| Ribosome-inactivating protein | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
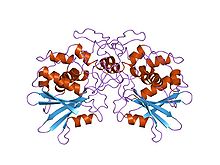 Structure of pokeweed antiviral protein.
[1] | |||||||||||
| Identifiers | |||||||||||
| Symbol | RIP | ||||||||||
| Pfam | PF00161 | ||||||||||
| InterPro | IPR001574 | ||||||||||
| PROSITE | PDOC00248 | ||||||||||
| SCOP2 | 1paf / SCOPe / SUPFAM | ||||||||||
| |||||||||||
A ribosome-inactivating protein (RIP) is a protein synthesis inhibitor that acts at the eukaryotic ribosome. [2] This protein family describes a large family of such proteins that work by acting as rRNA N-glycosylase (EC 3.2.2.22). They inactivate 60S ribosomal subunits by an N-glycosidic cleavage, which releases a specific adenine base from the sugar-phosphate backbone of 28S rRNA. [3] [4] [5] RIPs exist in bacteria and plants. [6]
Members of the family include shiga toxins, and type I (e.g. trichosanthin and luffin) and type II (e.g. ricin, agglutinin, and abrin) ribosome inactivating proteins (RIPs). All these toxins are structurally related. RIPs have been of considerable interest because of their potential use, conjugated with monoclonal antibodies, as immunotoxins to treat cancers. Further, trichosanthin has been shown to have potent activity against HIV-1-infected T cells and macrophages. [7] Elucidation of the structure-function relationships of RIPs has therefore become a major research effort. It is now known that RIPs are structurally related. A conserved glutamic residue has been implicated in the catalytic mechanism; [8] this lies near a conserved arginine, which also plays a role in catalysis. [9]
Only a minority of RIPs are toxic to humans when consumed, and proteins of this family are found in the vast majority of plants used for human consumption, such as Rice, Maize and Barley. In plants, they are thought to defend against pathogens and insects. [10]
Ribosome-inactivating proteins (RIPs) are separated into the following types based on protein domain composition: [11]
- Type I (A): RIPs-I are polypeptides composed of an A domain. This is the site of N-glycosidase activity.
- Type II (AB): RIPs-II are composed of an A domain with similar catalytic activity to Type I RIPs, and a B domain with carbohydrate-binding (lectin) properties. The B domain is able to bind galactosyl moieties on the cell surface which facilitates entry into the cell, thus making Type II particularly cytotoxic. The A and B domains are fused together by disulfide bonds. [11] [12] This group excludes bacterial AB5 toxins such as Shiga toxin, as the carbohydrate-binding ability evolved separately and these toxins are more similar to type I RIPs. [11]
- Type III: RIPs-III are separated into two subgroups. One subgroup (AC) contains the same original RIP domain (A), and a C-terminal with unknown functionality. The other subgroup (AD) is similar to Type I, but contains a site for inactivation. [11]
Examples include:
- Abrin
- Beetin
- Ricin
- Saporin
- Shiga toxin
- A Spiroplasma toxin [13]
- Trichosanthin
- Viscumin ( European mistletoe)
- Pokeweed antiviral protein ( Phytolacca americana) [14]
References
- ^ Monzingo AF, Collins EJ, Ernst SR, Irvin JD, Robertus JD (October 1993). "The 2.5 A structure of pokeweed antiviral protein". Journal of Molecular Biology. 233 (4): 705–15. doi: 10.1006/jmbi.1993.1547. PMID 8411176.
- ^ Ribosome+Inactivating+Proteins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- ^ Endo Y, Tsurugi K, Yutsudo T, Takeda Y, Ogasawara T, Igarashi K (January 1988). "Site of action of a Vero toxin (VT2) from Escherichia coli O157:H7 and of Shiga toxin on eukaryotic ribosomes. RNA N-glycosidase activity of the toxins". European Journal of Biochemistry. 171 (1–2): 45–50. doi: 10.1111/j.1432-1033.1988.tb13756.x. PMID 3276522.
- ^ May MJ, Hartley MR, Roberts LM, Krieg PA, Osborn RW, Lord JM (January 1989). "Ribosome inactivation by ricin A chain: a sensitive method to assess the activity of wild-type and mutant polypeptides". The EMBO Journal. 8 (1): 301–8. doi: 10.1002/j.1460-2075.1989.tb03377.x. PMC 400803. PMID 2714255.
- ^ Funatsu G, Islam MR, Minami Y, Sung-Sil K, Kimura M (1991). "Conserved amino acid residues in ribosome-inactivating proteins from plants". Biochimie. 73 (7–8): 1157–61. doi: 10.1016/0300-9084(91)90160-3. PMID 1742358.
- ^ Mak AN, Wong YT, An YJ, Cha SS, Sze KH, Au SW, et al. (2007). "Structure-function study of maize ribosome-inactivating protein: implications for the internal inactivation region and the sole glutamate in the active site". Nucleic Acids Research. 35 (18): 6259–67. doi: 10.1093/nar/gkm687. PMC 2094058. PMID 17855394.
- ^ Zhou K, Fu Z, Chen M, Lin Y, Pan K (May 1994). "Structure of trichosanthin at 1.88 A resolution". Proteins. 19 (1): 4–13. doi: 10.1002/prot.340190103. PMID 8066085. S2CID 21524411.
- ^ Hovde CJ, Calderwood SB, Mekalanos JJ, Collier RJ (April 1988). "Evidence that glutamic acid 167 is an active-site residue of Shiga-like toxin I". Proceedings of the National Academy of Sciences of the United States of America. 85 (8): 2568–72. Bibcode: 1988PNAS...85.2568H. doi: 10.1073/pnas.85.8.2568. PMC 280038. PMID 3357883.
- ^ Monzingo AF, Collins EJ, Ernst SR, Irvin JD, Robertus JD (October 1993). "The 2.5 A structure of pokeweed antiviral protein". Journal of Molecular Biology. 233 (4): 705–15. doi: 10.1006/jmbi.1993.1547. PMID 8411176.
- ^ Zhu, Feng; Zhou, Yang-Kai; Ji, Zhao-Lin; Chen, Xiao-Ren (9 February 2018). "The Plant Ribosome-Inactivating Proteins Play Important Roles in Defense against Pathogens and Insect Pest Attacks". Frontiers in Plant Science. 9: 146. doi: 10.3389/fpls.2018.00146. PMC 5811460. PMID 29479367.
- ^ a b c d Lapadula WJ, Ayub MJ (September 2017). "Ribosome Inactivating Proteins from an evolutionary perspective". Toxicon. 136: 6–14. doi: 10.1016/j.toxicon.2017.06.012. PMID 28651991. S2CID 9814488.
- ^ Fredriksson, Sten-Åke; Artursson, Elisabet; Bergström, Tomas; Östin, Anders; Nilsson, Calle; Åstot, Crister (December 2014). "Identification of RIP-II Toxins by Affinity Enrichment, Enzymatic Digestion and LC-MS". Analytical Chemistry. 87 (2): 967–974. doi: 10.1021/ac5032918. ISSN 0003-2700. PMID 25496503.
- ^ Hamilton PT, Peng F, Boulanger MJ, Perlman SJ (January 2016). "A ribosome-inactivating protein in a Drosophila defensive symbiont". Proceedings of the National Academy of Sciences of the United States of America. 113 (2): 350–5. Bibcode: 2016PNAS..113..350H. doi: 10.1073/pnas.1518648113. PMC 4720295. PMID 26712000.
- ^ Domashevskiy AV, Goss DJ (January 2015). "Pokeweed antiviral protein, a ribosome inactivating protein: activity, inhibition and prospects". Toxins. 7 (2): 274–98. doi: 10.3390/toxins7020274. PMC 4344624. PMID 25635465.