| |
| Names | |
|---|---|
|
IUPAC name
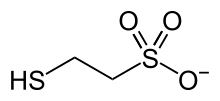
2-Sulfanylethanesulfonate
| |
|
Systematic IUPAC name
2-Sulfanylethanesulfonate | |
| Other names
2-mercaptoethylsulfonate; 2-mercaptoethanesulfonate; coenzyme M anion; H-S-CoM; AC1L1HCY; 2-sulfanylethane-1-sulfonate; CTK8A8912
| |
| Identifiers | |
| |
3D model (
JSmol)
|
|
| ChEBI | |
| ChemSpider | |
PubChem
CID
|
|
| UNII |
|
| |
| |
| Properties | |
| C2H5O3S2 | |
| Molar mass | 141.18 g·mol−1 |
Except where otherwise noted, data are given for materials in their
standard state (at 25 °C [77 °F], 100 kPa).
| |
Coenzyme M is a
coenzyme required for methyl-transfer reactions in the
metabolism of
archaeal
methanogens,
[1]
[2] and in the metabolism of other substrates in
bacteria.
[3] It is also a necessary cofactor in the metabolic pathway of alkene-oxidizing bacteria. CoM helps eliminate the toxic epoxides formed from the oxidation of alkenes such as propylene.
[4] The structure of this coenzyme was discovered by CD Taylor and RS Wolfe in 1974 while they were studying methanogenesis, the process by which carbon dioxide is transformed into methane in some anaerobic bacteria.
[5] The coenzyme is an
anion with the formula HSCH
2CH
2SO−
3. It is named 2-mercaptoethanesulfonate and abbreviated HS–CoM. The
cation is unimportant, but the
sodium salt is most available. Mercaptoethanesulfonate contains both a
thiol, which is the main site of reactivity, and a
sulfonate group, which confers solubility in aqueous media.
Biochemical role
Methanogenesis
The coenzyme is the C1 donor in
methanogenesis. It is converted to methyl-coenzyme M thioether, the
thioether CH
3SCH
2CH
2SO−
3, in the penultimate step to
methane formation.
[6] Methyl-coenzyme M reacts with
coenzyme B, 7-thioheptanoylthreoninephosphate, to give a heterodisulfide, releasing methane:
- CH3–S–CoM + HS–CoB → CH4 + CoB–S–S–CoM
This induction is catalyzed by the enzyme methyl-coenzyme M reductase, which restricts cofactor F430 as the prosthetic group.
Alkene metabolism
Coenzyme M is also used to make acetoacetate from CO2 and propylene or ethylene in aerobic bacteria. Specifically, in bacteria that oxidize alkenes into epoxides. After the propylene (or other alkene) undergoes epoxidation and becomes epoxypropane it becomes electrophilic and toxic. These epoxides react with DNA and proteins, affecting cell function. Alkene-oxidizing bacteria like Xanthobacter autotrophicus [4] use a metabolic pathway in which CoM is conjugated with an aliphatic epoxide. This step creates a nucleophilic compound which can react with CO2. The eventual carboxylation produces acetoacetate, breaking down the propylene. [4]
See also
- Mesna – a cancer chemotherapy adjuvant with the same structure
References
- ^ Balch WE, Wolfe RS (1979). "Specificity and biological distribution of coenzyme M (2-mercaptoethanesulfonic acid)". J. Bacteriol. 137 (1): 256–63. doi: 10.1128/JB.137.1.256-263.1979. PMC 218444. PMID 104960.
-
^ Taylor CD, Wolfe RS (10 August 1974).
"Structure and methylation of coenzyme M(HSCH
2CH
2SO
3)". J. Biol. Chem. 249 (15): 4879–85. doi: 10.1016/S0021-9258(19)42403-4. PMID 4367810. - ^ Partovi, Sarah E.; Mus, Florence; Gutknecht, Andrew E.; Martinez, Hunter A.; Tripet, Brian P.; Lange, Bernd Markus; DuBois, Jennifer L.; Peters, John W. (2018-04-06). "Coenzyme M biosynthesis in bacteria involves phosphate elimination by a functionally distinct member of the aspartase/fumarase superfamily". The Journal of Biological Chemistry. 293 (14): 5236–5246. doi: 10.1074/jbc.RA117.001234. ISSN 1083-351X. PMC 5892593. PMID 29414784.
- ^ a b c Krishnakumar, Arathi M.; Sliwa, Darius; Endrizzi, James A.; Boyd, Eric S.; Ensign, Scott A.; Peters, John W. (September 2008). "Getting a Handle on the Role of Coenzyme M in Alkene Metabolism". Microbiology and Molecular Biology Reviews. 72 (3): 445–456. doi: 10.1128/MMBR.00005-08. ISSN 1092-2172. PMC 2546864. PMID 18772284.
- ^ Parry, Ronald J. (1999-01-01), Barton, Sir Derek; Nakanishi, Koji; Meth-Cohn, Otto (eds.), "1.29 - Biosynthesis of Sulfur-containing Natural Products", Comprehensive Natural Products Chemistry, Oxford: Pergamon, pp. 825–863, doi: 10.1016/b978-0-08-091283-7.00031-x, ISBN 978-0-08-091283-7, retrieved 2022-05-10
- ^ Thauer, Rudolf K. (1998-09-01). "Biochemistry of methanogenesis: a tribute to Marjory Stephenson:1998 Marjory Stephenson Prize Lecture". Microbiology. 144 (9): 2377–2406. doi: 10.1099/00221287-144-9-2377. ISSN 1350-0872. PMID 9782487.