| Leucine Rich Repeat | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
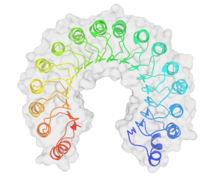 Top view of porcine ribonuclease inhibitor, showing its horseshoe shape.
[1] The outer layer is composed of
α-helices and the inner layer of
parallel β-strands. The inner and outer diameters are roughly 2.1 nm and 6.7 nm, respectively. | |||||||||||
| Identifiers | |||||||||||
| Symbol | LRR_1 | ||||||||||
| Pfam | PF00560 | ||||||||||
| Pfam clan | CL0022 | ||||||||||
| InterPro | IPR003590 | ||||||||||
| SMART | SM00368 | ||||||||||
| SCOP2 | 1bnh / SCOPe / SUPFAM | ||||||||||
| |||||||||||
Ribonuclease inhibitor (RI) is a large (~450 residues, ~49 kDa), acidic (pI ~4.7), leucine-rich repeat protein that forms extremely tight complexes with certain ribonucleases. It is a major cellular protein, comprising ~0.1% of all cellular protein by weight, and appears to play an important role in regulating the lifetime of RNA. [2]
RI has a surprisingly high cysteine content (~6.5%, cf. 1.7% in typical proteins) and is sensitive to oxidation. RI is also rich in leucine (21.5%, compared to 9% in typical proteins) and commensurately lower in other hydrophobic residues, esp. valine, isoleucine, methionine, tyrosine, and phenylalanine.
Structure

RI is the classic leucine-rich repeat protein, consisting of alternating α-helices and β-strands along its backbone. These secondary structure elements wrap around in a curved, right-handed solenoid that resembles a horseshoe. The parallel β-strands and α-helices form the inner and outer wall of the horseshoe, respectively. The structure appears to be stabilized by buried asparagines at the base of each turn, as it passes from α-helix to β-strand. The αβ repeats alternate between 28 and 29 residues in length, effectively forming a 57-residue unit that corresponds to its genetic structure (each exon codes for a 57-residue unit).
Binding to ribonucleases
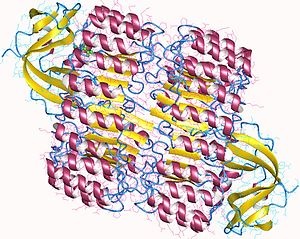
The affinity of RI for ribonucleases is among the highest for any protein-protein interaction; the dissociation constant of the RI- RNase A complex is in the femtomolar (fM) range under physiological conditions. Despite this high affinity, RI is able to bind a wide variety of RNases A despite their relatively low sequence identity. Both biochemical studies and crystallographic structures of RI-RNase A complexes suggest that the interaction is governed largely by electrostatic interactions, but also involves substantial buried surface area. [3] [4] RI's affinity for ribonucleases is important, since many ribonucleases have cytotoxic and cytostatic effects that correlate well with ability to bind RI. [5]
Mammalian RIs are unable to bind certain pancreatic ribonuclease family members from other species. In particular, amphibian RNases, such ranpirnase and amphinase from the Northern leopard frog, escape mammalian RI and have been noted to have differential cytotoxicity against cancer cells. [6]
See also
- Guanidinium thiocyanate - a chemical RNase inhibitor.
References
- ^ a b PDB: 2BNH; Kobe B, Deisenhofer J (1993). "Crystal structure of porcine ribonuclease inhibitor, a protein with leucine-rich repeats". Nature. 366 (6457): 751–6. doi: 10.1038/366751a0. PMID 8264799. S2CID 34579479.
- ^ Shapiro R (2001). "Cytoplasmic ribonuclease inhibitor". Ribonucleases - Part A. Methods in Enzymology. Vol. 341. pp. 611–28. doi: 10.1016/S0076-6879(01)41180-3. ISBN 9780121822422. PMID 11582809.
- ^ Lee FS, Shapiro R, Vallee BL (Jan 1989). "Tight-binding inhibition of angiogenin and ribonuclease A by placental ribonuclease inhibitor". Biochemistry. 28 (1): 225–30. doi: 10.1021/bi00427a031. PMID 2706246.
- ^ Papageorgiou AC, Shapiro R, Acharya KR (Sep 1997). "Molecular recognition of human angiogenin by placental ribonuclease inhibitor--an X-ray crystallographic study at 2.0 A resolution". The EMBO Journal. 16 (17): 5162–77. doi: 10.1093/emboj/16.17.5162. PMC 1170149. PMID 9311977.
- ^ Makarov AA, Ilinskaya ON (April 2003). "Cytotoxic ribonucleases: molecular weapons and their targets". FEBS Letters. 540 (1–3): 15–20. doi: 10.1016/s0014-5793(03)00225-4. PMID 12681476. S2CID 30324366.
- ^ Ardelt W, Shogen K, Darzynkiewicz Z (Jun 2008). "Onconase and amphinase, the antitumor ribonucleases from Rana pipiens oocytes". Current Pharmaceutical Biotechnology. 9 (3): 215–25. doi: 10.2174/138920108784567245. PMC 2586917. PMID 18673287.
Further reading
- Kobe B, Deisenhofer J (Mar 1995). "A structural basis of the interactions between leucine-rich repeats and protein ligands". Nature. 374 (6518): 183–6. doi: 10.1038/374183a0. PMID 7877692. S2CID 4364436.
- Kobe B, Deisenhofer J (Dec 1996). "Mechanism of ribonuclease inhibition by ribonuclease inhibitor protein based on the crystal structure of its complex with ribonuclease A". Journal of Molecular Biology. 264 (5): 1028–43. doi: 10.1006/jmbi.1996.0694. PMID 9000628.
- Papageorgiou AC, Shapiro R, Acharya KR (Sep 1997). "Molecular recognition of human angiogenin by placental ribonuclease inhibitor--an X-ray crystallographic study at 2.0 A resolution". The EMBO Journal. 16 (17): 5162–77. doi: 10.1093/emboj/16.17.5162. PMC 1170149. PMID 9311977.
- Suzuki M, Saxena SK, Boix E, Prill RJ, Vasandani VM, Ladner JE, Sung C, Youle RJ (Mar 1999). "Engineering receptor-mediated cytotoxicity into human ribonucleases by steric blockade of inhibitor interaction". Nature Biotechnology. 17 (3): 265–70. doi: 10.1038/7010. PMID 10096294. S2CID 23140257.
- Shapiro R, Ruiz-Gutierrez M, Chen CZ (Sep 2000). "Analysis of the interactions of human ribonuclease inhibitor with angiogenin and ribonuclease A by mutagenesis: importance of inhibitor residues inside versus outside the C-terminal "hot spot"". Journal of Molecular Biology. 302 (2): 497–519. doi: 10.1006/jmbi.2000.4075. PMID 10970748.
- Bretscher LE, Abel RL, Raines RT (Apr 2000). "A ribonuclease A variant with low catalytic activity but high cytotoxicity". The Journal of Biological Chemistry. 275 (14): 9893–6. doi: 10.1074/jbc.275.14.9893. PMID 10744660.
- Yakovlev GI, Mitkevich VA, Makarov AA (2006). "Ribonuclease inhibitors". Molecular Biology. 40 (6): 867–874. doi: 10.1134/S0026893306060045. S2CID 31887913.