| |
| Names | |
|---|---|
|
IUPAC name
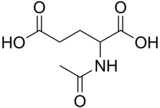
2-Acetamidopentanedioic acid
[1]
| |
| Other names
Acetylglutamic acid[
citation needed]
| |
| Identifiers | |
3D model (
JSmol)
|
|
| 3DMet | |
| Abbreviations |
|
| 1727473 S | |
| ChEBI | |
| ChemSpider | |
| DrugBank | |
| ECHA InfoCard | 100.024.899 |
| EC Number |
|
| KEGG | |
| MeSH | N-acetylglutamate |
PubChem
CID
|
|
| RTECS number |
|
| UNII | |
| |
| |
| Properties | |
| C7H11NO5 | |
| Molar mass | 189.167 g·mol−1 |
| Appearance | White crystals |
| Density | 1 g mL−1 |
| Melting point | 191 to 194 °C (376 to 381 °F; 464 to 467 K) |
| 36 g L−1 | |
| Hazards | |
| Lethal dose or concentration (LD, LC): | |
LD50 (
median dose)
|
>7 g kg−1 (oral, rat) |
| Related compounds | |
Related alkanoic acids
|
|
Related compounds
|
|
Except where otherwise noted, data are given for materials in their
standard state (at 25 °C [77 °F], 100 kPa).
| |
N-Acetylglutamic acid (also referred to as N-acetylglutamate, abbreviated NAG, chemical formula C7H11NO5) [2] is biosynthesized from glutamate and acetylornithine by ornithine acetyltransferase, and from glutamic acid and acetyl-CoA by the enzyme N-acetylglutamate synthase. The reverse reaction, hydrolysis of the acetyl group, is catalyzed by a specific hydrolase. It is the first intermediate involved in the biosynthesis of arginine in prokaryotes and simple eukaryotes and a regulator in the process known as the urea cycle that converts toxic ammonia to urea for excretion from the body in vertebrates.
Discovery
N-Acetylglutamic acid is an extracellular metabolite isolated from the prokaryote Rhizobium trifolii that was characterized using many structure determination techniques such as proton nuclear magnetic resonance (1H NMR) spectroscopy, Fourier-transform infrared spectroscopy, and gas chromatography-mass spectrometry.
In Rhizobium, extracellular build-up of N-acetylglutamic acid is due to metabolism involving nod factor genes on a symbiotic plasmid. When the nod factors are mutated, less N-acetylglutamic acid is produced. [3]
Biosynthesis
Prokaryotes and simple eukaryotes
In prokaryotes and simple eukaryotes, N-acetylglutamic acid can be produced by N-acetylglutamate synthase (NAGS) or ornithine acetyltransferase (OAT).
Ornithine acetyltransferase (OAT) synthesis
OAT synthesizes N-acetylglutamic acid from glutamate and acetylornithine and is the method of choice for production in prokaryotes that have the ability to synthesize the compound ornithine. [4]
N-Acetylglutamate synthase (NAGS) synthesis
N-Acetylglutamate synthase is an enzyme that serves as a replenisher of N-acetylglutamic acid to supplement any N-acetylglutamic acid lost by the cell through mitosis or degradation. NAGS synthesizes N-acetylglutamic acid by catalyzing the addition of an acetyl group from acetyl-coenzyme A to glutamate. In prokaryotes with non-cyclic ornithine production, NAGS is the sole method of N-acetylglutamic acid synthesis and is inhibited by arginine. [4] Acetylation of glutamate is thought to prevent glutamate from being used by proline biosynthesis. [5]
Vertebrates
In contrast to prokaryotes, NAGS in mammals is enhanced by arginine, along with protamines. It is inhibited by N-acetylglutamic acid and its analogues (other N-acetylated compounds). [4]
The brain also contains N-acetylglutamic acid at trace amounts, however no expression of NAGS is found. This suggests that N-acetylglutamic acid is produced by another enzyme in the brain that is yet to be determined. [4]
Biological roles
Vertebrates and mammals
In vertebrae and mammals, N-acetylglutamic acid is the allosteric activator molecule to mitochondrial carbamyl phosphate synthetase I (CPSI) which is the first enzyme in the urea cycle. [6] It triggers the production of the first urea cycle intermediate, carbamyl phosphate. CPSI is inactive when N-acetylglutamic acid is not present. In the liver and small intestines, N-acetylglutamic acid-dependent CPSI produces citrulline, the second intermediate in the urea cycle. Liver cell distribution of N-acetylglutamic acid is highest in the mitochondria at 56% of total N-acetylglutamic acid availability, 24% in the nucleus, and the remaining 20% in the cytosol. Aminoacylase I in liver and kidney cells degrades N-acetylglutamic acid to glutamate and acetate. [7] In contrast, N-acetylglutamic acid is not the allosteric cofactor to carbamyl phosphate synthetase found in the cytoplasm, which is involved in pyrimidine synthesis. [8]
N-acetylglutamic acid concentrations increase when protein consumption increases due to the accumulation of ammonia that must be secreted through the urea cycle, which supports the role of N-acetylglutamic acid as the cofactor for CPSI. Furthermore, N-acetylglutamic acid can be found in many commonly consumed foods such as soy, corn, and coffee, with cocoa powder containing a notably high concentration. [9]
Deficiency in N-acetylglutamic acid in humans is an autosomal recessive disorder that results in blockage of urea production which ultimately increases the concentration of ammonia in the blood ( hyperammonemia). Deficiency can be caused by defects in the NAGS coding gene or by deficiencies in the precursors essential for synthesis. [4]
Bacteria
N-Acetylglutamic acid is the second intermediate in the arginine production pathway in Escherichia coli and is produced via NAGS. [5] In this pathway, N-acetylglutamic acid kinase (NAGK) catalyzes the phosphorylation of the gamma (third) carboxyl group of N-acetylglutamic acid using the phosphate produced by hydrolysis of adenosine triphosphate (ATP). [10]
White clover seedling roots
Rhizobium can form a symbiotic relationship with white clover seedling roots and form colonies. The extracellular N-acetylglutamic acid produced by these bacteria have three morphological effects on the white clover seedling roots: branching of root hairs, swelling of root tips, and increase in the number of cell divisions in undifferentiated cells found on the outer-most cell layer of the root. This suggests that N-acetylglutamic acid is involved in the stimulation of mitosis. The same effects were observed on the strawberry clover, but not in legumes. The effects of N-acetylglutamic acid on the clover species were more potent than the effects from glutamine, glutamate, arginine, or ammonia. [4]
Structure

N-Acetylglutamic acid is composed of two carboxylic acid groups and an amide group protruding from the second carbon. The structure of N-acetylglutamic acid at physiological pH (7.4) has all carboxyl groups deprotonated.
Proton NMR spectroscopy
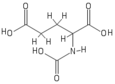

The molecular structure of N-acetylglutamic acid was determined using proton NMR spectroscopy. [3] Proton NMR reveals the presence and functional group location of protons based on chemical shifts recorded on the spectrum. [11]
13C NMR spectroscopy
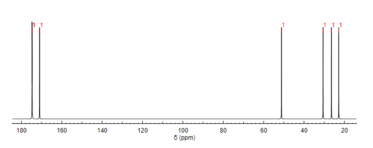
Like proton NMR, carbon-13 (13C) NMR spectroscopy is a method used in molecular structure determination. 13C NMR reveals the types of carbons present in a molecule based on chemical shifts that correspond to certain functional groups. N-Acetylglutamic acid exhibits carbonyl carbons most distinctly due to the three carbonyl-containing substituents. [12]
See also
References
- ^ "N-Acetyl-DL-glutamic acid - Compound Summary". PubChem Compound. USA: National Center for Biotechnology Information. 25 March 2005. Identification. Retrieved 25 June 2012.
- ^ Pubchem. "N-Acetyl L-glutamic acid". pubchem.ncbi.nlm.nih.gov. Retrieved 2018-06-03.
- ^ a b Philip-Hollingsworth S, Hollingsworth RI, Dazzo FB (September 1991). "N-Acetylglutamic acid: an extracellular nod signal of Rhizobium trifolii ANU843 that induces root hair branching and nodule-like primordia in white clover roots". The Journal of Biological Chemistry. 266 (25): 16854–8. doi: 10.1016/S0021-9258(18)55380-1. PMID 1885611.
- ^ a b c d e f Caldovic L, Tuchman M (June 2003). "N-Acetylglutamate and its changing role through evolution". The Biochemical Journal. 372 (Pt 2): 279–90. doi: 10.1042/BJ20030002. PMC 1223426. PMID 12633501.
- ^ a b Caldara M, Dupont G, Leroy F, Goldbeter A, De Vuyst L, Cunin R (March 2008). "Arginine biosynthesis in Escherichia coli: experimental perturbation and mathematical modeling". The Journal of Biological Chemistry. 283 (10): 6347–58. doi: 10.1074/jbc.M705884200. PMID 18165237.
- ^ Auditore, Joseph V.; Wade, Littleton; Olson, Erik J. (November 1966). "Occurrence of N-acetyl-L-glutamic Acid in the Human Brain". Journal of Neurochemistry. 13 (11): 1149–1155. doi: 10.1111/j.1471-4159.1966.tb04272.x. ISSN 0022-3042. PMID 5924663. S2CID 43263361.
- ^ Harper MS, Amanda Shen Z, Barnett JF, Krsmanovic L, Myhre A, Delaney B (November 2009). "N-Acetyl-glutamic acid: evaluation of acute and 28-day repeated dose oral toxicity and genotoxicity". Food and Chemical Toxicology. 47 (11): 2723–9. doi: 10.1016/j.fct.2009.07.036. PMID 19654033.
- ^ Pelley JW (2007). "Chapter 14: Purine, Pyrimidine, and Single-Carbon Metabolism". Elsevier's Integrated Biochemistry. Elsevier. pp. 117–122. doi: 10.1016/b978-0-323-03410-4.50020-1. ISBN 978-0-323-03410-4.
- ^ Hession AO, Esrey EG, Croes RA, Maxwell CA (October 2008). "N-Acetylglutamate and N-acetylaspartate in soybeans (Glycine max L.), maize (Zea mays L.), [corrected] and other foodstuffs". Journal of Agricultural and Food Chemistry. 56 (19): 9121–6. doi: 10.1021/jf801523c. PMID 18781757.
-
^ Gil Ortiz F, Ramón Maiques S, Fita I, Rubio V (August 2003). "The course of phosphorus in the reaction of N-acetyl-L-glutamate kinase, determined from the structures of crystalline complexes, including a complex with an AlF−
4 transition state mimic". Journal of Molecular Biology. 331 (1): 231–44. doi: 10.1016/S0022-2836(03)00716-2. PMID 12875848. - ^ "Predict 1H proton NMR spectra". www.nmrdb.org. Retrieved 2018-06-03.
- ^ "Predict 13C carbon NMR spectra". www.nmrdb.org. Retrieved 2018-06-03.