Genomic structural variation is the variation in structure of an organism's chromosome, such as deletions, duplications, copy-number variants, insertions, inversions and translocations. Originally, a structure variation affects a sequence length about 1kb to 3Mb, which is larger than SNPs and smaller than chromosome abnormality (though the definitions have some overlap). [1] However, the operational range of structural variants has widened to include events > 50bp. [2] Some structural variants are associated with genetic diseases, however most are not. [3] [4] Approximately 13% of the human genome is defined as structurally variant in the normal population, and there are at least 240 genes that exist as homozygous deletion polymorphisms in human populations, suggesting these genes are dispensable in humans. [4] While humans carry a median of 3.6 Mbp in SNPs (compared to a reference genome), a median of 8.9 Mbp is affected by structual variation which thus causes most genetic differences between humans in terms of raw sequence data. [4]
Microscopic structural variation
Microscopic means that it can be detected with optical microscopes, such as aneuploidies, marker chromosome, gross rearrangements and variation in chromosome size. [5] [6] The frequency in human population is thought to be underestimated due to the fact that some of these are not actually easy to identify. These structural abnormalities exist in 1 of every 375 live births by putative information. [7]
Sub-microscopic structural variation
Sub-microscopic structural variants are much harder to detect owing to their small size. The first study in 2004 that used DNA microarrays could detect tens of genetic loci that exhibited copy number variation, deletions and duplications, greater than 100 kilobases in the human genome. [8] However, by 2015 whole genome sequencing studies could detect around 5,000 of structural variants as small as 100 base pairs encompassing approximately 20 megabases in each individual genome. [3] [4] These structural variants include deletions, tandem duplications, inversions, mobile element insertions. The mutation rate is also much higher than microscopic structural variants, estimated by two studies at 16% and 20% respectively, both of which are probably underestimates due to the challenges of accurately detecting structural variants. [3] [9] It has also been shown that the generation of spontaneous structural variants significantly increases the likelihood of generating further spontaneous single nucleotide variants or indels within 100 kilobases of the structural variation event. [3]
Copy-number variation
Copy-number variation (CNV) is a large category of structural variation, which includes insertions, deletions and duplications. In recent studies, copy-number variations are tested on people who do not have genetic diseases, using methods that are used for quantitative SNP genotyping. Results show that 28% of the suspected regions in the individuals actually do contain copy number variations. [10] [11] Also, CNVs in human genome affect more nucleotides than Single Nucleotide Polymorphism (SNP). It is also noteworthy that many of CNVs are not in coding regions. Because CNVs are usually caused by unequal recombination, widespread similar sequences such as LINEs and SINEs may be a common mechanism of CNV creation. [12] [13]
Inversion
There are several inversions known which are related to human disease. For instance, recurrent 400kb inversion in factor VIII gene is a common cause of haemophilia A, [14] and smaller inversions affecting idunorate 2-sulphatase (IDS) will cause Hunter syndrome. [15] More examples include Angelman syndrome and Sotos syndrome. However, recent research shows that one person can have 56 putative inversions, thus the non-disease inversions are more common than previously supposed. Also in this study it's indicated that inversion breakpoints are commonly associated with segmental duplications. [16] One 900 kb inversion in the chromosome 17 is under positive selection and are predicted to increase its frequency in European population. [17]
Other structural variants
More complex structural variants can occur include a combination of the above in a single event. [3] The most common type of complex structural variation are non-tandem duplications, where sequence is duplicated and inserted in inverted or direct orientation into another part of the genome. [3] Other classes of complex structural variant include deletion-inversion-deletions, duplication-inversion-duplications, and tandem duplications with nested deletions. [3] There are also cryptic translocations and segmental uniparental disomy (UPD). There are increasing reports of these variations, but are more difficult to detect than traditional variations because these variants are balanced and array-based or PCR-based methods are not able to locate them. [18]
Structural variation and phenotypes
Some genetic diseases are suspected to be caused by structural variations, but the relation is not very certain. It is not plausible to divide these variants into two classes as "normal" or "disease", because the actual output of the same variant will also vary. Also, a few of the variants are actually positively selected for (mentioned above). A series of studies have shown that gene disrupting spontaneous (de novo) CNVs disrupt genes approximately four times more frequently in autism than in controls and contribute to approximately 5–10% of cases. [3] [19] [20] [21] [22] Inherited variants also contribute to around 5–10% of cases of autism. [3]
Structural variations also have its function in population genetics. Different frequency of a same variation can be used as a genetic mark to infer relationship between populations in different areas. A complete comparison between human and chimpanzee structural variation also suggested that some of these may be fixed in one species because of its adaptative function. [23] There are also deletions related to resistance against malaria and AIDS. [24] [25] Also, some highly variable segments are thought to be caused by balancing selection, but there are also studies against this hypothesis. [26]
Database of structural variation
Some of genome browsers and bioinformatic databases have a list of structural variations in human genome with an emphasis on CNVs, and can show them in the genome browsing page, for example, UCSC Genome Browser. [27] Under the page viewing a part of the genome, there are "Common Cell CNVs" and "Structural Var" which can be enabled. On NCBI, there is a special page [28] for structural variation. In that system, both "inner" and "outer" coordinates are shown; they are both not actual breakpoints, but surmised minimal and maximum range of sequence affected by the structural variation. The types are classified as insertion, loss, gain, inversion, LOH, everted, transchr and UPD.[ citation needed]
Methods of detection
New methods have been developed to analyze human genetic structural variation at high resolutions. The methods used to test the genome are in either a specific targeted way or in a genome wide manner. For Genome wide tests, array-based comparative genome hybridization approaches bring the best genome wide scans to find new copy number variants. [30] These techniques use DNA fragments that are labeled from a genome of interest and are hybridized, with another genome labeled differently, to arrays spotted with cloned DNA fragments. This reveals copy number differences between two genomes. [30]
For targeted genome examinations, the best assays for checking specific areas of the genome are primarily PCR based. The best established of the PCR based methods is real time quantitative polymerase chain reaction (qPCR). [30] A different approach is to specifically check certain areas that surround known segmental duplications since they are usually areas of copy number variation. [30] An SNP genotyping method that offers independent fluorescence intensities for two alleles can be used to target the nucleotides in between two copies of a segmental duplication. [30] From this, an increase in intensity from one of the alleles compared to the other can be observed.
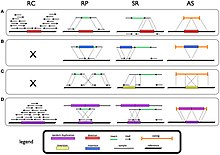
With the development of next-generation sequencing (NGS) technology, four classes of strategies for the detection of structural variants with NGS data have been reported, with each being based on patterns that are diagnostic of different classes of SV. [31] [29] [32] [33]
- Read-depth or read-count methods assume a random distribution (e.g. Poisson distribution) of reads from short read sequencing. The divergence from this distribution is investigated to discover duplications and deletions. Regions with duplication will show higher read depth while those with deletion will result in lower read depth.
- Split-read methods enable detection of insertions (including mobile element insertions) and deletions down to single base-pair resolution. The presence of a SV is identified from discontinuous alignment to the reference genome. A gap in the read marks a deletion and in the reference marks an insertion.
- Read pair methods examine the length and orientation of paired-end reads from short read sequencing data. For example, read pairs further apart than expected indicate a deletion. Translocations, inversions and tandem duplications can likewise be discovered using read-pairs.
- De novo sequence assembly may be applied with reads that are accurate enough. While, in practice, use of this method is limited by the length of sequence reads, long read based genome assemblies offer structural variation discovery for classes such as insertions that escape detection when using other methods. [34]
See also
References
- ^ Feuk, Lars; Carson, Andrew R.; Scherer, Stephen W. (2006). "Structural variation in the human genome". Nature Reviews Genetics. 7 (2): 85–97. doi: 10.1038/nrg1767. PMID 16418744. S2CID 17255998.
- ^ Alkan, Can; Coe, Bradley P.; Eichler, Evan E. (2011-03-01). "Genome structural variation discovery and genotyping". Nature Reviews Genetics. 12 (5): 363–376. doi: 10.1038/nrg2958. ISSN 1471-0056. PMC 4108431. PMID 21358748.
- ^ a b c d e f g h i Brandler, William M.; Antaki, Danny; Gujral, Madhusudan; Noor, Amina; Rosanio, Gabriel; Chapman, Timothy R.; Barrera, Daniel J.; Lin, Guan Ning; Malhotra, Dheeraj; Watts, Amanda C.; Wong, Lawrence C.; Estabillo, Jasper A.; Gadomski, Therese E.; Hong, Oanh; Fajardo, Karin V. Fuentes; Bhandari, Abhishek; Owen, Renius; Baughn, Michael; Yuan, Jeffrey; Solomon, Terry; Moyzis, Alexandra G.; Maile, Michelle S.; Sanders, Stephan J.; Reiner, Gail E.; Vaux, Keith K.; Strom, Charles M.; Zhang, Kang; Muotri, Alysson R.; Akshoomoff, Natacha; Leal, Suzanne M.; Pierce, Karen; Courchesne, Eric; Iakoucheva, Lilia M.; Corsello, Christina; Sebat, Jonathan (24 March 2016). "Frequency and Complexity of De Novo Structural Mutation in Autism". The American Journal of Human Genetics. 98 (4): 667–79. doi: 10.1016/j.ajhg.2016.02.018. PMC 4833290. PMID 27018473.
- ^ a b c d Sudmant, Peter H.; Rausch, Tobias; Gardner, Eugene J.; Handsaker, Robert E.; Abyzov, Alexej; Huddleston, John; Zhang, Yan; Ye, Kai; Jun, Goo; Hsi-Yang Fritz, Markus; Konkel, Miriam K.; Malhotra, Ankit; Stütz, Adrian M.; Shi, Xinghua; Paolo Casale, Francesco; Chen, Jieming; Hormozdiari, Fereydoun; Dayama, Gargi; Chen, Ken; Malig, Maika; Chaisson, Mark J. P.; Walter, Klaudia; Meiers, Sascha; Kashin, Seva; Garrison, Erik; Auton, Adam; Lam, Hugo Y. K.; Jasmine Mu, Xinmeng; Alkan, Can; Antaki, Danny; Bae, Taejeong; Cerveira, Eliza; Chines, Peter; Chong, Zechen; Clarke, Laura; Dal, Elif; Ding, Li; Emery, Sarah; Fan, Xian; Gujral, Madhusudan; Kahveci, Fatma; Kidd, Jeffrey M.; Kong, Yu; Lameijer, Eric-Wubbo; McCarthy, Shane; Flicek, Paul; Gibbs, Richard A.; Marth, Gabor; Mason, Christopher E.; Menelaou, Androniki; Muzny, Donna M.; Nelson, Bradley J.; Noor, Amina; Parrish, Nicholas F.; Pendleton, Matthew; Quitadamo, Andrew; Raeder, Benjamin; Schadt, Eric E.; Romanovitch, Mallory; Schlattl, Andreas; Sebra, Robert; Shabalin, Andrey A.; Untergasser, Andreas; Walker, Jerilyn A.; Wang, Min; Yu, Fuli; Zhang, Chengsheng; Zhang, Jing; Zheng-Bradley, Xiangqun; Zhou, Wanding; Zichner, Thomas; Sebat, Jonathan; Batzer, Mark A.; McCarroll, Steven A.; Mills, Ryan E.; Gerstein, Mark B.; Bashir, Ali; Stegle, Oliver; Devine, Scott E.; Lee, Charles; Eichler, Evan E.; Korbel, Jan O. (30 September 2015). "An integrated map of structural variation in 2,504 human genomes". Nature. 526 (7571): 75–81. Bibcode: 2015Natur.526...75.. doi: 10.1038/nature15394. PMC 4617611. PMID 26432246.
- ^ Reich, David E.; Schaffner, Stephen F.; Daly, Mark J.; McVean, Gil; Mullikin, James C.; Higgins, John M.; Richter, Daniel J.; Lander, Eric S.; Altshuler, David (2002). "Human genome sequence variation and the influence of gene history, mutation and recombination". Nature Genetics. 32 (1): 135–42. doi: 10.1038/ng947. PMID 12161752. S2CID 16822751.
- ^ Gripenberg, Ulla (1964). "Size variation and orientation of the human Y chromosome". Chromosoma. 15 (5): 618–29. doi: 10.1007/BF00319995. PMID 14333154. S2CID 26549548.
- ^ Wyandt, H. E.; Tonk, V. S. (2004). Atlas of Human Chromosome Heteromorphisms. Netherlands: Kluwer Academic. ISBN 978-90-481-6296-3.[ page needed]
- ^ Sebat, J. (23 July 2004). "Large-Scale Copy Number Polymorphism in the Human Genome". Science. 305 (5683): 525–528. Bibcode: 2004Sci...305..525S. doi: 10.1126/science.1098918. PMID 15273396. S2CID 20357402.
- ^ Kloosterman, Wigard P.; Francioli, Laurent C.; Hormozdiari, Fereydoun; Marschall, Tobias; Hehir-Kwa, Jayne Y.; Abdellaoui, Abdel; Lameijer, Eric-Wubbo; Moed, Matthijs H.; Koval, Vyacheslav; Renkens, Ivo; van Roosmalen, Markus J.; Arp, Pascal; Karssen, Lennart C.; Coe, Bradley P.; Handsaker, Robert E.; Suchiman, Eka D.; Cuppen, Edwin; Thung, Djie Tjwan; McVey, Mitch; Wendl, Michael C.; Uitterlinden, André; van Duijn, Cornelia M.; Swertz, Morris A.; Wijmenga, Cisca; van Ommen, GertJan B.; Slagboom, P. Eline; Boomsma, Dorret I.; Schönhuth, Alexander; Eichler, Evan E.; de Bakker, Paul I.W.; Ye, Kai; Guryev, Victor (June 2015). "Characteristics of de novo structural changes in the human genome". Genome Research. 25 (6): 792–801. doi: 10.1101/gr.185041.114. PMC 4448676. PMID 25883321.
- ^ Sebat, J.; Lakshmi, B; Troge, J; Alexander, J; Young, J; Lundin, P; Månér, S; Massa, H; et al. (2004). "Large-Scale Copy Number Polymorphism in the Human Genome". Science. 305 (5683): 525–8. Bibcode: 2004Sci...305..525S. doi: 10.1126/science.1098918. PMID 15273396. S2CID 20357402.
- ^ Iafrate, A John; Feuk, Lars; Rivera, Miguel N; Listewnik, Marc L; Donahoe, Patricia K; Qi, Ying; Scherer, Stephen W; Lee, Charles (2004). "Detection of large-scale variation in the human genome". Nature Genetics. 36 (9): 949–51. doi: 10.1038/ng1416. PMID 15286789.
- ^ Lupski, James R. (2010). "Retrotransposition and Structural Variation in the Human Genome". Cell. 141 (7): 1110–2. doi: 10.1016/j.cell.2010.06.014. PMID 20602993. S2CID 2047696.
- ^ Lam, Hugo YK; Mu, Xinmeng Jasmine; Stutz, Adrian M; Tanzer, Andrea; Cayting, Philip D; Snyder, Michael; Kim, Philip M; Korbel, Jan O; Gerstein, Mark B (2010). "Nucleotide-resolution analysis of structural variants using BreakSeq and a breakpoint library". Nature Biotechnology. 28 (1): 47–55. doi: 10.1038/nbt.1600. PMC 2951730. PMID 20037582.
- ^ Lakich, Delia; Kazazian, Haig H.; Antonarakis, Stylianos E.; Gitschier, Jane (1993). "Inversions disrupting the factor VIII gene are a common cause of severe haemophilia A". Nature Genetics. 5 (3): 236–41. doi: 10.1038/ng1193-236. PMID 8275087. S2CID 25636383.
- ^ Bondeson, Maire-Louise; Dahl, Niklas; Malmgren, Helena; Kleijer, Wim J.; Tönnesen, Tönne; Carlberg, Britt-Marie; Pettersson, Ulf (1995). "Inversion of the IDS gene resulting from recombination with IDS-related sequences in a common cause of the Hunter syndrome". Human Molecular Genetics. 4 (4): 615–21. doi: 10.1093/hmg/4.4.615. PMID 7633410.
- ^ Tuzun, Eray; Sharp, Andrew J; Bailey, Jeffrey A; Kaul, Rajinder; Morrison, V Anne; Pertz, Lisa M; Haugen, Eric; Hayden, Hillary; et al. (2005). "Fine-scale structural variation of the human genome". Nature Genetics. 37 (7): 727–32. doi: 10.1038/ng1562. PMID 15895083. S2CID 14162962.
- ^ Stefansson, Hreinn; Helgason, Agnar; Thorleifsson, Gudmar; Steinthorsdottir, Valgerdur; Masson, Gisli; Barnard, John; Baker, Adam; Jonasdottir, Aslaug; et al. (2005). "A common inversion under selection in Europeans". Nature Genetics. 37 (2): 129–37. doi: 10.1038/ng1508. PMID 15654335. S2CID 120515.
-
^ Sung, Wing-Kin (18 May 2017).
Algorithms for next-generation sequencing. Boca Raton. p. 215.
ISBN
978-1-4665-6551-7.
OCLC
987790994.
{{ cite book}}: CS1 maint: location missing publisher ( link) - ^ Sebat, J.; Lakshmi, B.; Malhotra, D.; Troge, J.; Lese-Martin, C.; Walsh, T.; Yamrom, B.; Yoon, S.; Krasnitz, A.; Kendall, J.; Leotta, A.; Pai, D.; Zhang, R.; Lee, Y.-H.; Hicks, J.; Spence, S. J.; Lee, A. T.; Puura, K.; Lehtimaki, T.; Ledbetter, D.; Gregersen, P. K.; Bregman, J.; Sutcliffe, J. S.; Jobanputra, V.; Chung, W.; Warburton, D.; King, M.-C.; Skuse, D.; Geschwind, D. H.; Gilliam, T. C.; Ye, K.; Wigler, M. (20 April 2007). "Strong Association of De Novo Copy Number Mutations with Autism". Science. 316 (5823): 445–449. Bibcode: 2007Sci...316..445S. doi: 10.1126/science.1138659. PMC 2993504. PMID 17363630.
- ^ Pinto, Dalila; Delaby, Elsa; Merico, Daniele; Barbosa, Mafalda; Merikangas, Alison; Klei, Lambertus; Thiruvahindrapuram, Bhooma; Xu, Xiao; Ziman, Robert; Wang, Zhuozhi; Vorstman, Jacob A.S.; Thompson, Ann; Regan, Regina; Pilorge, Marion; Pellecchia, Giovanna; Pagnamenta, Alistair T.; Oliveira, Bárbara; Marshall, Christian R.; Magalhaes, Tiago R.; Lowe, Jennifer K.; Howe, Jennifer L.; Griswold, Anthony J.; Gilbert, John; Duketis, Eftichia; Dombroski, Beth A.; De Jonge, Maretha V.; Cuccaro, Michael; Crawford, Emily L.; Correia, Catarina T.; et al. (May 2014). "Convergence of Genes and Cellular Pathways Dysregulated in Autism Spectrum Disorders". The American Journal of Human Genetics. 94 (5): 677–694. doi: 10.1016/j.ajhg.2014.03.018. PMC 4067558. PMID 24768552.
- ^ Levy, Dan; Ronemus, Michael; Yamrom, Boris; Lee, Yoon-ha; Leotta, Anthony; Kendall, Jude; Marks, Steven; Lakshmi, B.; Pai, Deepa; Ye, Kenny; Buja, Andreas; Krieger, Abba; Yoon, Seungtai; Troge, Jennifer; Rodgers, Linda; Iossifov, Ivan; Wigler, Michael (June 2011). "Rare De Novo and Transmitted Copy-Number Variation in Autistic Spectrum Disorders". Neuron. 70 (5): 886–897. doi: 10.1016/j.neuron.2011.05.015. PMID 21658582. S2CID 11132936.
- ^ Sanders, Stephan J.; Ercan-Sencicek, A. Gulhan; Hus, Vanessa; Luo, Rui; Murtha, Michael T.; Moreno-De-Luca, Daniel; Chu, Su H.; Moreau, Michael P.; Gupta, Abha R.; Thomson, Susanne A.; Mason, Christopher E.; Bilguvar, Kaya; Celestino-Soper, Patricia B.S.; Choi, Murim; Crawford, Emily L.; Davis, Lea; Davis Wright, Nicole R.; Dhodapkar, Rahul M.; DiCola, Michael; DiLullo, Nicholas M.; Fernandez, Thomas V.; Fielding-Singh, Vikram; Fishman, Daniel O.; Frahm, Stephanie; Garagaloyan, Rouben; Goh, Gerald S.; Kammela, Sindhuja; Klei, Lambertus; Lowe, Jennifer K.; Lund, Sabata C.; McGrew, Anna D.; Meyer, Kyle A.; Moffat, William J.; Murdoch, John D.; O'Roak, Brian J.; Ober, Gordon T.; Pottenger, Rebecca S.; Raubeson, Melanie J.; Song, Youeun; Wang, Qi; Yaspan, Brian L.; Yu, Timothy W.; Yurkiewicz, Ilana R.; Beaudet, Arthur L.; Cantor, Rita M.; Curland, Martin; Grice, Dorothy E.; Günel, Murat; Lifton, Richard P.; Mane, Shrikant M.; Martin, Donna M.; Shaw, Chad A.; Sheldon, Michael; Tischfield, Jay A.; Walsh, Christopher A.; Morrow, Eric M.; Ledbetter, David H.; Fombonne, Eric; Lord, Catherine; Martin, Christa Lese; Brooks, Andrew I.; Sutcliffe, James S.; Cook, Edwin H.; Geschwind, Daniel; Roeder, Kathryn; Devlin, Bernie; State, Matthew W. (June 2011). "Multiple Recurrent De Novo CNVs, Including Duplications of the 7q11.23 Williams Syndrome Region, Are Strongly Associated with Autism". Neuron. 70 (5): 863–885. doi: 10.1016/j.neuron.2011.05.002. PMC 3939065. PMID 21658581.
- ^ Johnson, Matthew E.; Viggiano, Luigi; Bailey, Jeffrey A.; Abdul-Rauf, Munah; Goodwin, Graham; Rocchi, Mariano; Eichler, Evan E. (2001). "Positive selection of a gene family during the emergence of humans and African apes". Nature. 413 (6855): 514–9. Bibcode: 2001Natur.413..514J. doi: 10.1038/35097067. PMID 11586358. S2CID 4327069.
- ^ Redon, Richard; Ishikawa, Shumpei; Fitch, Karen R.; Feuk, Lars; Perry, George H.; Andrews, T. Daniel; Fiegler, Heike; Shapero, Michael H.; et al. (2006). "Global variation in copy number in the human genome". Nature. 444 (7118): 444–54. Bibcode: 2006Natur.444..444R. doi: 10.1038/nature05329. PMC 2669898. PMID 17122850.
- ^ Gonzalez, E.; Kulkarni, H; Bolivar, H; Mangano, A; Sanchez, R; Catano, G; Nibbs, RJ; Freedman, BI; et al. (2005). "The Influence of CCL3L1 Gene-Containing Segmental Duplications on HIV-1/AIDS Susceptibility". Science. 307 (5714): 1434–40. Bibcode: 2005Sci...307.1434G. doi: 10.1126/science.1101160. PMID 15637236. S2CID 8815153.
- ^ Bubb, K. L.; Bovee, D; Buckley, D; Haugen, E; Kibukawa, M; Paddock, M; Palmieri, A; Subramanian, S; et al. (2006). "Scan of Human Genome Reveals No New Loci Under Ancient Balancing Selection". Genetics. 173 (4): 2165–77. doi: 10.1534/genetics.106.055715. PMC 1569689. PMID 16751668.
- ^ "Human hg38 chr1:11,102,837-11,267,747 UCSC Genome Browser v374".
- ^ "Overview of Structural Variation".
- ^ a b Tattini, Lorenzo; D'Aurizio, Romina; Magi, Alberto (2015). "Detection of genomic structural variants from next-generation sequencing data". Frontiers in Bioengineering and Biotechnology. 3 (92): 92. doi: 10.3389/fbioe.2015.00092. PMC 4479793. PMID 26161383.
- ^ a b c d e Feuk, L.; Carson, A.R.; Schere, S.W. (2006). "Structural variation in the human genome". Nature Reviews Genetics. 7 (2): 85–97. doi: 10.1038/nrg1767. PMID 16418744. S2CID 17255998.
- ^ Korbel JO, Urban AE, Affourtit JP, Godwin B, Grubert F, Simons JF, Kim PM, Palejev D, Carriero NJ, Du L, Taillon BE, Chen Z, Tanzer A, Saunders AC, Chi J, Yang F, Carter NP, Hurles ME, Weissman SM, Harkins TT, Gerstein MB, Egholm M, Snyder M (October 2007). "Paired-end mapping reveals extensive structural variation in the human genome". Science. 318 (5849): 420–6. Bibcode: 2007Sci...318..420K. doi: 10.1126/science.1149504. PMC 2674581. PMID 17901297.
- ^ Alkan, Can; Coe, Bradley P.; Eichler, Evan E. (2011). "Genome structural variation discovery and genotyping". Nature Reviews Genetics. 12 (5): 363–376. doi: 10.1038/nrg2958. PMC 4108431. PMID 21358748.
- ^ Kuzniar, Arnold; Maassen, Jason; Verhoeven, Stefan; Santuari, Luca; Shneider, Carl; Kloosterman, Wigard P.; de Ridder, Jeroen (2020). "sv-callers: a highly portable parallel workflow for structural variant detection in whole-genome sequence data". PeerJ. 8 (5): 2167–8359. doi: 10.7717/peerj.8214. PMC 6951283. PMID 31934500.
- ^ Ebert P, Audano PA, Zhu Q, Rodriguez-Martin B, Porubsky D, Bonder MJ, Sulovari A, Ebler J, Zhou W, Serra Mari R, Yilmaz F, Zhao X, Hsieh P, Lee J, Kumar S, Lin J, Rausch T, Chen Y, Ren J, Santamarina M, Höps W, Ashraf H, Chuang NT, Yang X, Munson KM, Lewis AP, Fairley S, Tallon LJ, Clarke WE, Basile AO, Byrska-Bishop M, Corvelo A, Evani US, Lu TY, Chaisson MJ, Chen J, Li C, Brand H, Wenger AM, Ghareghani M, Harvey WT, Raeder B, Hasenfeld P, Regier AA, Abel HJ, Hall IM, Flicek P, Stegle O, Gerstein MB, Tubio JM, Mu Z, Li YI, Shi X, Hastie AR, Ye K, Chong Z, Sanders AD, Zody MC, Talkowski ME, Mills RE, Devine SE, Lee C, Korbel JO, Marschall T, Eichler EE (April 2021). "Haplotype-resolved diverse human genomes and integrated analysis of structural variation". Science. 372 (6537). doi: 10.1126/science.abf7117. PMC 8026704. PMID 33632895.