
A toxin-antitoxin system consists of a "toxin" and a corresponding "antitoxin", usually encoded by closely linked genes. The toxin is usually a protein while the antitoxin can be a protein or an RNA. Toxin-antitoxin systems are widely distributed in prokaryotes, and organisms often have them in multiple copies. [2] [3] When these systems are contained on plasmids – transferable genetic elements – they ensure that only the daughter cells that inherit the plasmid survive after cell division. If the plasmid is absent in a daughter cell, the unstable antitoxin is degraded and the stable toxic protein kills the new cell; this is known as 'post-segregational killing' (PSK). [4] [5]
Toxin-antitoxin systems are typically classified according to how the antitoxin neutralises the toxin. In a type I toxin-antitoxin system, the translation of messenger RNA (mRNA) that encodes the toxin is inhibited by the binding of a small non-coding RNA antitoxin that binds the toxin mRNA. The toxic protein in a type II system is inhibited post-translationally by the binding of an antitoxin protein. Type III toxin-antitoxin systems consist of a small RNA that binds directly to the toxin protein and inhibits its activity. [6] There are also types IV-VI, which are less common. [7] Toxin-antitoxin genes are often inherited through horizontal gene transfer [8] [9] and are associated with pathogenic bacteria, having been found on plasmids conferring antibiotic resistance and virulence. [1]
Chromosomal toxin-antitoxin systems also exist, some of which are thought to perform cell functions such as responding to stresses, causing cell cycle arrest and bringing about programmed cell death. [1] [10] In evolutionary terms, toxin-antitoxin systems can be considered selfish DNA in that the purpose of the systems are to replicate, regardless of whether they benefit the host organism or not. Some have proposed adaptive theories to explain the evolution of toxin-antitoxin systems; for example, chromosomal toxin-antitoxin systems could have evolved to prevent the inheritance of large deletions of the host genome. [11] Toxin-antitoxin systems have several biotechnological applications, such as maintaining plasmids in cell lines, targets for antibiotics, and as positive selection vectors. [12]
Biological functions
Stabilization and fitness of mobile DNA
As stated above, toxin-antitoxin systems are well characterized as plasmid addiction modules. It was also proposed that toxin-antitoxin systems have evolved as plasmid exclusion modules. A cell that would carry two plasmids from the same incompatibility group will eventually generate two daughters cells carrying either plasmid. Should one of these plasmids encode for a TA system, its "displacement" by another TA-free plasmid system will prevent its inheritance and thus induce post-segregational killing. [13] This theory was corroborated through computer modelling. [14] Toxin-antitoxin systems can also be found on other mobile genetic elements such as conjugative transposons and temperate bacteriophages and could be implicated in the maintenance and competition of these elements. [15]
Genome stabilization

Toxin-antitoxin systems could prevent harmful large deletions in a bacterial genome, though arguably deletions of large coding regions are fatal to a daughter cell regardless. [11] In Vibrio cholerae, multiple type II toxin-antitoxin systems located in a super-integron were shown to prevent the loss of gene cassettes. [18]
Altruistic cell death
mazEF, a toxin-antitoxin locus found in E. coli and other bacteria, was proposed to induce programmed cell death in response to starvation, specifically a lack of amino acids. [19] This would release the cell's contents for absorption by neighbouring cells, potentially preventing the death of close relatives, and thereby increasing the inclusive fitness of the cell that perished. This would be an example of altruism and how bacterial colonies could resemble multicellular organisms. [14] However, the "mazEF-mediated PCD" has largely been refuted by several studies. [20] [21] [22]
Stress tolerance
Another theory states that chromosomal toxin-antitoxin systems are designed to be bacteriostatic rather than bactericidal. [23] RelE, for example, is a global inhibitor of translation, is induced during nutrient stress. By shutting down translation under stress, it could reduce the chance of starvation by lowering the cell's nutrient requirements. [24] However, it was shown that several toxin-antitoxin systems, including relBE, do not give any competitive advantage under any stress condition. [21]
Anti-addiction
It has been proposed that chromosomal homologues of plasmid toxin-antitoxin systems may serve as anti- addiction modules, which would allow progeny to lose a plasmid without suffering the effects of the toxin it encodes. [9] For example, a chromosomal copy of the ccdA antitoxin encoded in the chromosome of Erwinia chrysanthemi is able to neutralize the ccdB toxin encoded on the F plasmid and thus, prevent toxin activation when such a plasmid is lost. [25] Similarly, the ataR antitoxin encoded on the chromosome of E. coli O157:H7 is able neutralize the ataTP toxin encoded on plasmids found in other enterohemorragic E. coli. [26]
Phage protection
Type III toxin-antitoxin (AbiQ) systems have been shown to protect bacteria from bacteriophages altruistically. [27] [28] During an infection, bacteriophages hijack transcription and translation, which could prevent antitoxin replenishment and release toxin, triggering what is called an "abortive infection". [27] [28] Similar protective effects have been observed with type I, [29] type II, [30] and type IV (AbiE) [31] toxin-antitoxin systems.
Abortive initiation (Abi) can also happen without toxin-antitoxin systems, and many Abi proteins of other types exist. This mechanism serves to halt the replication of phages, protecting the overall population from harm. [32]
Antimicrobial persistence
When bacteria are challenged with antibiotics, a small and distinct subpopulation of cells is able to withstand the treatment by a phenomenon dubbed as "persistence" (not to be confused with resistance). [33] Due to their bacteriostatic properties, type II toxin-antitoxin systems have previously been thought to be responsible for persistence, by switching a fraction of the bacterial population to a dormant state. [34] However, this hypothesis has been widely invalidated. [35] [36] [37]
Selfish DNA
Toxin-antitoxin systems have been used as examples of selfish DNA as part of the gene centered view of evolution. It has been theorised that toxin-antitoxin loci serve only to maintain their own DNA, at the expense of the host organism. [1] [38] Thus, chromosomal toxin-antitoxin systems would serve no purpose and could be treated as "junk DNA". For example, the ccdAB system encoded in the chromosome of E. coli O157:H7 has been shown to be under negative selection, albeit at a slow rate due to its addictive properties. [8]
System types
Type I
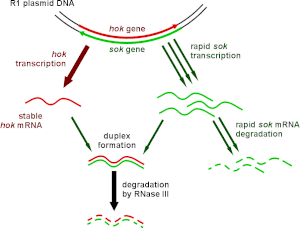
Type I toxin-antitoxin systems rely on the base-pairing of complementary antitoxin RNA with the toxin mRNA. Translation of the mRNA is then inhibited either by degradation via RNase III or by occluding the Shine-Dalgarno sequence or ribosome binding site of the toxin mRNA. Often the toxin and antitoxin are encoded on opposite strands of DNA. The 5' or 3' overlapping region between the two genes is the area involved in complementary base-pairing, usually with between 19–23 contiguous base pairs. [39]
Toxins of type I systems are small, hydrophobic proteins that confer toxicity by damaging cell membranes. [1] Few intracellular targets of type I toxins have been identified, possibly due to the difficult nature of analysing proteins that are poisonous to their bacterial hosts. [10] Also, the detection of small proteins has been challenging due to technical issues, a problem that remains to be solved with large-scale analysis. [40]
Type I systems sometimes include a third component. In the case of the well-characterised hok/sok system, in addition to the hok toxin and sok antitoxin, there is a third gene, called mok. This open reading frame almost entirely overlaps that of the toxin, and the translation of the toxin is dependent on the translation of this third component. [5] Thus the binding of antitoxin to toxin is sometimes a simplification, and the antitoxin in fact binds a third RNA, which then affects toxin translation. [39]
Example systems
| Toxin | Antitoxin | Notes | Ref. |
|---|---|---|---|
| hok | sok | The original and best-understood type I toxin-antitoxin system (pictured), which stabilises plasmids in a number of gram-negative bacteria | [39] |
| fst | RNAII | The first type I system to be identified in gram-positive bacteria | [41] |
| tisB | istR | A chromosomal system induced in the SOS response | [42] |
| dinQ | agrB | A chromosomal system induced in the SOS response | [43] |
| ldrD | rdlD | A chromosomal system in Enterobacteriaceae | [44] |
| flmA | flmB | A hok/sok homologue, which also stabilises the F plasmid | [45] |
| ibs | sib | Discovered in E. coli intergenic regions, the antitoxin was originally named QUAD RNA | [46] |
| txpA/brnT | ratA | Ensures the inheritance of the skin element during sporulation in Bacillus subtilis | [47] |
| symE | symR | A chromosomal system induced in the SOS response | [3] |
| XCV2162 | ptaRNA1 | A system identified in Xanthomonas campestris with erratic phylogenetic distribution. | [48] |
| timP | timR | A chromosomal system identified in Salmonella | [49] |
| aapA1 | isoA1 | A type 1 TA module in Helicobacter pylori | [50] |
| sprA1 | sprA1as | Located within S. aureus small Pathogenicity island (SaPI). SprA1 encodes for a small cytotoxic peptide, PepA1, which disrupts both S. aureus membranes and host erythrocytes. | [51] [52] |
Type II

Type II toxin-antitoxin systems are generally better-understood than type I. [39] In this system a labile proteic antitoxin tightly binds and inhibits the activity of a stable toxin. [10] The largest family of type II toxin-antitoxin systems is vapBC, [53] which has been found through bioinformatics searches to represent between 37 and 42% of all predicted type II loci. [16] [17] Type II systems are organised in operons with the antitoxin protein typically being located upstream of the toxin, which helps to prevent expression of the toxin without the antitoxin. [54] The proteins are typically around 100 amino acids in length, [39] and exhibit toxicity in a number of ways: CcdB, for example, affects DNA replication by poisoning DNA gyrase [55] whereas toxins from the MazF family are endoribonucleases that cleave cellular mRNAs, [56] [57] tRNAs [58] [59] or rRNAs [60] at specific sequence motifs. The most common toxic activity is the protein acting as an endonuclease, also known as an interferase. [61] [62]
One of the key features of the TAs is the autoregulation. The antitoxin and toxin protein complex bind to the operator that is present upstream of the TA genes. This results in repression of the TA operon. The key to the regulation are (i) the differential translation of the TA proteins and (ii) differential proteolysis of the TA proteins. As explained by the "Translation-reponsive model", [63] the degree of expression is inversely proportional to the concentration of the repressive TA complex. The TA complex concentration is directly proportional to the global translation rate. The higher the rate of translation more TA complex and less transcription of TA mRNA. Lower the rate of translation, lesser the TA complex and higher the expression. Hence, the transcriptional expression of TA operon is inversely proportional to translation rate.
A third protein can sometimes be involved in type II toxin-antitoxin systems. in the case of the ω-ε-ζ (omega-epsilon-zeta) system, the omega protein is a DNA binding protein that negatively regulates the transcription of the whole system. [64] Similarly, the paaR2 protein regulates the expression of the paaR2-paaA2-parE2 toxin-antitoxin system. [65] Other toxin-antitoxin systems can be found with a chaperone as a third component. [66] This chaperone is essential for proper folding of the antitoxin, thus making the antitoxin addicted to its cognate chaperone.
Example systems
| Toxin | Antitoxin | Notes | Ref. |
|---|---|---|---|
| ccdB | ccdA | Found on the F plasmid of Escherichia coli | [55] |
| parE | parD | Found in multiple copies in Caulobacter crescentus | [67] |
| mazF | mazE | Found in E. coli and in chromosomes of other bacteria | [29] |
| yafO | yafN | A system induced by the SOS response to DNA damage in E. coli | [68] |
| hicA | hicB | Found in archaea and bacteria | [69] |
| kid | kis | Stabilises the R1 plasmid and is related to the CcdB/A system | [23] |
| ζ | ε | Found mostly in Gram-positive bacteria | [64] |
| ataT | ataR | Found in enterohemorragic E. coli and Klebsiella spp. | [70] |
Type III
| ToxN_toxin | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | ToxN, type III toxin-antitoxin system | ||||||||
| Pfam | PF13958 | ||||||||
| |||||||||
Type III toxin-antitoxin systems rely on direct interaction between a toxic protein and an RNA antitoxin. The toxic effects of the protein are neutralised by the RNA gene. [6] One example is the ToxIN system from the bacterial plant pathogen Erwinia carotovora. The toxic ToxN protein is approximately 170 amino acids long and has been shown to be toxic to E. coli. The toxic activity of ToxN is inhibited by ToxI RNA, an RNA with 5.5 direct repeats of a 36 nucleotide motif (AGGTGATTTGCTACCTTTAAGTGCAGCTAGAAATTC). [27] [71] Crystallographic analysis of ToxIN has found that ToxN inhibition requires the formation of a trimeric ToxIN complex, whereby three ToxI monomers bind three ToxN monomers; the complex is held together by extensive protein-RNA interactions. [72]
Type IV
Type IV toxin-antitoxin systems are similar to type II systems, because they consist of two proteins. Unlike type II systems, the antitoxin in type IV toxin-antitoxin systems counteracts the activity of the toxin, and the two proteins do not necessarily interact directly. DarTG1 and DarTG2 are type IV toxin-antitoxin systems that modify DNA. Their toxins add ADP-ribose to guanosine bases (DarT1 toxin) or thymidine bases (DarT2 toxin), and their antitoxins remove the toxic modifications (NADAR antitoxin from guanosine and DarG antitoxin from thymidine). [73] [74] [75] [76]
Type V
ghoST is a type V toxin-antitoxin system, in which the antitoxin (GhoS) cleaves the ghoT mRNA. This system is regulated by a type II system, mqsRA. [77]
Type VI
socAB is a type VI toxin-antitoxin system that was discovered in Caulobacter crescentus. The antitoxin, SocA, promotes degradation of the toxin, SocB, by the protease ClpXP. [78]
Type VII
Type VII has been proposed to include systems hha/tomB, tglT/takA and hepT/mntA, all of which neutralise toxin activity by post-translational chemical modification of amino acid residues. [79]
Type VIII
Type VIII includes the system creTA. In this system, the antitoxin creA serves as a guide RNA for a CRISPR-Cas system. Due to incomplete complementarity between the creA guide and the creAT promoter, the Cas complex does not cleave the DNA, but instead remains at the site, where it blocks access by RNA polymerase, preventing expression of the creT toxin (a natural instance of CRISPRi). When expressed, the creT RNA will sequester the rare arginine codon tRNAUCU, stalling translation and halting cell metabolism. [80]
Biotechnological applications
The biotechnological applications of toxin-antitoxin systems have begun to be realised by several biotechnology organisations. [12] [23] A primary usage is in maintaining plasmids in a large bacterial cell culture. In an experiment examining the effectiveness of the hok/sok locus, it was found that segregational stability of an inserted plasmid expressing beta-galactosidase was increased by between 8 and 22 times compared to a control culture lacking a toxin-antitoxin system. [81] [82] In large-scale microorganism processes such as fermentation, progeny cells lacking the plasmid insert often have a higher fitness than those who inherit the plasmid and can outcompete the desirable microorganisms. A toxin-antitoxin system maintains the plasmid thereby maintaining the efficiency of the industrial process. [12]
Additionally, toxin-antitoxin systems may be a future target for antibiotics. Inducing suicide modules against pathogens could help combat the growing problem of multi-drug resistance. [83]
Ensuring a plasmid accepts an insert is a common problem of DNA cloning. Toxin-antitoxin systems can be used to positively select for only those cells that have taken up a plasmid containing the inserted gene of interest, screening out those that lack the inserted gene. An example of this application comes from the ccdB-encoded toxin, which has been incorporated into plasmid vectors. [84] The gene of interest is then targeted to recombine into the ccdB locus, inactivating the transcription of the toxic protein. Thus, cells containing the plasmid but not the insert perish due to the toxic effects of CcdB protein, and only those that incorporate the insert survive. [12]
Another example application involves both the CcdB toxin and CcdA antitoxin. CcdB is found in recombinant bacterial genomes and an inactivated version of CcdA is inserted into a linearised plasmid vector. A short extra sequence is added to the gene of interest that activates the antitoxin when the insertion occurs. This method ensures orientation-specific gene insertion. [84]
Genetically modified organisms must be contained in a pre-defined area during research. [83] Toxin-antitoxin systems can cause cell suicide in certain conditions, such as a lack of a lab-specific growth medium they would not encounter outside of the controlled laboratory set-up. [23] [85]
See also
References
- ^ a b c d e Van Melderen L, Saavedra De Bast M (March 2009). Rosenberg SM (ed.). "Bacterial toxin-antitoxin systems: more than selfish entities?". PLOS Genetics. 5 (3): e1000437. doi: 10.1371/journal.pgen.1000437. PMC 2654758. PMID 19325885.
- ^ Fozo EM, Makarova KS, Shabalina SA, Yutin N, Koonin EV, Storz G (June 2010). "Abundance of type I toxin-antitoxin systems in bacteria: searches for new candidates and discovery of novel families". Nucleic Acids Research. 38 (11): 3743–59. doi: 10.1093/nar/gkq054. PMC 2887945. PMID 20156992.
- ^ a b Gerdes K, Wagner EG (April 2007). "RNA antitoxins". Current Opinion in Microbiology. 10 (2): 117–24. doi: 10.1016/j.mib.2007.03.003. PMID 17376733.
- ^ Gerdes K (February 2000). "Toxin-antitoxin modules may regulate synthesis of macromolecules during nutritional stress". Journal of Bacteriology. 182 (3): 561–72. doi: 10.1128/JB.182.3.561-572.2000. PMC 94316. PMID 10633087.
- ^ a b Faridani OR, Nikravesh A, Pandey DP, Gerdes K, Good L (2006). "Competitive inhibition of natural antisense Sok-RNA interactions activates Hok-mediated cell killing in Escherichia coli". Nucleic Acids Research. 34 (20): 5915–22. doi: 10.1093/nar/gkl750. PMC 1635323. PMID 17065468.
- ^ a b Labrie SJ, Samson JE, Moineau S (May 2010). "Bacteriophage resistance mechanisms". Nature Reviews. Microbiology. 8 (5): 317–27. doi: 10.1038/nrmicro2315. PMID 20348932. S2CID 205497795.
- ^ Page R, Peti W (April 2016). "Toxin-antitoxin systems in bacterial growth arrest and persistence". Nature Chemical Biology. 12 (4): 208–14. doi: 10.1038/nchembio.2044. PMID 26991085.
- ^ a b Mine N, Guglielmini J, Wilbaux M, Van Melderen L (April 2009). "The decay of the chromosomally encoded ccdO157 toxin-antitoxin system in the Escherichia coli species". Genetics. 181 (4): 1557–66. doi: 10.1534/genetics.108.095190. PMC 2666520. PMID 19189956.
- ^ a b Ramisetty BC, Santhosh RS (February 2016). "Horizontal gene transfer of chromosomal Type II toxin-antitoxin systems of Escherichia coli". FEMS Microbiology Letters. 363 (3): fnv238. doi: 10.1093/femsle/fnv238. PMID 26667220.
- ^ a b c Hayes F (September 2003). "Toxins-antitoxins: plasmid maintenance, programmed cell death, and cell cycle arrest". Science. 301 (5639): 1496–9. Bibcode: 2003Sci...301.1496H. doi: 10.1126/science.1088157. PMID 12970556. S2CID 10028255.
- ^ a b Rowe-Magnus DA, Guerout AM, Biskri L, Bouige P, Mazel D (March 2003). "Comparative analysis of superintegrons: engineering extensive genetic diversity in the Vibrionaceae". Genome Research. 13 (3): 428–42. doi: 10.1101/gr.617103. PMC 430272. PMID 12618374.
- ^ a b c d Stieber D, Gabant P, Szpirer C (September 2008). "The art of selective killing: plasmid toxin/antitoxin systems and their technological applications". BioTechniques. 45 (3): 344–6. doi: 10.2144/000112955. PMID 18778262.
- ^ Cooper TF, Heinemann JA (November 2000). "Postsegregational killing does not increase plasmid stability but acts to mediate the exclusion of competing plasmids". Proceedings of the National Academy of Sciences of the United States of America. 97 (23): 12643–8. Bibcode: 2000PNAS...9712643C. doi: 10.1073/pnas.220077897. PMC 18817. PMID 11058151.
- ^ a b Mochizuki A, Yahara K, Kobayashi I, Iwasa Y (February 2006). "Genetic addiction: selfish gene's strategy for symbiosis in the genome". Genetics. 172 (2): 1309–23. doi: 10.1534/genetics.105.042895. PMC 1456228. PMID 16299387.
- ^ Magnuson RD (September 2007). "Hypothetical functions of toxin-antitoxin systems". Journal of Bacteriology. 189 (17): 6089–92. doi: 10.1128/JB.00958-07. PMC 1951896. PMID 17616596.
- ^ a b Pandey DP, Gerdes K (2005). "Toxin-antitoxin loci are highly abundant in free-living but lost from host-associated prokaryotes". Nucleic Acids Research. 33 (3): 966–76. doi: 10.1093/nar/gki201. PMC 549392. PMID 15718296.
- ^ a b c Sevin EW, Barloy-Hubler F (2007). "RASTA-Bacteria: a web-based tool for identifying toxin-antitoxin loci in prokaryotes". Genome Biology. 8 (8): R155. doi: 10.1186/gb-2007-8-8-r155. PMC 2374986. PMID 17678530.
- ^ Szekeres S, Dauti M, Wilde C, Mazel D, Rowe-Magnus DA (March 2007). "Chromosomal toxin-antitoxin loci can diminish large-scale genome reductions in the absence of selection". Molecular Microbiology. 63 (6): 1588–605. doi: 10.1111/j.1365-2958.2007.05613.x. PMID 17367382. S2CID 28191383.
- ^ Aizenman E, Engelberg-Kulka H, Glaser G (June 1996). "An Escherichia coli chromosomal "addiction module" regulated by guanosine [corrected] 3',5'-bispyrophosphate: a model for programmed bacterial cell death". Proceedings of the National Academy of Sciences of the United States of America. 93 (12): 6059–63. Bibcode: 1996PNAS...93.6059A. doi: 10.1073/pnas.93.12.6059. PMC 39188. PMID 8650219.
- ^ Ramisetty BC, Natarajan B, Santhosh RS (February 2015). "mazEF-mediated programmed cell death in bacteria: "what is this?"". Critical Reviews in Microbiology. 41 (1): 89–100. doi: 10.3109/1040841X.2013.804030. PMID 23799870. S2CID 34286252.
- ^ a b Tsilibaris V, Maenhaut-Michel G, Mine N, Van Melderen L (September 2007). "What is the benefit to Escherichia coli of having multiple toxin-antitoxin systems in its genome?". Journal of Bacteriology. 189 (17): 6101–8. doi: 10.1128/JB.00527-07. PMC 1951899. PMID 17513477.
- ^ Ramisetty BC, Raj S, Ghosh D (December 2016). "Escherichia coli MazEF toxin-antitoxin system does not mediate programmed cell death". Journal of Basic Microbiology. 56 (12): 1398–1402. doi: 10.1002/jobm.201600247. PMID 27259116. S2CID 1685755.
- ^ a b c d Diago-Navarro E, Hernandez-Arriaga AM, López-Villarejo J, Muñoz-Gómez AJ, Kamphuis MB, Boelens R, Lemonnier M, Díaz-Orejas R (August 2010). "parD toxin-antitoxin system of plasmid R1--basic contributions, biotechnological applications and relationships with closely-related toxin-antitoxin systems". The FEBS Journal. 277 (15): 3097–117. doi: 10.1111/j.1742-4658.2010.07722.x. PMID 20569269.
- ^ Christensen SK, Mikkelsen M, Pedersen K, Gerdes K (December 2001). "RelE, a global inhibitor of translation, is activated during nutritional stress". Proceedings of the National Academy of Sciences of the United States of America. 98 (25): 14328–33. Bibcode: 2001PNAS...9814328C. doi: 10.1073/pnas.251327898. PMC 64681. PMID 11717402.
- ^ Saavedra De Bast M, Mine N, Van Melderen L (July 2008). "Chromosomal toxin-antitoxin systems may act as antiaddiction modules". Journal of Bacteriology. 190 (13): 4603–9. doi: 10.1128/JB.00357-08. PMC 2446810. PMID 18441063.
- ^ Jurėnas D, Garcia-Pino A, Van Melderen L (September 2017). "Novel toxins from type II toxin-antitoxin systems with acetyltransferase activity". Plasmid. 93: 30–35. doi: 10.1016/j.plasmid.2017.08.005. PMID 28941941.
- ^ a b c Fineran PC, Blower TR, Foulds IJ, Humphreys DP, Lilley KS, Salmond GP (January 2009). "The phage abortive infection system, ToxIN, functions as a protein-RNA toxin-antitoxin pair". Proceedings of the National Academy of Sciences of the United States of America. 106 (3): 894–9. Bibcode: 2009PNAS..106..894F. doi: 10.1073/pnas.0808832106. PMC 2630095. PMID 19124776.
- ^ a b Emond E, Dion E, Walker SA, Vedamuthu ER, Kondo JK, Moineau S (December 1998). "AbiQ, an abortive infection mechanism from Lactococcus lactis". Applied and Environmental Microbiology. 64 (12): 4748–56. Bibcode: 1998ApEnM..64.4748E. doi: 10.1128/AEM.64.12.4748-4756.1998. PMC 90918. PMID 9835558.
- ^ a b Hazan R, Engelberg-Kulka H (September 2004). "Escherichia coli mazEF-mediated cell death as a defense mechanism that inhibits the spread of phage P1". Molecular Genetics and Genomics. 272 (2): 227–34. doi: 10.1007/s00438-004-1048-y. PMID 15316771. S2CID 28840747.
- ^ Pecota DC, Wood TK (April 1996). "Exclusion of T4 phage by the hok/sok killer locus from plasmid R1". Journal of Bacteriology. 178 (7): 2044–50. doi: 10.1128/jb.178.7.2044-2050.1996. PMC 177903. PMID 8606182.
- ^ Dy RL, Przybilski R, Semeijn K, Salmond GP, Fineran PC (April 2014). "A widespread bacteriophage abortive infection system functions through a Type IV toxin-antitoxin mechanism". Nucleic Acids Research. 42 (7): 4590–605. doi: 10.1093/nar/gkt1419. PMC 3985639. PMID 24465005.
- ^ Seed KD (June 2015). "Battling Phages: How Bacteria Defend against Viral Attack". PLOS Pathogens. 11 (6): e1004847. doi: 10.1371/journal.ppat.1004847. PMC 4465916. PMID 26066799.
- ^ Kussell E, Kishony R, Balaban NQ, Leibler S (April 2005). "Bacterial persistence: a model of survival in changing environments". Genetics. 169 (4): 1807–14. doi: 10.1534/genetics.104.035352. PMC 1449587. PMID 15687275.
- ^ Maisonneuve E, Gerdes K (April 2014). "Molecular mechanisms underlying bacterial persisters". Cell. 157 (3): 539–48. doi: 10.1016/j.cell.2014.02.050. PMID 24766804.
- ^ Ramisetty BC, Ghosh D, Roy Chowdhury M, Santhosh RS (2016). "What Is the Link between Stringent Response, Endoribonuclease Encoding Type II Toxin-Antitoxin Systems and Persistence?". Frontiers in Microbiology. 7: 1882. doi: 10.3389/fmicb.2016.01882. PMC 5120126. PMID 27933045.
- ^ Harms A, Fino C, Sørensen MA, Semsey S, Gerdes K (December 2017). "Prophages and Growth Dynamics Confound Experimental Results with Antibiotic-Tolerant Persister Cells". mBio. 8 (6): e01964–17. doi: 10.1128/mBio.01964-17. PMC 5727415. PMID 29233898.
- ^ Goormaghtigh F, Fraikin N, Putrinš M, Hallaert T, Hauryliuk V, Garcia-Pino A, Sjödin A, Kasvandik S, Udekwu K, Tenson T, Kaldalu N, Van Melderen L (June 2018). "Reassessing the Role of Type II Toxin-Antitoxin Systems in Formation of Escherichia coli Type II Persister Cells". mBio. 9 (3): e00640–18. doi: 10.1128/mBio.00640-18. PMC 6016239. PMID 29895634.
- ^ Ramisetty BC, Santhosh RS (July 2017). "Endoribonuclease type II toxin-antitoxin systems: functional or selfish?". Microbiology. 163 (7): 931–939. doi: 10.1099/mic.0.000487. PMID 28691660. S2CID 3879598.
- ^ a b c d e Fozo EM, Hemm MR, Storz G (December 2008). "Small toxic proteins and the antisense RNAs that repress them". Microbiology and Molecular Biology Reviews. 72 (4): 579–89, Table of Contents. doi: 10.1128/MMBR.00025-08. PMC 2593563. PMID 19052321.
- ^ Sberro H, Fremin BJ, Zlitni S, Edfors F, Greenfield N, Snyder MP, et al. (August 2019). "Large-Scale Analyses of Human Microbiomes Reveal Thousands of Small, Novel Genes". Cell. 178 (5): 1245–1259.e14. doi: 10.1016/j.cell.2019.07.016. PMC 6764417. PMID 31402174.
- ^ Greenfield TJ, Ehli E, Kirshenmann T, Franch T, Gerdes K, Weaver KE (August 2000). "The antisense RNA of the par locus of pAD1 regulates the expression of a 33-amino-acid toxic peptide by an unusual mechanism". Molecular Microbiology. 37 (3): 652–60. doi: 10.1046/j.1365-2958.2000.02035.x. PMID 10931358. (subscription required)
- ^ Vogel J, Argaman L, Wagner EG, Altuvia S (December 2004). "The small RNA IstR inhibits synthesis of an SOS-induced toxic peptide". Current Biology. 14 (24): 2271–6. doi: 10.1016/j.cub.2004.12.003. PMID 15620655.
- ^ Weel-Sneve R, Kristiansen KI, Odsbu I, Dalhus B, Booth J, Rognes T, Skarstad K, Bjørås M (February 7, 2013). "Single transmembrane peptide DinQ modulates membrane-dependent activities". PLOS Genetics. 9 (2): e1003260. doi: 10.1371/journal.pgen.1003260. PMC 3567139. PMID 23408903.
- ^ Kawano M, Oshima T, Kasai H, Mori H (July 2002). "Molecular characterization of long direct repeat (LDR) sequences expressing a stable mRNA encoding for a 35-amino-acid cell-killing peptide and a cis-encoded small antisense RNA in Escherichia coli". Molecular Microbiology. 45 (2): 333–49. doi: 10.1046/j.1365-2958.2002.03042.x. PMID 12123448. (subscription required)
- ^ Loh SM, Cram DS, Skurray RA (June 1988). "Nucleotide sequence and transcriptional analysis of a third function (Flm) involved in F-plasmid maintenance". Gene. 66 (2): 259–68. doi: 10.1016/0378-1119(88)90362-9. PMID 3049248.
- ^ Fozo EM, Kawano M, Fontaine F, Kaya Y, Mendieta KS, Jones KL, Ocampo A, Rudd KE, Storz G (December 2008). "Repression of small toxic protein synthesis by the Sib and OhsC small RNAs". Molecular Microbiology. 70 (5): 1076–93. doi: 10.1111/j.1365-2958.2008.06394.x. PMC 2597788. PMID 18710431. (subscription required)
- ^ Silvaggi JM, Perkins JB, Losick R (October 2005). "Small untranslated RNA antitoxin in Bacillus subtilis". Journal of Bacteriology. 187 (19): 6641–50. doi: 10.1128/JB.187.19.6641-6650.2005. PMC 1251590. PMID 16166525.
- ^ Findeiss S, Schmidtke C, Stadler PF, Bonas U (March 2010). "A novel family of plasmid-transferred anti-sense ncRNAs". RNA Biology. 7 (2): 120–4. doi: 10.4161/rna.7.2.11184. PMID 20220307.
- ^ Andresen L, Martínez-Burgo Y, Nilsson Zangelin J, Rizvanovic A, Holmqvist E (November 2020). "Salmonella Protein TimP Targets the Cytoplasmic Membrane and Is Repressed by the Small RNA TimR". mBio. 11 (6): e01659–20, /mbio/11/6/mBio.01659–20.atom. doi: 10.1128/mBio.01659-20. PMC 7667032. PMID 33172998.
- ^ Arnion H, Korkut DN, Masachis Gelo S, Chabas S, Reignier J, Iost I, Darfeuille F (May 2017). "Mechanistic insights into type I toxin antitoxin systems in Helicobacter pylori: the importance of mRNA folding in controlling toxin expression". Nucleic Acids Research. 45 (8): 4782–4795. doi: 10.1093/nar/gkw1343. PMC 5416894. PMID 28077560.
- ^ Sayed N, Jousselin A, Felden B (December 2011). "A cis-antisense RNA acts in trans in Staphylococcus aureus to control translation of a human cytolytic peptide". Nature Structural & Molecular Biology. 19 (1): 105–12. doi: 10.1038/nsmb.2193. PMID 22198463. S2CID 8217681.
- ^ Sayed N, Nonin-Lecomte S, Réty S, Felden B (December 2012). "Functional and structural insights of a Staphylococcus aureus apoptotic-like membrane peptide from a toxin-antitoxin module". The Journal of Biological Chemistry. 287 (52): 43454–63. doi: 10.1074/jbc.M112.402693. PMC 3527932. PMID 23129767.
- ^ Robson J, McKenzie JL, Cursons R, Cook GM, Arcus VL (July 2009). "The vapBC operon from Mycobacterium smegmatis is an autoregulated toxin-antitoxin module that controls growth via inhibition of translation". Journal of Molecular Biology. 390 (3): 353–67. doi: 10.1016/j.jmb.2009.05.006. PMID 19445953.
- ^ Deter HS, Jensen RV, Mather WH, Butzin NC (July 2017). "Mechanisms for Differential Protein Production in Toxin-Antitoxin Systems". Toxins. 9 (7): 211. doi: 10.3390/toxins9070211. PMC 5535158. PMID 28677629.
- ^ a b Bernard P, Couturier M (August 1992). "Cell killing by the F plasmid CcdB protein involves poisoning of DNA-topoisomerase II complexes". Journal of Molecular Biology. 226 (3): 735–45. doi: 10.1016/0022-2836(92)90629-X. PMID 1324324.
- ^ Zhang Y, Zhang J, Hoeflich KP, Ikura M, Qing G, Inouye M (October 2003). "MazF cleaves cellular mRNAs specifically at ACA to block protein synthesis in Escherichia coli". Molecular Cell. 12 (4): 913–23. doi: 10.1016/s1097-2765(03)00402-7. PMID 14580342.
- ^ Culviner PH, Laub MT (June 2018). "Global Analysis of the E. coli Toxin MazF Reveals Widespread Cleavage of mRNA and the Inhibition of rRNA Maturation and Ribosome Biogenesis". Molecular Cell. 70 (5): 868–880.e10. doi: 10.1016/j.molcel.2018.04.026. PMC 8317213. PMID 29861158.
- ^ Barth VC, Zeng JM, Vvedenskaya IO, Ouyang M, Husson RN, Woychik NA (July 2019). "Toxin-mediated ribosome stalling reprograms the Mycobacterium tuberculosis proteome". Nature Communications. 10 (1): 3035. Bibcode: 2019NatCo..10.3035B. doi: 10.1038/s41467-019-10869-8. PMC 6620280. PMID 31292443.
- ^ Barth VC, Woychik NA (2019). "The Sole Mycobacterium smegmatis MazF Toxin Targets tRNALys to Impart Highly Selective, Codon-Dependent Proteome Reprogramming". Frontiers in Genetics. 10: 1356. doi: 10.3389/fgene.2019.01356. PMC 7033543. PMID 32117414.
- ^ Schifano JM, Edifor R, Sharp JD, Ouyang M, Konkimalla A, Husson RN, Woychik NA (May 2013). "Mycobacterial toxin MazF-mt6 inhibits translation through cleavage of 23S rRNA at the ribosomal A site". Proceedings of the National Academy of Sciences of the United States of America. 110 (21): 8501–6. Bibcode: 2013PNAS..110.8501S. doi: 10.1073/pnas.1222031110. PMC 3666664. PMID 23650345.
- ^ Christensen-Dalsgaard M, Overgaard M, Winther KS, Gerdes K (2008). "Chapter 25 RNA Decay by Messenger RNA Interferases". RNA Turnover in Bacteria, Archaea and Organelles. Methods in Enzymology. Vol. 447. pp. 521–35. doi: 10.1016/S0076-6879(08)02225-8. ISBN 978-0-12-374377-0. PMID 19161859.
- ^ Yamaguchi Y, Inouye M (2009). "Chapter 12 mRNA Interferases, Sequence-Specific Endoribonucleases from the Toxin–Antitoxin Systems". mRNA interferases, sequence-specific endoribonucleases from the toxin-antitoxin systems. Progress in Molecular Biology and Translational Science. Vol. 85. pp. 467–500. doi: 10.1016/S0079-6603(08)00812-X. ISBN 978-0-12-374761-7. PMID 19215780.
- ^ Ramisetty BC (2020). "Regulation of Type II Toxin-Antitoxin Systems: The Translation-Responsive Model". Frontiers in Microbiology. 11: 895. doi: 10.3389/fmicb.2020.00895. PMC 7214741. PMID 32431690.
- ^ a b Mutschler H, Meinhart A (December 2011). "ε/ζ systems: their role in resistance, virulence, and their potential for antibiotic development". Journal of Molecular Medicine. 89 (12): 1183–94. doi: 10.1007/s00109-011-0797-4. PMC 3218275. PMID 21822621.
- ^ Hallez R, Geeraerts D, Sterckx Y, Mine N, Loris R, Van Melderen L (May 2010). "New toxins homologous to ParE belonging to three-component toxin-antitoxin systems in Escherichia coli O157:H7" (PDF). Molecular Microbiology. 76 (3): 719–32. doi: 10.1111/j.1365-2958.2010.07129.x. PMID 20345661.
- ^ Bordes P, Cirinesi AM, Ummels R, Sala A, Sakr S, Bitter W, Genevaux P (May 2011). "SecB-like chaperone controls a toxin-antitoxin stress-responsive system in Mycobacterium tuberculosis". Proceedings of the National Academy of Sciences of the United States of America. 108 (20): 8438–43. Bibcode: 2011PNAS..108.8438B. doi: 10.1073/pnas.1101189108. PMC 3100995. PMID 21536872.
- ^ Fiebig A, Castro Rojas CM, Siegal-Gaskins D, Crosson S (July 2010). "Interaction specificity, toxicity and regulation of a paralogous set of ParE/RelE-family toxin-antitoxin systems". Molecular Microbiology. 77 (1): 236–51. doi: 10.1111/j.1365-2958.2010.07207.x. PMC 2907451. PMID 20487277. (subscription required)
- ^ Singletary LA, Gibson JL, Tanner EJ, McKenzie GJ, Lee PL, Gonzalez C, Rosenberg SM (December 2009). "An SOS-regulated type 2 toxin-antitoxin system". Journal of Bacteriology. 191 (24): 7456–65. doi: 10.1128/JB.00963-09. PMC 2786605. PMID 19837801.
- ^ Jørgensen MG, Pandey DP, Jaskolska M, Gerdes K (February 2009). "HicA of Escherichia coli defines a novel family of translation-independent mRNA interferases in bacteria and archaea". Journal of Bacteriology. 191 (4): 1191–9. doi: 10.1128/JB.01013-08. PMC 2631989. PMID 19060138.
- ^ Jurėnas D, Chatterjee S, Konijnenberg A, Sobott F, Droogmans L, Garcia-Pino A, Van Melderen L (June 2017). "fMet" (PDF). Nature Chemical Biology. 13 (6): 640–646. doi: 10.1038/nchembio.2346. PMID 28369041.
- ^ Blower TR, Fineran PC, Johnson MJ, Toth IK, Humphreys DP, Salmond GP (October 2009). "Mutagenesis and functional characterization of the RNA and protein components of the toxIN abortive infection and toxin-antitoxin locus of Erwinia". Journal of Bacteriology. 191 (19): 6029–39. doi: 10.1128/JB.00720-09. PMC 2747886. PMID 19633081.
- ^ Blower TR, Pei XY, Short FL, Fineran PC, Humphreys DP, Luisi BF, Salmond GP (February 2011). "A processed noncoding RNA regulates an altruistic bacterial antiviral system". Nature Structural & Molecular Biology. 18 (2): 185–90. doi: 10.1038/nsmb.1981. PMC 4612426. PMID 21240270.
- ^ Brown JM, Shaw KJ (November 2003). "A novel family of Escherichia coli toxin-antitoxin gene pairs". Journal of Bacteriology. 185 (22): 6600–8. doi: 10.1128/jb.185.22.6600-6608.2003. PMC 262102. PMID 14594833.
- ^ Jankevicius G, Ariza A, Ahel M, Ahel I (December 2016). "The Toxin-Antitoxin System DarTG Catalyzes Reversible ADP-Ribosylation of DNA". Molecular Cell. 64 (6): 1109–1116. doi: 10.1016/j.molcel.2016.11.014. PMC 5179494. PMID 27939941.
- ^ Schuller M, Butler RE, Ariza A, Tromans-Coia C, Jankevicius G, Claridge TD, et al. (August 2021). "Molecular basis for DarT ADP-ribosylation of a DNA base". Nature. 596 (7873): 597–602. Bibcode: 2021Natur.596..597S. doi: 10.1038/s41586-021-03825-4. hdl: 2299/25013. PMID 34408320. S2CID 237214909.
- ^ Schuller, Marion; Raggiaschi, Roberto; Mikolcevic, Petra; Rack, Johannes G. M.; Ariza, Antonio; Zhang, YuGeng; Ledermann, Raphael; Tang, Christoph; Mikoc, Andreja; Ahel, Ivan (2023-07-06). "Molecular basis for the reversible ADP-ribosylation of guanosine bases". Molecular Cell. 83 (13): 2303–2315.e6. doi: 10.1016/j.molcel.2023.06.013. ISSN 1097-2765. PMID 37390817. S2CID 259304277.
- ^ Wang X, Lord DM, Hong SH, Peti W, Benedik MJ, Page R, Wood TK (June 2013). "Type II toxin/antitoxin MqsR/MqsA controls type V toxin/antitoxin GhoT/GhoS". Environmental Microbiology. 15 (6): 1734–44. Bibcode: 2013EnvMi..15.1734W. doi: 10.1111/1462-2920.12063. PMC 3620836. PMID 23289863.
- ^ Aakre CD, Phung TN, Huang D, Laub MT (December 2013). "A bacterial toxin inhibits DNA replication elongation through a direct interaction with the β sliding clamp". Molecular Cell. 52 (5): 617–28. doi: 10.1016/j.molcel.2013.10.014. PMC 3918436. PMID 24239291.
- ^ Wang X, Yao J, Sun YC, Wood TK (May 2021). "Type VII Toxin/Antitoxin Classification System for Antitoxins that Enzymatically Neutralize Toxins". Trends in Microbiology. 29 (5): 388–393. doi: 10.1016/j.tim.2020.12.001. PMID 33342606. S2CID 229341165.
- ^ Li, Ming; Gong, Luyao; Cheng, Feiyue; Yu, Haiying; Zhao, Dahe; Wang, Rui; Wang, Tian; Zhang, Shengjie; Zhou, Jian; Shmakov, Sergey A.; Koonin, Eugene V.; Xiang, Hua (2021-04-30). "Toxin-antitoxin RNA pairs safeguard CRISPR-Cas systems". Science. 372 (6541): eabe5601. doi: 10.1126/science.abe5601. ISSN 0036-8075. PMID 33926924. S2CID 233448823.
- ^ Wu K, Jahng D, Wood TK (1994). "Temperature and growth rate effects on the hok/sok killer locus for enhanced plasmid stability". Biotechnology Progress. 10 (6): 621–9. doi: 10.1021/bp00030a600. PMID 7765697. S2CID 34815594.
- ^ Pecota DC, Kim CS, Wu K, Gerdes K, Wood TK (May 1997). "Combining the hok/sok, parDE, and pnd postsegregational killer loci to enhance plasmid stability". Applied and Environmental Microbiology. 63 (5): 1917–24. Bibcode: 1997ApEnM..63.1917P. doi: 10.1128/AEM.63.5.1917-1924.1997. PMC 168483. PMID 9143123.
- ^ a b Gerdes K, Christensen SK, Løbner-Olesen A (May 2005). "Prokaryotic toxin-antitoxin stress response loci". Nature Reviews. Microbiology. 3 (5): 371–82. doi: 10.1038/nrmicro1147. PMID 15864262. S2CID 13417307.
- ^ a b Bernard P, Gabant P, Bahassi EM, Couturier M (October 1994). "Positive-selection vectors using the F plasmid ccdB killer gene". Gene. 148 (1): 71–4. doi: 10.1016/0378-1119(94)90235-6. PMID 7926841.
- ^ Torres B, Jaenecke S, Timmis KN, García JL, Díaz E (December 2003). "A dual lethal system to enhance containment of recombinant micro-organisms". Microbiology. 149 (Pt 12): 3595–601. doi: 10.1099/mic.0.26618-0. PMID 14663091.
External links
- RASTA – Rapid Automated Scan for Toxins and Antitoxins in Bacteria