm has no eyes, it has been found to be sensitive to light due to a third type of light-sensitive animal photoreceptor protein, LITE-1, which is 10 to 100 times more efficient at absorbing light than the other two types of photopigments ( opsins and cryptochromes) found in the animal kingdom. [122]
C. elegans is remarkably adept at tolerating acceleration. It can withstand 400,000 g's, according to geneticists at the University of São Paulo in Brazil. In an experiment, 96% of them were still alive without adverse effects after an hour in an ultracentrifuge. [123]
Spaceflight research
C. elegans made news when specimens were discovered to have survived the Space Shuttle Columbia disaster in February 2003. [124] Later, in January 2009, live samples of C. elegans from the University of Nottingham were announced to be spending two weeks on the International Space Station that October, in a space research project to explore the effects of zero gravity on muscle development and physiology. The research was primarily about genetic basis of muscle atrophy, which relates to spaceflight or being bed-ridden, geriatric, or diabetic. [125] Descendants of the worms aboard Columbia in 2003 were launched into space on Endeavour for the STS-134 mission. [126] Additional experiments on muscle dystrophy during spaceflight were carried on board the ISS starting in 2018. [127] It was shown that the genes affecting muscles attachment were expressed less in space. However, it has yet to be seen if this affects muscle strength.
Genetics
Genome

Karyotype of C. elegans explanation of colors
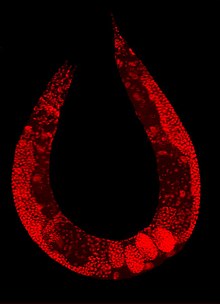
Mitotic chromosomes of C. elegans. DNA (red)/
Kinetochores (green).
Holocentric organisms, including C. elegans, assemble diffuse kinetochores along the entire poleward face of each sister chromatid. | |
| NCBI genome ID | 41 |
|---|---|
| Ploidy | diploid |
| Genome size | 101.169 Mb (haploid) |
| Number of chromosomes | 5 pairs of autosomes (I, II, III, IV and V) + 1 or 2 sex chromosomes (X [128]) |
| Year of completion | 1998 |
| Sequenced organelle | mitochondrion |
| Organelle size | 0,01 Mb |
C. elegans was the first multicellular organism to have its whole genome sequenced. The sequence was published in 1998, [129] although some small gaps were present; the last gap was finished by October 2002.[ citation needed] In the run up to the whole genome the C. elegans Sequencing Consortium/C. elegans Genome Project released several partial scans including Wilson et al. 1994. [130] [131] [132]
Size and gene content
The C. elegans genome is about 100 million base pairs long and consists of six pairs of chromosomes in hermaphrodites or five pairs of autosomes with XO chromosome in male C. elegans and a mitochondrial genome. Its gene density is about one gene per five kilo-base pairs. Introns make up 26% and intergenic regions 47% of the genome. Many genes are arranged in clusters and how many of these are operons is unclear. [133] C. elegans and other nematodes are among the few eukaryotes currently known to have operons; these include trypanosomes, flatworms (notably the trematode Schistosoma mansoni), and a primitive chordate tunicate Oikopleura dioica. Many more organisms are likely to be shown to have these operons. [134]
The genome contains an estimated 20,470 protein-coding genes. [135] About 35% of C. elegans genes have human homologs. Remarkably, human genes have been shown repeatedly to replace their C. elegans homologs when introduced into C. elegans. Conversely, many C. elegans genes can function similarly to mammalian genes. [48]
The number of known RNA genes in the genome has increased greatly due to the 2006 discovery of a new class called 21U-RNA genes, [136] and the genome is now believed to contain more than 16,000 RNA genes, up from as few as 1,300 in 2005. [137]
Scientific curators continue to appraise the set of known genes; new gene models continue to be added and incorrect ones modified or removed.
The reference C. elegans genome sequence continues to change as new evidence reveals errors in the original sequencing. Most changes are minor, adding or removing only a few base pairs of DNA. For example, the WS202 release of WormBase (April 2009) added two base pairs to the genome sequence. [138] Sometimes, more extensive changes are made as noted in the WS197 release of December 2008, which added a region of over 4,300 bp to the sequence. [139] [140]
The C. elegans Genome Project's Wilson et al. 1994 found CelVav [132] and a von Willebrand factor A domain [130] and with Wilson et al. 1998 provides the first credible evidence for an aryl hydrocarbon receptor (AHR) homolog outside of vertebrates. [131] 2
Related genomes
In 2003, the genome sequence of the related nematode C. briggsae was also determined, allowing researchers to study the comparative genomics of these two organisms. [141] The genome sequences of more nematodes from the same genus e.g., C. remanei, [142] C. japonica [143] and C. brenneri (named after Brenner), have also been studied using the shotgun sequencing technique. [144] These sequences have now been completed. [145] [146]
Other genetic studies
As of 2014, C. elegans is the most basal species in the 'Elegans' group (10 species) of the 'Elegans' supergroup (17 species) in phylogenetic studies. It forms a branch of its own distinct to any other species of the group. [147]
Tc1 transposon is a DNA transposon active in C. elegans.
Scientific community
In 2002, the Nobel Prize in Physiology or Medicine was awarded to Sydney Brenner, H. Robert Horvitz, and John Sulston for their work on the genetics of organ development and programmed cell death in C. elegans. The 2006 Nobel Prize in Physiology or Medicine was awarded to Andrew Fire and Craig C. Mello for their discovery of RNA interference in C. elegans. [148] In 2008, Martin Chalfie shared a Nobel Prize in Chemistry for his work on green fluorescent protein; some of the research involved the use of C. elegans.
Many scientists who research C. elegans closely connect to Sydney Brenner, with whom almost all research in this field began in the 1970s; they have worked as either a postdoctoral or a postgraduate researcher in Brenner's lab or in the lab of someone who previously worked with Brenner. Most who worked in his lab later established their own worm research labs, thereby creating a fairly well-documented "lineage" of C. elegans scientists, which was recorded into the WormBase database in some detail at the 2003 International Worm Meeting. [149]
See also
References
- ^ Maupas, É (1900). "Modes et formes de reproduction des nématodes". Archives de Zoologie Expérimentale et Générale. 8: 463–624.
- ^ Nigon V (1949). "Les modalités de la reproduction et le déterminisme du sexe chez quelques nematodes libres". Ann. Sci. Nat. Zool. Biol. Anim. 11: 1–132.
- ^ Moerman DG, Waterston RH (December 1984). "Spontaneous unstable unc-22 IV mutations in C. elegans var. Bergerac". Genetics. 108 (4): 859–77. doi: 10.1093/genetics/108.4.859. PMC 1224270. PMID 6096205.
- ^ Babity JM, Starr TV, Rose AM (June 1990). "Tc1 transposition and mutator activity in a Bristol strain of Caenorhabditis elegans". Molecular & General Genetics. 222 (1): 65–70. doi: 10.1007/bf00283024. PMID 1978238. S2CID 11275388.
- ^ Harris LJ, Rose AM (July 1989). "Structural analysis of Tc1 elements in Caenorhabditis elegans var. Bristol (strain N2)". Plasmid. 22 (1): 10–21. doi: 10.1016/0147-619x(89)90031-0. PMID 2550981.
- ^ "Caenorhabditis". Merriam-Webster.com Dictionary.
- ^ Wood, WB (1988). The Nematode Caenorhabditis elegans. Cold Spring Harbor Laboratory Press. p. 1. ISBN 978-0-87969-433-3.
- ^ Sudhaus W, Kiontke K (2009). "Phylogeny of Rhabditis subgenus Caenorhabditis (Rhabditidae, Nematoda)". Journal of Zoological Systematics and Evolutionary Research. 34 (4): 217–233. doi: 10.1111/j.1439-0469.1996.tb00827.x.
- ^ καινός (caenos) = new, recent; ῥάβδος (rhabdos) = rod, wand.
- ^ Ferris, H (30 November 2013). "Caenorhabditis elegans". University of California, Davis. Archived from the original on 9 December 2013. Retrieved 2013-11-19.
- ^ Wallace RL, Ricci C, Melone G (1996). "A cladistic analysis of pseudocoelomate (aschelminth) morphology". Invertebrate Biology. 115 (2): 104–112. doi: 10.2307/3227041. JSTOR 3227041.
- ^ "Introduction to sex determination". www.wormbook.org. Retrieved 2017-03-15.
- ^ a b Brenner S (May 1974). "The genetics of Caenorhabditis elegans". Genetics. 77 (1): 71–94. doi: 10.1093/genetics/77.1.71. PMC 1213120. PMID 4366476.
- ^ a b Cook, SJ; Jarrell, TA; Brittin, CA; Wang, Y; Bloniarz, AE; Yakovlev, MA; Nguyen, KCQ; Tang, Lt-H; Bayer, EA; Duerr, JS; Bulow, HE; Hobert, O; Hall, DH; Emmons, SW (3 December 2019). "Whole-animal connectomes of both Caenorhabditis elegans sexes". Nature. 571 (7763). US National Library of Medicine, National Institutes of Health: 63–71. Bibcode: 2019Natur.571...63C. doi: 10.1038/s41586-019-1352-7. PMC 6889226. PMID 31270481.
- ^ White JG, Southgate E, Thomson JN, Brenner S (November 1986). "The structure of the nervous system of the nematode Caenorhabditis elegans". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 314 (1165): 1–340. Bibcode: 1986RSPTB.314....1W. doi: 10.1098/rstb.1986.0056. PMID 22462104.
- ^ White JG (June 2013). "Getting into the mind of a worm--a personal view". WormBook: 1–10. doi: 10.1895/wormbook.1.158.1. PMC 4781474. PMID 23801597.
- ^ Jabr F (2012-10-02). "The Connectome Debate: Is Mapping the Mind of a Worm Worth It?". Scientific American. Retrieved 2014-01-18.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2007). Molecular Biology of the Cell (5th ed.). Garland Science. p. 1321. ISBN 978-0-8153-4105-5.
- ^ Lemieux GA, Ashrafi K (August 2016). "Investigating Connections between Metabolism, Longevity, and Behavior in Caenorhabditis elegans". Trends in Endocrinology and Metabolism. 27 (8): 586–596. doi: 10.1016/j.tem.2016.05.004. PMC 4958586. PMID 27289335.
- ^ "The C. elegans pharynx: a model for organogenesis". www.wormbook.org. Retrieved 2017-03-15.
- ^ Soares FA, Fagundez DA, Avila DS (2017). "Neurodegeneration Induced by Metals in Caenorhabditis elegans". Neurotoxicity of Metals. Advances in Neurobiology. Vol. 18. pp. 55–383. doi: 10.1007/978-3-319-60189-2_18. ISBN 978-3-319-60188-5. PMID 28889277.
- ^ Nonet, M. (2004) About the nematode Caenorhabdtis elegans
- ^ Hobert, Oliver (2005). "Specification of the nervous system". WormBook: 1–19. doi: 10.1895/wormbook.1.12.1. PMC 4781215. PMID 18050401.
- ^ Coburn C, Gems D (2013). "The mysterious case of the C. elegans gut granule: death fluorescence, anthranilic acid and the kynurenine pathway". Frontiers in Genetics. 4: 151. doi: 10.3389/fgene.2013.00151. PMC 3735983. PMID 23967012.
- ^ Nayak S, Goree J, Schedl T (January 2005). "fog-2 and the evolution of self-fertile hermaphroditism in Caenorhabditis". PLOS Biology. 3 (1): e6. doi: 10.1371/journal.pbio.0030006. PMC 539060. PMID 15630478.
- ^ Loer CM, Kenyon CJ (December 1993). "Serotonin-deficient mutants and male mating behavior in the nematode Caenorhabditis elegans". The Journal of Neuroscience. 13 (12): 5407–17. doi: 10.1523/jneurosci.13-12-05407.1993. PMC 6576401. PMID 8254383.
- ^ Ma X, Zhao Y, Sun W, Shimabukuro K, Miao L (October 2012). "Transformation: how do nematode sperm become activated and crawl?". Protein & Cell. 3 (10): 755–61. doi: 10.1007/s13238-012-2936-2. PMC 4875351. PMID 22903434.
- ^ a b c Starostina NG, Lim JM, Schvarzstein M, Wells L, Spence AM, Kipreos ET (July 2007). "A CUL-2 ubiquitin ligase containing three FEM proteins degrades TRA-1 to regulate C. elegans sex determination". Developmental Cell. 13 (1): 127–39. doi: 10.1016/j.devcel.2007.05.008. PMC 2064902. PMID 17609115.
- ^ a b c d e f g h Haag ES (December 2005). "The evolution of nematode sex determination: C. elegans as a reference point for comparative biology". WormBook: The Online Review of C. Elegans Biology: 1–14. doi: 10.1895/wormbook.1.120.1. PMC 4781019. PMID 18050417.
- ^ a b "Handbook - Male Introduction". www.wormatlas.org. Retrieved 2021-03-30.
- ^ Frézal L, Félix MA (March 2015). "C. elegans outside the Petri dish". eLife. 4: e05849. doi: 10.7554/eLife.05849. PMC 4373675. PMID 25822066.
- ^ a b c d Conradt, Barbara; Horvitz, H.Robert (1999-08-06). "The TRA-1A Sex Determination Protein of C. elegans Regulates Sexually Dimorphic Cell Deaths by Repressing the egl-1 Cell Death Activator Gene". Cell. 98 (3): 317–327. doi: 10.1016/S0092-8674(00)81961-3. ISSN 0092-8674. PMID 10458607. S2CID 14951719.
- ^ Miller LM, Plenefisch JD, Casson LP, Meyer BJ (October 1988). "xol-1: a gene that controls the male modes of both sex determination and X chromosome dosage compensation in C. elegans". Cell. 55 (1): 167–83. doi: 10.1016/0092-8674(88)90019-0. PMID 3167975. S2CID 5005906.
- ^ "mix-1 (gene) - WormBase : Nematode Information Resource". wormbase.org. Retrieved 2021-04-23.
- ^ a b Cutter AD, Ward S (January 2005). "Sexual and temporal dynamics of molecular evolution in C. elegans development". Molecular Biology and Evolution. 22 (1): 178–88. doi: 10.1093/molbev/msh267. PMID 15371532.
- ^ Goldstein B, Hird SN (May 1996). "Specification of the anteroposterior axis in Caenorhabditis elegans". Development. 122 (5): 1467–74. doi: 10.1242/dev.122.5.1467. PMID 8625834.
- ^ Gilbert SF (2016). Developmental biology (11th ed.). Sinauer. p. 268. ISBN 9781605354705.
- ^ Guo S, Kemphues KJ (May 1995). "par-1, a gene required for establishing polarity in C. elegans embryos, encodes a putative Ser/Thr kinase that is asymmetrically distributed". Cell. 81 (4): 611–20. doi: 10.1016/0092-8674(95)90082-9. PMID 7758115.
- ^ a b Gönczy P, Rose LS (October 2005). "Asymmetric cell division and axis formation in the embryo". WormBook: 1–20. doi: 10.1895/wormbook.1.30.1. PMC 4780927. PMID 18050411.
- ^ Kimble J, Crittenden SL. Germline proliferation and its control. 2005 Aug 15. In: WormBook: The Online Review of C. elegans Biology [Internet]. Pasadena (CA): WormBook; 2005-. Available from: https://www.ncbi.nlm.nih.gov/books/NBK19769/
- ^ a b "WBbt:0006773 (anatomy term)". WormBase (WS242 ed.). May 14, 2014. WBbt:0006773.
- ^ Gilbert SF (2016). Developmental biology (11th ed.). Sinauer. p. 272. ISBN 9781605354705.
- ^ Thorpe CJ, Schlesinger A, Carter JC, Bowerman B (August 1997). "Wnt signaling polarizes an early C. elegans blastomere to distinguish endoderm from mesoderm". Cell. 90 (4): 695–705. doi: 10.1016/s0092-8674(00)80530-9. PMID 9288749.
- ^ Pohl C, Bao Z (September 2010). "Chiral forces organize left-right patterning in C. elegans by uncoupling midline and anteroposterior axis". Developmental Cell. 19 (3): 402–12. doi: 10.1016/j.devcel.2010.08.014. PMC 2952354. PMID 20833362. Villares JC, Carlini EA (1988). "[Quantification of sebaceous excretion in volunteers: influence of chronological age, sex and race]". Med Cutan Ibero Lat Am. 16 (6): 439–44. PMID 3073266. Gilbert SF (2016). Developmental biology (11th ed.). Sinauer. p. 269. ISBN 9781605354705.
- ^ Skiba F, Schierenberg E (June 1992). "Cell lineages, developmental timing, and spatial pattern formation in embryos of free-living soil nematodes". Developmental Biology. 151 (2): 597–610. doi: 10.1016/0012-1606(92)90197-o. PMID 1601187.
- ^ Gilbert SF (2016). Developmental biology (11th ed.). Sinauer. p. 273. ISBN 9781605354705.
- ^ Ludewig, Andreas H.; Schroeder, Frank C. (2013-01-18). "Ascaroside signaling in C. elegans". WormBook: 1–22. doi: 10.1895/wormbook.1.155.1. ISSN 1551-8507. PMC 3758900. PMID 23355522.
- ^ a b "Introduction to C. Elegans". C. Elegans as a model organism. Rutgers University. Archived from the original on 2002-08-18. Retrieved August 15, 2014.
- ^ "Handbook - Introduction".
- ^ "Dauer". www.wormbook.org. Retrieved 2018-09-27.
- ^ Resnick TD, McCulloch KA, Rougvie AE (May 2010). "miRNAs give worms the time of their lives: small RNAs and temporal control in Caenorhabditis elegans". Developmental Dynamics. 239 (5): 1477–89. doi: 10.1002/dvdy.22260. PMC 4698981. PMID 20232378.
-
^ Rougvie AE, Moss EG (2013). "Developmental Transitions in C. Elegans Larval Stages".
Developmental transitions in C. elegans larval stages. Developmental Timing. Vol. 105. Academic Press. pp. 153–80.
doi:
10.1016/B978-0-12-396968-2.00006-3.
ISBN
9780123969682.
PMID
23962842.
{{ cite book}}:|journal=ignored ( help) - ^ Lee RC, Feinbaum RL, Ambros V (December 1993). "The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14". Cell. 75 (5): 843–54. doi: 10.1016/0092-8674(93)90529-y. PMID 8252621.
- ^ Banerjee D, Kwok A, Lin SY, Slack FJ (February 2005). "Developmental timing in C. elegans is regulated by kin-20 and tim-1, homologs of core circadian clock genes". Developmental Cell. 8 (2): 287–95. doi: 10.1016/j.devcel.2004.12.006. PMID 15691769.
- ^ Sulston, J.E.; Horvitz, H.R. (March 1977). "Post-embryonic cell lineages of the nematode, Caenorhabditis elegans". Developmental Biology. 56 (1): 110–156. doi: 10.1016/0012-1606(77)90158-0. PMID 838129.
- ^ Sulston, J.E.; Schierenberg, E.; White, J.G.; Thomson, J.N. (November 1983). "The embryonic cell lineage of the nematode Caenorhabditis elegans". Developmental Biology. 100 (1): 64–119. doi: 10.1016/0012-1606(83)90201-4. PMID 6684600.
- ^ a b Sammut, Michele; Cook, Steven J.; Nguyen, Ken C. Q.; Felton, Terry; Hall, David H.; Emmons, Scott W.; Poole, Richard J.; Barrios, Arantza (October 2015). "Glia-derived neurons are required for sex-specific learning in C. elegans". Nature. 526 (7573): 385–390. Bibcode: 2015Natur.526..385S. doi: 10.1038/nature15700. ISSN 0028-0836. PMC 4650210. PMID 26469050.
- ^ Ohno, Hayao; Bao, Zhirong (2020-11-14). "Small RNAs couple embryonic developmental programs to gut microbes". bioRxiv. doi: 10.1101/2020.11.13.381830. S2CID 227060212.
- ^ Ruppert EE, Fox RS, Barnes RD (2004). Invertebrate Zoology (7th ed.). Cengage Learning. p. 753. ISBN 978-81-315-0104-7.
- ^ Hu, Minqi; Ma, Yeping; Chua, Song Lin (2024-01-16). "Bacterivorous nematodes decipher microbial iron siderophores as prey cue in predator–prey interactions". Proceedings of the National Academy of Sciences. 121 (3): e2314077121. doi: 10.1073/pnas.2314077121. ISSN 0027-8424. PMC 10801909. PMID 38190542.
- ^ Félix MA, Braendle C (November 2010). "The natural history of Caenorhabditis elegans". Current Biology. 20 (22): R965–9. doi: 10.1016/j.cub.2010.09.050. PMID 21093785.
- ^ Chan, Shepherd Yuen; Liu, Sylvia Yang; Wu, Rongben; Wei, Wei; Fang, James Kar-Hei; Chua, Song Lin (2023-06-02). "Simultaneous Dissemination of Nanoplastics and Antibiotic Resistance by Nematode Couriers". Environmental Science & Technology. 57 (23): 8719–8727. Bibcode: 2023EnST...57.8719C. doi: 10.1021/acs.est.2c07129. ISSN 0013-936X. PMID 37267481. S2CID 259047038.
- ^ Mylonakis E, Ausubel FM, Perfect JR, Heitman J, Calderwood SB (November 2002). "Killing of Caenorhabditis elegans by Cryptococcus neoformans as a model of yeast pathogenesis". Proceedings of the National Academy of Sciences of the United States of America. 99 (24): 15675–80. Bibcode: 2002PNAS...9915675M. doi: 10.1073/pnas.232568599. PMC 137775. PMID 12438649.
- ^ Sifri CD, Begun J, Ausubel FM, Calderwood SB (April 2003). "Caenorhabditis elegans as a model host for Staphylococcus aureus pathogenesis". Infection and Immunity. 71 (4): 2208–17. doi: 10.1128/IAI.71.4.2208-2217.2003. PMC 152095. PMID 12654843.
- ^ Tan MW, Mahajan-Miklos S, Ausubel FM (January 1999). "Killing of Caenorhabditis elegans by Pseudomonas aeruginosa used to model mammalian bacterial pathogenesis". Proceedings of the National Academy of Sciences of the United States of America. 96 (2): 715–20. Bibcode: 1999PNAS...96..715T. doi: 10.1073/pnas.96.2.715. PMC 15202. PMID 9892699.
- ^ Garsin DA, Villanueva JM, Begun J, Kim DH, Sifri CD, Calderwood SB, et al. (June 2003). "Long-lived C. elegans daf-2 mutants are resistant to bacterial pathogens". Science. 300 (5627): 1921. doi: 10.1126/science.1080147. PMID 12817143. S2CID 37703980.
- ^ Chan, Shepherd Yuen; Liu, Sylvia Yang; Seng, Zijing; Chua, Song Lin (January 2021). "Biofilm matrix disrupts nematode motility and predatory behavior". The ISME Journal. 15 (1): 260–269. Bibcode: 2021ISMEJ..15..260C. doi: 10.1038/s41396-020-00779-9. ISSN 1751-7370. PMC 7852553. PMID 32958848.
- ^ Li, Shaoyang; Liu, Sylvia Yang; Chan, Shepherd Yuen; Chua, Song Lin (May 2022). "Biofilm matrix cloaks bacterial quorum sensing chemoattractants from predator detection". The ISME Journal. 16 (5): 1388–1396. Bibcode: 2022ISMEJ..16.1388L. doi: 10.1038/s41396-022-01190-2. ISSN 1751-7370. PMC 9038794. PMID 35034106.
- ^ Kiontke K, Sudhaus W (January 2006). "Ecology of Caenorhabditis species". WormBook: 1–14. doi: 10.1895/wormbook.1.37.1. PMC 4780885. PMID 18050464.
- ^ Gal TZ, Glazer I, Koltai H (November 2004). "An LEA group 3 family member is involved in survival of C. elegans during exposure to stress". FEBS Letters. 577 (1–2): 21–6. doi: 10.1016/j.febslet.2004.09.049. PMID 15527756. S2CID 21960486.
- ^ Elaine R. Ingham Soil biology primer USDA
- ^ Bowen, N. J. (1999). "Genomic Analysis of Caenorhabditis elegans Reveals Ancient Families of Retroviral-like Elements". Genome Research. 9 (10): 924–935. doi: 10.1101/gr.9.10.924. PMID 10523521.
- ^ Cuomo CA, Desjardins CA, Bakowski MA, Goldberg J, Ma AT, Becnel JJ, Didier ES, Fan L, Heiman DI, Levin JZ, Young S, Zeng Q, Troemel ER (December 2012). "Microsporidian genome analysis reveals evolutionary strategies for obligate intracellular growth". Genome Research. 22 (12): 2478–88. doi: 10.1101/gr.142802.112. PMC 3514677. PMID 22813931.
- ^ Niu XM, Zhang KQ (2011). "Arthrobotrys oligospora: A model organism for understanding the interaction between fungi and nematodes". Mycology. 2 (2): 59–78. doi: 10.1080/21501203.2011.562559.
- ^ Clare JJ, Tate SN, Nobbs M, Romanos MA (November 2000). "Voltage-gated sodium channels as therapeutic targets". Drug Discovery Today. 5 (11): 506–520. doi: 10.1016/S1359-6446(00)01570-1. PMID 11084387.
- ^ Kosinski RA, Zaremba M (2007). "Dynamics of the Model of the Caenorhabditis Elegans Neural Network". Acta Physica Polonica B. 38 (6): 2201. Bibcode: 2007AcPPB..38.2201K.
- ^ Brouillette, Monique (21 April 2022). "Mapping the brain to understand the mind". Knowable Magazine | Annual Reviews. doi: 10.1146/knowable-042122-1.
- ^ Watts DJ, Strogatz SH (June 1998). "Collective dynamics of 'small-world' networks". Nature. 393 (6684): 440–2. Bibcode: 1998Natur.393..440W. doi: 10.1038/30918. PMID 9623998. S2CID 4429113.
- ^ Schafer WR (September 2005). "Deciphering the neural and molecular mechanisms of C. elegans behavior". Current Biology. 15 (17): R723–9. doi: 10.1016/j.cub.2005.08.020. PMID 16139205.
- ^ Cook SJ, Jarrell TA, Brittin CA, Wang Y, Bloniarz AE, Yakovlev MA, et al. (July 2019). "Whole-animal connectomes of both Caenorhabditis elegans sexes". Nature. 571 (7763): 63–71. Bibcode: 2019Natur.571...63C. doi: 10.1038/s41586-019-1352-7. PMC 6889226. PMID 31270481.
- ^ a b Alcántar-Fernández J, Navarro RE, Salazar-Martínez AM, Pérez-Andrade ME, Miranda-Ríos J (2018). "Caenorhabditis elegans respond to high-glucose diets through a network of stress-responsive transcription factors". PLOS ONE. 13 (7): e0199888. Bibcode: 2018PLoSO..1399888A. doi: 10.1371/journal.pone.0199888. PMC 6039004. PMID 29990370.
- ^ Avery, L. "Sydney Brenner". Southwestern Medical Center. Archived from the original on August 15, 2011. Alt. URL Archived 2013-12-08 at the Wayback Machine
- ^ Brenner, S (1974). "The genetics of Caenorhabditis elegans". Genetics. 77 (1): 71–94. doi: 10.1093/genetics/77.1.71. PMC 1213120. PMID 4366476.
- ^ "Behavior". www.wormbook.org. Retrieved 2018-09-26.
- ^ Sulston JE, Horvitz HR (March 1977). "Post-embryonic cell lineages of the nematode, Caenorhabditis elegans". Developmental Biology. 56 (1): 110–56. doi: 10.1016/0012-1606(77)90158-0. PMID 838129.
- ^ Kimble J, Hirsh D (June 1979). "The postembryonic cell lineages of the hermaphrodite and male gonads in Caenorhabditis elegans". Developmental Biology. 70 (2): 396–417. doi: 10.1016/0012-1606(79)90035-6. PMID 478167.
- ^ Peden E, Killian DJ, Xue D (August 2008). "Cell death specification in C. elegans". Cell Cycle. 7 (16): 2479–84. doi: 10.4161/cc.7.16.6479. PMC 2651394. PMID 18719375.
- ^ NIDDK, National Institute of Diabetes and Digestive and Kidney Diseases (March 5, 2015). "Injection of C. elegans Gonads". YouTube. Archived from the original on 2021-11-17. Retrieved March 21, 2020.
- ^ Kamath RS, Fraser AG, Dong Y, Poulin G, Durbin R, Gotta M, Kanapin A, Le Bot N, Moreno S, Sohrmann M, Welchman DP, Zipperlen P, Ahringer J (January 2003). "Systematic functional analysis of the Caenorhabditis elegans genome using RNAi". Nature. 421 (6920): 231–7. Bibcode: 2003Natur.421..231K. doi: 10.1038/nature01278. hdl: 10261/63159. PMID 12529635. S2CID 15745225.
- ^ Fortunato A, Fraser AG (2005). "Uncover genetic interactions in Caenorhabditis elegans by RNA interference". Bioscience Reports. 25 (5–6): 299–307. doi: 10.1007/s10540-005-2892-7. PMID 16307378. S2CID 6983519.
- ^ Félix MA (November 2008). "RNA interference in nematodes and the chance that favored Sydney Brenner". Journal of Biology. 7 (9): 34. doi: 10.1186/jbiol97. PMC 2776389. PMID 19014674.
- ^ Winston WM, Sutherlin M, Wright AJ, Feinberg EH, Hunter CP (June 2007). "Caenorhabditis elegans SID-2 is required for environmental RNA interference". Proceedings of the National Academy of Sciences of the United States of America. 104 (25): 10565–70. Bibcode: 2007PNAS..10410565W. doi: 10.1073/pnas.0611282104. PMC 1965553. PMID 17563372.
- ^ Takanami T, Mori A, Takahashi H, Higashitani A (November 2000). "Hyper-resistance of meiotic cells to radiation due to a strong expression of a single recA-like gene in Caenorhabditis elegans". Nucleic Acids Research. 28 (21): 4232–6. doi: 10.1093/nar/28.21.4232. PMC 113154. PMID 11058122.
- ^ Takanami T, Zhang Y, Aoki H, Abe T, Yoshida S, Takahashi H, Horiuchi S, Higashitani A (September 2003). "Efficient repair of DNA damage induced by heavy ion particles in meiotic prophase I nuclei of Caenorhabditis elegans". Journal of Radiation Research. 44 (3): 271–6. Bibcode: 2003JRadR..44..271T. doi: 10.1269/jrr.44.271. PMID 14646232.
- ^ Chin GM, Villeneuve AM (March 2001). "C. elegans mre-11 is required for meiotic recombination and DNA repair but is dispensable for the meiotic G(2) DNA damage checkpoint". Genes & Development. 15 (5): 522–34. doi: 10.1101/gad.864101. PMC 312651. PMID 11238374.
- ^ Barrière A, Félix MA (July 2005). "High local genetic diversity and low outcrossing rate in Caenorhabditis elegans natural populations". Current Biology. 15 (13): 1176–84. arXiv: q-bio/0508003. Bibcode: 2005q.bio.....8003B. doi: 10.1016/j.cub.2005.06.022. PMID 16005289. S2CID 2229622.
- ^ Bernstein H, Bernstein C (July 2010) "Evolutionary Origin of Recombination during Meiosis," BioScience 60(7), 498-505. https://doi.org/10.1525/bio.2010.60.7.5
- ^ Feng Z, Li W, Ward A, Piggott BJ, Larkspur ER, Sternberg PW, Xu XZ (November 2006). "A C. elegans model of nicotine-dependent behavior: regulation by TRP-family channels". Cell. 127 (3): 621–33. doi: 10.1016/j.cell.2006.09.035. PMC 2859215. PMID 17081982.
- ^ "Caenorhabditis isolation guide". WormBase. Archived from the original on November 7, 2007. Retrieved 2007-08-30. Alt. URL Archived 2014-09-05 at the Wayback Machine
- ^ Dolgin, E. (2007). "Slime for a Dime". Science. 317 (5842): 1157b. doi: 10.1126/science.317.5842.1157b. S2CID 85163742.
- ^ Wolkow CA, Kimura KD, Lee MS, Ruvkun G (October 2000). "Regulation of C. elegans life-span by insulinlike signaling in the nervous system". Science. 290 (5489): 147–50. Bibcode: 2000Sci...290..147W. doi: 10.1126/science.290.5489.147. PMID 11021802.
- ^ Ewald CY, Landis JN, Porter Abate J, Murphy CT, Blackwell TK (March 2015). "Dauer-independent insulin/IGF-1-signalling implicates collagen remodelling in longevity". Nature. 519 (7541): 97–101. Bibcode: 2015Natur.519...97E. doi: 10.1038/nature14021. PMC 4352135. PMID 25517099.
- ^ Venz, Richard; Pekec, Tina; Katic, Iskra; Ciosk, Rafal; Ewald, Collin Yvès (2021-09-10). Leiser, Scott F; Kaeberlein, Matt; Alcedo, Joy (eds.). "End-of-life targeted degradation of DAF-2 insulin/IGF-1 receptor promotes longevity free from growth-related pathologies". eLife. 10: e71335. doi: 10.7554/eLife.71335. ISSN 2050-084X. PMC 8492056. PMID 34505574.
- ^ a b Hyun, Moonjung; Lee, Jihyun; Lee, Kyungjin; May, Alfred; Bohr, Vilhelm A.; Ahn, Byungchan (2008). "Longevity and resistance to stress correlate with DNA repair capacity in Caenorhabditis elegans". Nucleic Acids Research. 36 (4): 1380–1389. doi: 10.1093/nar/gkm1161. PMC 2275101. PMID 18203746.
- ^ Meyer, Joel N.; Boyd, Windy A.; Azzam, Gregory A.; Haugen, Astrid C.; Freedman, Jonathan H.; Van Houten, Bennett (2007). "Decline of nucleotide excision repair capacity in aging Caenorhabditis elegans". Genome Biology. 8 (5): R70. doi: 10.1186/gb-2007-8-5-r70. PMC 1929140. PMID 17472752.
- ^ McColl G, Killilea DW, Hubbard AE, Vantipalli MC, Melov S, Lithgow GJ (January 2008). "Pharmacogenetic analysis of lithium-induced delayed aging in Caenorhabditis elegans". The Journal of Biological Chemistry. 283 (1): 350–7. doi: 10.1074/jbc.M705028200. PMC 2739662. PMID 17959600.
- ^ Zarse K, Terao T, Tian J, Iwata N, Ishii N, Ristow M (August 2011). "Low-dose lithium uptake promotes longevity in humans and metazoans". European Journal of Nutrition. 50 (5): 387–9. doi: 10.1007/s00394-011-0171-x. PMC 3151375. PMID 21301855.
- ^ Ewald CY, Li C (March 2010). "Understanding the molecular basis of Alzheimer's disease using a Caenorhabditis elegans model system". Brain Structure & Function. 214 (2–3): 263–83. doi: 10.1007/s00429-009-0235-3. PMC 3902020. PMID 20012092.
- ^ Hanazawa M, Yonetani M, Sugimoto A (March 2011). "PGL proteins self associate and bind RNPs to mediate germ granule assembly in C. elegans". The Journal of Cell Biology. 192 (6): 929–37. doi: 10.1083/jcb.201010106. PMC 3063142. PMID 21402787.
- ^ Coutts, Fiona; Palmos, Alish B.; Duarte, Rodrigo R. R.; de Jong, Simone; Lewis, Cathryn M.; Dima, Danai; Powell, Timothy R. (March 2019). "The polygenic nature of telomere length and the anti-ageing properties of lithium". Neuropsychopharmacology. 44 (4): 757–765. doi: 10.1038/s41386-018-0289-0. ISSN 1740-634X. PMC 6372618. PMID 30559463.
- ^ Raices, Marcela; Maruyama, Hugo; Dillin, Andrew; Karlseder, Jan (September 2005). "Uncoupling of longevity and telomere length in C. elegans". PLOS Genetics. 1 (3): e30. doi: 10.1371/journal.pgen.0010030. ISSN 1553-7404. PMC 1200426. PMID 16151516.
- ^ Lulkiewicz, M.; Bajsert, J.; Kopczynski, P.; Barczak, W.; Rubis, B. (September 2020). "Telomere length: how the length makes a difference". Molecular Biology Reports. 47 (9): 7181–7188. doi: 10.1007/s11033-020-05551-y. ISSN 1573-4978. PMC 7561533. PMID 32876842.
- ^ Pardue, Mary-Lou; DeBaryshe, P. G. (2011-12-20). "Retrotransposons that maintain chromosome ends". Proceedings of the National Academy of Sciences of the United States of America. 108 (51): 20317–20324. doi: 10.1073/pnas.1100278108. ISSN 1091-6490. PMC 3251079. PMID 21821789.
- ^ Meier, Bettina; Clejan, Iuval; Liu, Yan; Lowden, Mia; Gartner, Anton; Hodgkin, Jonathan; Ahmed, Shawn (February 2006). "trt-1 is the Caenorhabditis elegans catalytic subunit of telomerase". PLOS Genetics. 2 (2): e18. doi: 10.1371/journal.pgen.0020018. ISSN 1553-7404. PMC 1361356. PMID 16477310.
- ^ Cesare, Anthony J.; Reddel, Roger R. (May 2010). "Alternative lengthening of telomeres: models, mechanisms and implications". Nature Reviews. Genetics. 11 (5): 319–330. doi: 10.1038/nrg2763. ISSN 1471-0064. PMID 20351727. S2CID 19224032.
- ^ Ijomone, Omamuyovwi M.; Miah, Mahfuzur R.; Peres, Tanara V.; Nwoha, Polycarp U.; Aschner, Michael (December 2016). "Null allele mutants of trt-1, the catalytic subunit of telomerase in Caenorhabditis elegans, are less sensitive to Mn-induced toxicity and DAergic degeneration". Neurotoxicology. 57: 54–60. doi: 10.1016/j.neuro.2016.08.016. ISSN 1872-9711. PMID 27593554.
- ^ Shtessel, Ludmila; Lowden, Mia Rochelle; Cheng, Chen; Simon, Matt; Wang, Kyle; Ahmed, Shawn (February 2013). "Caenorhabditis elegans POT-1 and POT-2 repress telomere maintenance pathways". G3: Genes, Genomes, Genetics. 3 (2): 305–313. doi: 10.1534/g3.112.004440. ISSN 2160-1836. PMC 3564990. PMID 23390606.
- ^ Kwon, Mi-Sun; Min, Jaewon; Jeon, Hee-Yeon; Hwang, Kwangwoo; Kim, Chuna; Lee, Junho; Joung, Je-Gun; Park, Woong-Yang; Lee, Hyunsook (October 2016). "Paradoxical delay of senescence upon depletion of BRCA2 in telomerase-deficient worms". FEBS Open Bio. 6 (10): 1016–1024. doi: 10.1002/2211-5463.12109. ISSN 2211-5463. PMC 5055038. PMID 27761361.
- ^ Bayat, Melih; Tanny, Robyn E.; Wang, Ye; Herden, Carla; Daniel, Jens; Andersen, Erik C.; Liebau, Eva; Waschk, Daniel E. J. (2020-03-30). "Effects of telomerase overexpression in the model organism Caenorhabditis elegans". Gene. 732: 144367. doi: 10.1016/j.gene.2020.144367. ISSN 1879-0038. PMID 31954861. S2CID 210829489.
- ^ Iwanir S, Tramm N, Nagy S, Wright C, Ish D, Biron D (March 2013). "The microarchitecture of C. elegans behavior during lethargus: homeostatic bout dynamics, a typical body posture, and regulation by a central neuron". Sleep. 36 (3): 385–95. doi: 10.5665/Sleep.2456. PMC 3571756. PMID 23449971.
- ^ Hill AJ, Mansfield R, Lopez JM, Raizen DM, Van Buskirk C (October 2014). "Cellular stress induces a protective sleep-like state in C. elegans". Current Biology. 24 (20): 2399–405. doi: 10.1016/j.cub.2014.08.040. PMC 4254280. PMID 25264259.
- ^ Teensy, Eyeless Worms Have Completely New Light-Detecting Cells
- ^ Scientific American, August 2018, page 14
- ^ "Worms survived Columbia disaster". BBC News. 1 May 2003. Retrieved 2008-07-11.
- ^ "University sends worms into space". BBC News. 17 January 2009. Retrieved 2009-07-09.
- ^ Klotz, I (16 May 2011). "Legacy Space Worms Flying on Shuttle". Discovery News. Archived from the original on 2012-06-16. Retrieved 2011-05-17.
- ^ Soni, Purushottam; Anupom, Taslim; Lesanpezeshki, Leila; Rahman, Mizanur; Hewitt, Jennifer E.; Vellone, Matthew; Stodieck, Louis; Blawzdziewicz, Jerzy; Szewczyk, Nathaniel J.; Vanapalli, Siva A. (2022-11-07). "Microfluidics-integrated spaceflight hardware for measuring muscle strength of Caenorhabditis elegans on the International Space Station". npj Microgravity. 8 (1): 50. Bibcode: 2022npjMG...8...50S. doi: 10.1038/s41526-022-00241-4. ISSN 2373-8065. PMC 9640571. PMID 36344513.
- ^ Strome S, Kelly WG, Ercan S, Lieb JD (March 2014). "Regulation of the X chromosomes in Caenorhabditis elegans". Cold Spring Harbor Perspectives in Biology. 6 (3): a018366. doi: 10.1101/cshperspect.a018366. PMC 3942922. PMID 24591522.
- ^ The C. elegans Sequencing Consortium (December 1998). "Genome sequence of the nematode C. elegans: a platform for investigating biology". Science. 282 (5396): 2012–8. Bibcode: 1998Sci...282.2012.. doi: 10.1126/science.282.5396.2012. PMID 9851916.
- ^ a b Sadler, J. Evan (1998). "Biochemistry and genetics of von Willebrand factor". Annual Review of Biochemistry. 67 (1). Annual Reviews: 395–424. doi: 10.1146/annurev.biochem.67.1.395. ISSN 0066-4154. PMID 9759493.
- ^ a b Hahn, Mark E. (2002). "Aryl hydrocarbon receptors: diversity and evolution". Chemico-Biological Interactions. 141 (1–2). Elsevier: 131–160. Bibcode: 2002CBI...141..131H. doi: 10.1016/s0009-2797(02)00070-4. ISSN 0009-2797. PMID 12213389.
- ^ a b Bustelo, Xosé R. (2000). "Regulatory and Signaling Properties of the Vav Family". Molecular and Cellular Biology. 20 (5). American Society for Microbiology: 1461–1477. doi: 10.1128/mcb.20.5.1461-1477.2000. ISSN 0270-7306. PMC 85310. PMID 10669724.
- ^ Blumenthal T, Evans D, Link CD, Guffanti A, Lawson D, Thierry-Mieg J, Thierry-Mieg D, Chiu WL, Duke K, Kiraly M, Kim SK (June 2002). "A global analysis of Caenorhabditis elegans operons". Nature. 417 (6891): 851–4. Bibcode: 2002Natur.417..851B. doi: 10.1038/nature00831. PMID 12075352. S2CID 4351788.
- ^ Blumenthal T (November 2004). "Operons in eukaryotes". Briefings in Functional Genomics & Proteomics. 3 (3): 199–211. doi: 10.1093/bfgp/3.3.199. PMID 15642184.
- ^ "WS227 Release Letter". WormBase. 10 August 2011. Archived from the original on 28 November 2013. Retrieved 2013-11-19.
- ^ Ruby JG, Jan C, Player C, Axtell MJ, Lee W, Nusbaum C, Ge H, Bartel DP (December 2006). "Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans". Cell. 127 (6): 1193–207. doi: 10.1016/j.cell.2006.10.040. PMID 17174894.
- ^ Stricklin SL, Griffiths-Jones S, Eddy SR (June 2005). "C. elegans noncoding RNA genes". WormBook: 1–7. doi: 10.1895/wormbook.1.1.1. PMC 4781554. PMID 18023116.
- ^ "WS202 Release Letter". WormBase. 29 May 2009. Archived from the original on December 1, 2013. Retrieved 2013-11-19.
- ^ "WS197 Release Letter". WormBase. 27 November 2008. Archived from the original on 17 October 2019. Retrieved 2013-11-19.
- ^ "Genome sequence changes". WormBase. 15 June 2011. Archived from the original on 17 October 2019. Retrieved 2011-08-13.
- ^ Stein LD, Bao Z, Blasiar D, Blumenthal T, Brent MR, Chen N, Chinwalla A, Clarke L, Clee C, Coghlan A, Coulson A, D'Eustachio P, Fitch DH, Fulton LA, Fulton RE, Griffiths-Jones S, Harris TW, Hillier LW, Kamath R, Kuwabara PE, Mardis ER, Marra MA, Miner TL, Minx P, Mullikin JC, Plumb RW, Rogers J, Schein JE, Sohrmann M, Spieth J, Stajich JE, Wei C, Willey D, Wilson RK, Durbin R, Waterston RH (November 2003). "The genome sequence of Caenorhabditis briggsae: a platform for comparative genomics". PLOS Biology. 1 (2): E45. doi: 10.1371/journal.pbio.0000045. PMC 261899. PMID 14624247.
- ^ Genome Sequencing Center. "Caenorhabditis remanei: Background". Washington University School of Medicine. Archived from the original on 2008-06-16. Retrieved 2008-07-11.
- ^ Genome Sequencing Center. "Caenorhabditis japonica: Background". Washington University School of Medicine. Archived from the original on 2008-06-26. Retrieved 2008-07-11.
- ^ Staden R (June 1979). "A strategy of DNA sequencing employing computer programs". Nucleic Acids Research. 6 (7): 2601–10. doi: 10.1093/nar/6.7.2601. PMC 327874. PMID 461197.
- ^ "UCSC genome browser". Retrieved 8 July 2014.
- ^ Kuhn RM, Karolchik D, Zweig AS, Wang T, Smith KE, Rosenbloom KR, Rhead B, Raney BJ, Pohl A, Pheasant M, Meyer L, Hsu F, Hinrichs AS, Harte RA, Giardine B, Fujita P, Diekhans M, Dreszer T, Clawson H, Barber GP, Haussler D, Kent WJ (January 2009). "The UCSC Genome Browser Database: update 2009". Nucleic Acids Research. 37 (Database issue): D755–61. doi: 10.1093/nar/gkn875. PMC 2686463. PMID 18996895.
- ^ Félix MA, Braendle C, Cutter AD (2014). "A streamlined system for species diagnosis in Caenorhabditis (Nematoda: Rhabditidae) with name designations for 15 distinct biological species". PLOS ONE. 9 (4): e94723. Bibcode: 2014PLoSO...994723F. doi: 10.1371/journal.pone.0094723. PMC 3984244. PMID 24727800.
- ^ Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (February 1998). "Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans". Nature. 391 (6669): 806–11. Bibcode: 1998Natur.391..806F. doi: 10.1038/35888. PMID 9486653. S2CID 4355692.
- ^ Harris TW, Antoshechkin I, Bieri T, Blasiar D, Chan J, Chen WJ, et al. (January 2010). "WormBase: a comprehensive resource for nematode research". Nucleic Acids Research. 38 (Database issue): D463-7. doi: 10.1093/nar/gkp952. PMC 2808986. PMID 19910365.
Further reading
- Bird J, Bird AC (1991). The structure of nematodes. Academic Press. pp. 1, 69–70, 152–153, 165, 224–225. ISBN 978-0-12-099651-3.
- Hope, IA (1999). C. elegans: a practical approach. Oxford University Press. pp. 1–6. ISBN 978-0-19-963738-6.
- Riddle DL, Blumenthal T, Meyer RJ, Priess JR (1997). C. elegans II. Cold Spring Harbor Laboratory Press. pp. 1–4, 679–683. ISBN 978-0-87969-532-3. PMID 21413221.
External links
- Brenner S (2002) Nature's Gift to Science. In. http://nobelprize.org/nobel_prizes/medicine/laureates/2002/brenner-lecture.pdf (also Horvitz and Sulston lectures)
- WormBase – an extensive online database covering the biology and genomics of C. elegans and other nematodes
- WormAtlas – online database on all aspects of C. elegans anatomy with detailed explanations and high-quality images
- WormBook – online review of C. elegans biology
- AceView WormGenes – another genome database for C. elegans, maintained at the NCBI
- C. elegans II – a free online textbook.
- WormWeb Neural Network – an online tool for visualizing and navigating the connectome of C. elegans
- C. elegans movies – a visual introduction to C. elegans
- View the ce11 genome assembly in the UCSC Genome Browser.
- Caenorhabditis elegans at eppo.int ( EPPO code CAEOEL)
- Cylinder, Drew (2022).
"Using C. elegans to study anesthesia".
doi:
10.6084/m9.figshare.20963590.v1.
{{ cite journal}}: Cite journal requires|journal=( help)