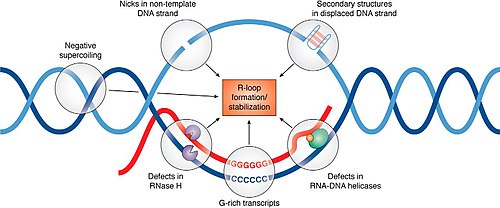
An R-loop is a three-stranded nucleic acid structure, composed of a DNA: RNA hybrid and the associated non-template single-stranded DNA. R-loops may be formed in a variety of circumstances and may be tolerated or cleared by cellular components. The term "R-loop" was given to reflect the similarity of these structures to D-loops; the "R" in this case represents the involvement of an RNA moiety.
In the laboratory, R-loops may also be created by the hybridization of mature mRNA with double-stranded DNA under conditions favoring the formation of a DNA-RNA hybrid; in this case, the intron regions (which have been spliced out of the mRNA) form single-stranded DNA loops, as they cannot hybridize with complementary sequence in the mRNA. [1]
History

R-looping was first described in 1976. [2] Independent R-looping studies from the laboratories of Richard J. Roberts and Phillip A. Sharp showed that protein coding adenovirus genes contained DNA sequences that were not present in the mature mRNA. [3] [4] Roberts and Sharp were awarded the Nobel Prize in 1993 for independently discovering introns. After their discovery in adenovirus, introns were found in a number of eukaryotic genes such as the eukaryotic ovalbumin gene (first by the O'Malley laboratory, then confirmed by other groups), [5] [6] hexon DNA, [3] and extrachromosomal rRNA genes of Tetrahymena thermophila. [7]
In the mid-1980s, development of an antibody that binds specifically to the R-loop structure opened the door for immunofluorescence studies, as well as genome-wide characterization of R-loop formation by DRIP-seq. [8]
R-loop mapping
R-loop mapping is a laboratory technique used to distinguish introns from exons in double-stranded DNA. [9] These R-loops are visualized by electron microscopy and reveal intron regions of DNA by creating unbound loops at these regions. [10]
R-loops in vivo
The potential for R-loops to serve as replication primers was demonstrated in 1980. [11] In 1994, R-loops were demonstrated to be present in vivo through analysis of plasmids isolated from E. coli mutants carrying mutations in topoisomerase. [12] This discovery of endogenous R-loops, in conjunction with rapid advances in genetic sequencing technologies, inspired a blossoming of R-loop research in the early 2000s that continues to this day. [13]
Regulation of R-loop formation and resolution
RNaseH enzymes are the primary proteins responsible for the dissolution of R-loops, acting to degrade the RNA moiety in order to allow the two complementary DNA strands to anneal. [14] Research over the past decade has identified more than 50 proteins that appear to influence R-loop accumulation, and while many of them are believed to contribute by sequestering or processing newly transcribed RNA to prevent re-annealing to the template, mechanisms of R-loop interaction for many of these proteins remain to be determined. [15]
Roles of R-loops in genetic regulation
R-loop formation is a key step in immunoglobulin class switching, a process that allows activated B cells to modulate antibody production. [16] They also appear to play a role in protecting some active promoters from methylation. [17] The presence of R-loops can also inhibit transcription. [18] Additionally, R-loop formation appears to be associated with “open” chromatin, characteristic of actively transcribed regions. [19] [20]
R-loops as genetic damage
When unscheduled R-loops form, they can cause damage by a number of different mechanisms. [21] Exposed single-stranded DNA can come under attack by endogenous mutagens, including DNA-modifying enzymes such as activation-induced cytidine deaminase, and can block replication forks to induce fork collapse and subsequent double-strand breaks. [22] As well, R-loops may induce unscheduled replication by acting as a primer. [11] [20]
R-loop accumulation has been associated with a number of diseases, including amyotrophic lateral sclerosis type 4 (ALS4), ataxia oculomotor apraxia type 2 (AOA2), Aicardi–Goutières syndrome, Angelman syndrome, Prader–Willi syndrome, and cancer. [13]
R-loops, Introns and DNA damage
Introns are non-coding regions within genes that are transcribed along with the coding regions of genes, but are subsequently removed from the primary RNA transcript by splicing. Actively transcribed regions of DNA often form R-loops that are vulnerable to DNA damage. Introns reduce R-loop formation and DNA damage in highly expressed yeast genes. [23] Genome-wide analysis showed that intron-containing genes display decreased R-loop levels and decreased DNA damage compared to intron-less genes of similar expression in both yeast and humans. [23] Inserting an intron within an R-loop prone gene can also suppress R-loop formation and recombination. Bonnet et al. (2017) [23] speculated that the function of introns in maintaining genetic stability may explain their evolutionary maintenance at certain locations, particularly in highly expressed genes.
See also
References
- ^ Wang, Kang; Wang, Honghong; Li, Conghui; Yin, Zhinang; Xiao, Ruijing; Li, Qiuzi; Xiang, Ying; Wang, Wen; Huang, Jian; Chen, Liang; Fang, Pingping; Liang, Kaiwei (2021-02-19). "Genomic profiling of native R loops with a DNA-RNA hybrid recognition sensor". Science Advances. 7 (8). Bibcode: 2021SciA....7.3516W. doi: 10.1126/sciadv.abe3516. ISSN 2375-2548. PMC 7888926. PMID 33597247.
- ^ Thomas M, White RL, Davis RW (July 1976). "Hybridization of RNA to double-stranded DNA: formation of R-loops". Proceedings of the National Academy of Sciences of the United States of America. 73 (7): 2294–8. Bibcode: 1976PNAS...73.2294T. doi: 10.1073/pnas.73.7.2294. PMC 430535. PMID 781674.
- ^ a b Berget SM, Moore C, Sharp PA (August 1977). "Spliced segments at the 5' terminus of adenovirus 2 late mRNA". Proceedings of the National Academy of Sciences of the United States of America. 74 (8): 3171–5. Bibcode: 1977PNAS...74.3171B. doi: 10.1073/pnas.74.8.3171. PMC 431482. PMID 269380.
- ^ Chow LT, Gelinas RE, Broker TR, Roberts RJ (September 1977). "An amazing sequence arrangement at the 5' ends of adenovirus 2 messenger RNA". Cell. 12 (1): 1–8. doi: 10.1016/0092-8674(77)90180-5. PMID 902310. S2CID 2099968.
- ^ Lai EC, Woo SL, Dugaiczyk A, Catterall JF, O'Malley BW (May 1978). "The ovalbumin gene: structural sequences in native chicken DNA are not contiguous". Proceedings of the National Academy of Sciences of the United States of America. 75 (5): 2205–9. Bibcode: 1978PNAS...75.2205L. doi: 10.1073/pnas.75.5.2205. PMC 392520. PMID 276861.
- ^ O'Hare K, Breathnach R, Benoist C, Chambon P (September 1979). "No more than seven interruptions in the ovalbumin gene: comparison of genomic and double-stranded cDNA sequences". Nucleic Acids Research. 7 (2): 321–34. doi: 10.1093/nar/7.2.321. PMC 328020. PMID 493147.
- ^ Cech TR, Rio DC (October 1979). "Localization of transcribed regions on extrachromosomal ribosomal RNA genes of Tetrahymena thermophila by R-loop mapping". Proceedings of the National Academy of Sciences of the United States of America. 76 (10): 5051–5. Bibcode: 1979PNAS...76.5051C. doi: 10.1073/pnas.76.10.5051. PMC 413077. PMID 291921.
- ^ Boguslawski SJ, Smith DE, Michalak MA, Mickelson KE, Yehle CO, Patterson WL, Carrico RJ (May 1986). "Characterization of monoclonal antibody to DNA.RNA and its application to immunodetection of hybrids". Journal of Immunological Methods. 89 (1): 123–30. doi: 10.1016/0022-1759(86)90040-2. PMID 2422282.
- ^ Woolford JL, Rosbash M (June 1979). "The use of R-looping for structural gene identification and mRNA purification". Nucleic Acids Research. 6 (7): 2483–97. doi: 10.1093/nar/6.7.2483. PMC 327867. PMID 379820.
- ^ King RC, Stansfield WD, Mulligan PK (2007). A Dictionary of Genetics. Oxford University Press 7.
- ^ a b Itoh T, Tomizawa J (May 1980). "Formation of an RNA primer for initiation of replication of ColE1 DNA by ribonuclease H". Proceedings of the National Academy of Sciences of the United States of America. 77 (5): 2450–4. Bibcode: 1980PNAS...77.2450I. doi: 10.1073/pnas.77.5.2450. PMC 349417. PMID 6156450.
- ^ Drolet M, Bi X, Liu LF (January 1994). "Hypernegative supercoiling of the DNA template during transcription elongation in vitro". The Journal of Biological Chemistry. 269 (3): 2068–74. doi: 10.1016/S0021-9258(17)42136-3. PMID 8294458.
- ^ a b Groh M, Gromak N (September 2014). "Out of balance: R-loops in human disease". PLOS Genetics. 10 (9): e1004630. doi: 10.1371/journal.pgen.1004630. PMC 4169248. PMID 25233079.
- ^ Cerritelli SM, Crouch RJ (March 2009). "Ribonuclease H: the enzymes in eukaryotes". The FEBS Journal. 276 (6): 1494–505. doi: 10.1111/j.1742-4658.2009.06908.x. PMC 2746905. PMID 19228196.
- ^ Chan YA, Aristizabal MJ, Lu PY, Luo Z, Hamza A, Kobor MS, Stirling PC, Hieter P (April 2014). "Genome-wide profiling of yeast DNA:RNA hybrid prone sites with DRIP-chip". PLOS Genetics. 10 (4): e1004288. doi: 10.1371/journal.pgen.1004288. PMC 3990523. PMID 24743342.
- ^ Roy D, Yu K, Lieber MR (January 2008). "Mechanism of R-loop formation at immunoglobulin class switch sequences". Molecular and Cellular Biology. 28 (1): 50–60. doi: 10.1128/mcb.01251-07. PMC 2223306. PMID 17954560.
- ^ Ginno PA, Lott PL, Christensen HC, Korf I, Chédin F (March 2012). "R-loop formation is a distinctive characteristic of unmethylated human CpG island promoters". Molecular Cell. 45 (6): 814–25. doi: 10.1016/j.molcel.2012.01.017. PMC 3319272. PMID 22387027.
- ^ D'Souza AD, Belotserkovskii BP, Hanawalt PC (February 2018). "A novel mode for transcription inhibition mediated by PNA-induced R-loops with a model in vitro system". Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms. 1861 (2): 158–166. doi: 10.1016/j.bbagrm.2017.12.008. PMC 5820110. PMID 29357316.
- ^ Castellano-Pozo M, Santos-Pereira JM, Rondón AG, Barroso S, Andújar E, Pérez-Alegre M, García-Muse T, Aguilera A (November 2013). "R loops are linked to histone H3 S10 phosphorylation and chromatin condensation". Molecular Cell. 52 (4): 583–90. doi: 10.1016/j.molcel.2013.10.006. PMID 24211264.
- ^ a b Costantino L, Koshland D (June 2015). "The Yin and Yang of R-loop biology". Current Opinion in Cell Biology. 34: 39–45. doi: 10.1016/j.ceb.2015.04.008. PMC 4522345. PMID 25938907.
- ^ Belotserkovskii BP, Tornaletti S, D'Souza AD, Hanawalt PC (November 2018). "R-loop generation during transcription: Formation, processing and cellular outcomes". DNA Repair. 71: 69–81. doi: 10.1016/j.dnarep.2018.08.009. PMC 6340742. PMID 30190235.
- ^ Sollier J, Cimprich KA (September 2015). "Breaking bad: R-loops and genome integrity". Trends in Cell Biology. 25 (9): 514–22. doi: 10.1016/j.tcb.2015.05.003. PMC 4554970. PMID 26045257.
- ^ a b c Bonnet A, Grosso AR, Elkaoutari A, Coleno E, Presle A, Sridhara SC, Janbon G, Géli V, de Almeida SF, Palancade B (August 2017). "Introns Protect Eukaryotic Genomes from Transcription-Associated Genetic Instability". Molecular Cell. 67 (4): 608–621.e6. doi: 10.1016/j.molcel.2017.07.002. PMID 28757210.