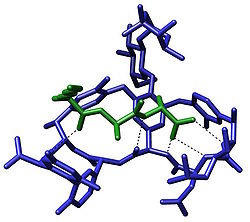


The term molecular recognition refers to the specific interaction between two or more molecules through noncovalent bonding such as hydrogen bonding, metal coordination, hydrophobic forces, [3] [4] van der Waals forces, π-π interactions, halogen bonding, or resonant interaction [5] effects. In addition to these direct interactions, solvents can play a dominant indirect role in driving molecular recognition in solution. [6] [7] The host and guest involved in molecular recognition exhibit molecular complementarity. Exceptions are molecular containers, [8] [9] including, e.g., nanotubes, in which portals essentially control selectivity. [10] [11] [12] [13]
Biological systems
Molecular recognition plays an important role in biological systems and is observed in between receptor-ligand, [14] [15] antigen- antibody, DNA- protein, sugar- lectin, RNA- ribosome, etc. An important example of molecular recognition is the antibiotic vancomycin that selectively binds with the peptides with terminal D-alanyl-D-alanine in bacterial cells through five hydrogen bonds. The vancomycin is lethal to the bacteria since once it has bound to these particular peptides they are unable to be used to construct the bacteria's cell wall.[ citation needed]
Synthetic molecular recognition
Recent work suggests that molecular recognition elements can be synthetically produced at the nano-scale, [16] circumventing the need for naturally occurring molecular recognition elements for the development of sensing tools for small molecules. Bio-mimetic polymers such as molecular imprinted polymers [17] and peptoids can be used to recognize larger biological targets such as proteins [18] and the conjugation of polymers to synthetic fluorescent nanomaterials can generate synthetic macromolecular structures that serve as synthetic antibodies for optical protein recognition and detection. [19] [20]
Supramolecular systems
Chemists have demonstrated that many artificial supramolecular systems can be designed that exhibit molecular recognition. [21] One of the earliest examples of such a system are crown ethers which are capable of selectively binding specific cations. However, a number of artificial systems have since been established.
Static vs. dynamic
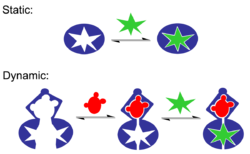
Molecular recognition can be subdivided into static molecular recognition and dynamic molecular recognition. Static molecular recognition is likened to the interaction between a key and a keyhole; it is a 1:1 type complexation reaction between a host molecule and a guest molecule to form a host–guest complex. To achieve advanced static molecular recognition, it is necessary to make recognition sites that are specific for guest molecules.
In the case of dynamic molecular recognition the binding of the first guest to the first binding site of a host affects the association constant of a second guest with a second binding site. leading to cooperativity of binding. [22] In the case of positive allosteric systems the binding of the first guest increases the association constant of the second guest. While for negative allosteric systems the binding of the first guest decreases the association constant with the second. The dynamic nature of this type of molecular recognition is particularly important since it provides a mechanism to regulate binding in biological systems. Dynamic molecular recognition may enhance the ability to discriminate between several competing targets via the conformational proofreading mechanism. Dynamic molecular recognition is also being studied for application in highly functional chemical sensors and molecular devices [23]
Complexity
A recent study based on molecular simulations and compliance constants describes molecular recognition as a phenomenon of organisation. Even for small molecules like carbohydrates, the recognition process can not be predicted or designed even assuming that each individual hydrogen bond's strength is exactly known. [24] However, as Mobley et al. [25] concluded, the accurate prediction of the molecular recognition events needs to go beyond the static snapshot of a single frame between the guest and the host. Entropies are key contributors to binding thermodynamics and need to be accounted for in order to predict more accurately the recognition process. [26] Entropies are rarely observable in single bound structures (static snapshot). In proteins, highly-specific recognition can be achieved by evolutionary fine-tuning of chemical interactions, conformational changes, and entropy contributions. [27]
Intragenic complementation
Jehle [28] pointed out that, when immersed in a liquid and intermingled with other molecules, charge fluctuation forces favor the association of identical molecules as nearest neighbors. In accord with this principle, the multiple copies of a polypeptide encoded by a gene often undergo molecular recognition with each other to form an ordered multi-polypeptide protein structure. When such a protein is formed from polypeptides produced by two different mutant alleles of a particular gene, the protein composed of a mixture of polypeptides may exhibit greater functional activity than the multi-polypeptide protein formed by each of the mutants alone. In such a case, the phenomenon is referred to as intragenic complementation.
Intragenic complementation (also called inter-allelic complementation) has been demonstrated in many different genes in a variety of organisms. [29] Crick and Orgel [30] analyzed of the results of such studies and came to the conclusion that intragenic complementation, in general, arises from the interaction of differently defective polypeptide monomers when they form an ordered aggregate they called a “multimer.”
See also
- Journal of Molecular Recognition
- SAMPL Challenge
- Noncovalent interactions
- Supramolecular chemistry
- Allostery
- Cooperativity
- Molecular assembler
References
- ^ Knox JR, Pratt RF (July 1990). "Different modes of vancomycin and D-alanyl-D-alanine peptidase binding to cell wall peptide and a possible role for the vancomycin resistance protein" (Free full text). Antimicrobial Agents and Chemotherapy. 34 (7): 1342–1347. doi: 10.1128/AAC.34.7.1342. PMC 175978. PMID 2386365.
- ^ Bielawski C, Chen Y, Zhang P, Prest P, Moore JS (1998). "A modular approach to constructing multi-site receptors for isophthalic acid". Chemical Communications (12): 1313–4. doi: 10.1039/a707262g.
- ^ Lockett MR, Lange H, Breiten B, Heroux A, Sherman W, Rappoport D, et al. (July 2013). "The binding of benzoarylsulfonamide ligands to human carbonic anhydrase is insensitive to formal fluorination of the ligand". Angewandte Chemie. 52 (30): 7714–7717. doi: 10.1002/anie.201301813. PMID 23788494. S2CID 1543705.
- ^ Breiten B, Lockett MR, Sherman W, Fujita S, Al-Sayah M, Lange H, et al. (October 2013). "Water networks contribute to enthalpy/entropy compensation in protein-ligand binding". Journal of the American Chemical Society. 135 (41): 15579–15584. CiteSeerX 10.1.1.646.8648. doi: 10.1021/ja4075776. PMID 24044696. S2CID 17554787.
- ^ Cosic I (December 1994). "Macromolecular bioactivity: is it resonant interaction between macromolecules?--Theory and applications". IEEE Transactions on Bio-Medical Engineering. 41 (12): 1101–1114. doi: 10.1109/10.335859. PMID 7851912. S2CID 23892544.
- ^ Baron R, Setny P, McCammon JA (September 2010). "Water in cavity-ligand recognition". Journal of the American Chemical Society. 132 (34): 12091–12097. doi: 10.1021/ja1050082. PMC 2933114. PMID 20695475.
- ^ Baron R, McCammon JA (2013). "Molecular recognition and ligand association". Annual Review of Physical Chemistry. 64: 151–175. Bibcode: 2013ARPC...64..151B. doi: 10.1146/annurev-physchem-040412-110047. PMID 23473376.
- ^ Cram DJ, Cram JM (1997). Container molecules and their guests. Cambridge: Royal Society of Chemistry. ISBN 978-0-85186-972-8.
- ^ Brotin T, Dutasta JP (January 2009). "Cryptophanes and their complexes--present and future". Chemical Reviews. 109 (1): 88–130. doi: 10.1021/cr0680437. PMID 19086781.
- ^ Lehn JM (1995). Supramolecular Chemistry. Weinheim: Wiley-VCH. ISBN 978-3-527-29312-4. OCLC 315928178.[ page needed]
- ^ Gellman SH (August 1997). "Introduction: Molecular Recognition". Chemical Reviews. 97 (5): 1231–1232. doi: 10.1021/cr970328j. PMID 11851448.
- ^ Chatterji D (2016). Basics of Molecular Recognition. Boca Raton, FL: CRC Press. ISBN 978-1-4822-1968-5.
- ^ Rotello V, Thayumanavan S, eds. (2008). Molecular recognition and polymers : control of polymer structure and self-assembly. Hoboken, N.J.: Wiley. ISBN 978-0-470-27738-6.
- ^ Lockett MR, Lange H, Breiten B, Heroux A, Sherman W, Rappoport D, et al. (July 2013). "The binding of benzoarylsulfonamide ligands to human carbonic anhydrase is insensitive to formal fluorination of the ligand". Angewandte Chemie. 52 (30): 7714–7717. doi: 10.1002/anie.201301813. PMID 23788494. S2CID 1543705.
- ^ Breiten B, Lockett MR, Sherman W, Fujita S, Al-Sayah M, Lange H, et al. (October 2013). "Water networks contribute to enthalpy/entropy compensation in protein-ligand binding". Journal of the American Chemical Society. 135 (41): 15579–15584. CiteSeerX 10.1.1.646.8648. doi: 10.1021/ja4075776. PMID 24044696. S2CID 17554787.
- ^ Zhang J, Landry MP, Barone PW, Kim JH, Lin S, Ulissi ZW, et al. (December 2013). "Molecular recognition using corona phase complexes made of synthetic polymers adsorbed on carbon nanotubes". Nature Nanotechnology. 8 (12): 959–968. Bibcode: 2013NatNa...8..959Z. doi: 10.1038/nnano.2013.236. PMC 5051352. PMID 24270641.
- ^ Sullivan MV, Dennison SR, Archontis G, Reddy SM, Hayes JM (July 2019). "Toward Rational Design of Selective Molecularly Imprinted Polymers (MIPs) for Proteins: Computational and Experimental Studies of Acrylamide Based Polymers for Myoglobin" (PDF). The Journal of Physical Chemistry B. 123 (26): 5432–5443. doi: 10.1021/acs.jpcb.9b03091. PMID 31150581. S2CID 172137800.
- ^ Mannige RV, Haxton TK, Proulx C, Robertson EJ, Battigelli A, Butterfoss GL, et al. (October 2015). "Peptoid nanosheets exhibit a new secondary-structure motif". Nature. 526 (7573): 415–420. Bibcode: 2015Natur.526..415M. doi: 10.1038/nature15363. PMID 26444241. S2CID 205245623.
- ^ Sullivan MV, Stockburn WJ, Hawes PC, Mercer T, Reddy SM (February 2021). "Green synthesis as a simple and rapid route to protein modified magnetic nanoparticles for use in the development of a fluorometric molecularly imprinted polymer-based assay for detection of myoglobin". Nanotechnology. 32 (9): 095502. Bibcode: 2021Nanot..32i5502S. doi: 10.1088/1361-6528/abce2d. PMC 8314874. PMID 33242844.
-
^ Beyene AG, Demirer GS, Landry MP (2009-01-01).
Nanoparticle-Templated Molecular Recognition Platforms for Detection of Biological Analytes. Vol. 8. John Wiley & Sons, Inc. pp. 197–223.
doi:
10.1002/cpch.10.
ISBN
9780470559277.
PMC
10539024.
PMID
27622569.
S2CID
5249925.
{{ cite book}}:|journal=ignored ( help) - ^ Biedermann F, Schneider HJ (May 2016). "Experimental Binding Energies in Supramolecular Complexes". Chemical Reviews. 116 (9): 5216–5300. doi: 10.1021/acs.chemrev.5b00583. PMID 27136957.
- ^ Shinkai S, Ikeda M, Sugasaki A, Takeuchi M (June 2001). "Positive allosteric systems designed on dynamic supramolecular scaffolds: toward switching and amplification of guest affinity and selectivity". Accounts of Chemical Research. 34 (6): 494–503. doi: 10.1021/ar000177y. PMID 11412086.
- ^ McBride JM, Eckmann JP, Tlusty T (November 2022). "General Theory of Specific Binding: Insights from a Genetic-Mechano-Chemical Protein Model". Molecular Biology and Evolution. 39 (11). doi: 10.1093/molbev/msac217. PMC 9641994. PMID 36208205.
- ^ Grunenberg J (June 2011). "Complexity in molecular recognition". Physical Chemistry Chemical Physics. 13 (21): 10136–10146. Bibcode: 2011PCCP...1310136G. doi: 10.1039/C1CP20097F. PMID 21503359.
- ^ Mobley DL, Dill KA (April 2009). "Binding of small-molecule ligands to proteins: "what you see" is not always "what you get"". Structure. 17 (4): 489–498. doi: 10.1016/j.str.2009.02.010. PMC 2756098. PMID 19368882.
- ^ Schmidtchen FP (October 2010). "Hosting anions. The energetic perspective". Chemical Society Reviews. 39 (10): 3916–3935. doi: 10.1039/C0CS00038H. PMID 20820595.
- ^ McBride JM, Eckmann JP, Tlusty T (November 2022). Echave J (ed.). "General Theory of Specific Binding: Insights from a Genetic-Mechano-Chemical Protein Model". Molecular Biology and Evolution. 39 (11): msac217. doi: 10.1093/molbev/msac217. PMC 9641994. PMID 36208205.
- ^ Jehle H (September 1963). "Intermolecular forces and biological specificity". Proceedings of the National Academy of Sciences of the United States of America. 50 (3): 516–24. Bibcode: 1963PNAS...50..516J. doi: 10.1073/pnas.50.3.516. PMC 221211. PMID 16578546.
- ^ Bernstein H, Edgar RS, Denhardt GH (June 1965). "Intragenic complementation among temperature sensitive mutants of bacteriophage T4D". Genetics. 51 (6): 987–1002. doi: 10.1093/genetics/51.6.987. PMC 1210828. PMID 14337770.
- ^ Crick FH, Orgel LE (January 1964). "The theory of inter-allelic complementation". Journal of Molecular Biology. 8: 161–5. doi: 10.1016/s0022-2836(64)80156-x. PMID 14149958.
External links
- "Special Issue on Molecular Recognition". International Journal of Molecular Sciences.