In organic chemistry, polyketides are a class of natural products derived from a precursor molecule consisting of a chain of alternating ketone (>C=O, or its reduced forms) and methylene (>CH2) groups: [−C(=O)−CH2−]n. [1] First studied in the early 20th century, discovery, biosynthesis, and application of polyketides has evolved. It is a large and diverse group of secondary metabolites caused by its complex biosynthesis which resembles that of fatty acid synthesis. Because of this diversity, polyketides can have various medicinal, agricultural, and industrial applications. Many polyketides are medicinal or exhibit acute toxicity. Biotechnology has enabled discovery of more naturally-occurring polyketides and evolution of new polyketides with novel or improved bioactivity.
History
Naturally produced polyketides by various plants and organisms have been used by humans since before studies on them began in the 19th and 20th century. In 1893, J. Norman Collie synthesized detectable amounts of orcinol by heating dehydracetic acid with barium hydroxide causing the pyrone ring to open into a triketide. [2] Further studies in 1903 by Collie on the triketone polyketide intermediate noted the condensation occurring amongst compounds with multiple keten groups coining the term polyketides. [3]
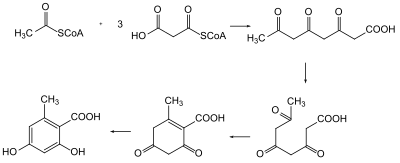
It wasn't until 1955 that the biosynthesis of polyketides were understood. [4] Arthur Birch used radioisotope labeling of carbon in acetate to trace the biosynthesis of 2-hydroxy-6-methylbenzoic acid in Penicillium patulum and demonstrate the head-to-tail linkage of acetic acids to form the polyketide. [5] In the 1980s and 1990s, advancements in genetics allowed for isolation of the genes associated to polyketides to understand the biosynthesis. [4]
Discovery
Polyketides can be produced in bacteria, fungi, plants, and certain marine organisms. [6] Earlier discovery of naturally occurring polyketides involved the isolation of the compounds being produced by the specific organism using organic chemistry purification methods based on bioactivity screens. [7] Later technology allowed for the isolation of the genes and heterologous expression of the genes to understand the biosynthesis. [8] In addition, further advancements in biotechnology have allowed for the use of metagenomics and genome mining to find new polyketides using similar enzymes to known polyketides. [9]
Biosynthesis
Polyketides are synthesized by multienzyme polypeptides that resemble eukaryotic fatty acid synthase but are often much larger. [4] They include acyl-carrier domains plus an assortment of enzymatic units that can function in an iterative fashion, repeating the same elongation/modification steps (as in fatty acid synthesis), or in a sequential fashion so as to generate more heterogeneous types of polyketides. [10]

Polyketide synthase
Polyketides are produced by polyketide synthases (PKSs). The core biosynthesis involves stepwise condensation of a starter unit (typically acetyl-CoA or propionyl-CoA) with an extender unit (either malonyl-CoA or methylmalonyl-CoA). The condensation reaction is accompanied by the decarboxylation of the extender unit, yielding a beta-keto functional group and releasing a carbon dioxide. [10] The first condensation yields an acetoacetyl group, a diketide. Subsequent condensations yield triketides, tetraketide, etc. [11] Other starter units attached to a coezyme A include isobutyrate, cyclohexanecarboxylate, malonate, and benzoate. [12]
PKSs are multi-domain enzymes or enzyme complex consisting of various domains. The polyketide chains produced by a minimal polyketide synthase (consisting of a acyltransferase and ketosynthase for the stepwise condensation of the starter unit and extender units) are almost invariably modified. [13] Each polyketide synthases is unique to each polyketide chain because they contain different combinations of domains that reduce the carbonyl group to a hydroxyl (via a ketoreductase), an olefin (via a dehydratase), or a methylene (via an enoylreductase). [14]
Termination of the polyketide scaffold biosynthesis can also vary. It is sometimes accompanied by a thioesterase that releases the polyketide via hydrating the thioester linkage (as in fatty acid synthesis) creating a linear polyketide scaffold. However, if water is not able to reach the active site, the hydrating reaction will not occur and an intramolecular reaction is more probable creating a macrocyclic polyketide. Another possibility is spontaneous hydrolysis without the aid of a thioesterase. [15]
Post-tailoring enzymes
Further possible modifications to the polyketide scaffolds can be made. This can include glycosylation via a glucosyltransferase or oxidation via a monooxygenase. [16] Similarly, cyclization and aromatization can be introduced via a cyclase, sometimes proceeded by the enol tautomers of the polyketide. [17] These enzymes are not part of the domains of the polyketide synthase. Instead, they are found in gene clusters in the genome close to the polyketide synthase genes. [18]
Classification
Polyketides are a structurally diverse family. [19] There are various subclasses of polyketides including: aromatics, macrolactones/ macrolides, decalin ring containing, polyether, and polyenes. [15]
Polyketide synthases are also broadly divided into three classes: Type I PKSs (multimodular megasynthases that are non-iterative, often producing macrocodes, polyethers, and polyenes), Type II PKSs (dissociated enzymes with iterative action, often producing aromatics), and Type III PKSs ( chalcone synthase-like, producing small aromatic molecules). [20]
In addition to these subclasses, there also exist polyketides that are hybridized with nonribosomal peptides (Hybrid NRP-PK and PK-NRP). Since nonribosomal peptide assembly lines use carrier proteins similar to those use in polyketide synthases, convergence of the two systems evolved to form hybrids, resulting in polypeptides with nitrogen in the skeletal structure and complex function groups similar to those found in amino acids. [21]
Applications
Polyketide antibiotics, [22] antifungals, [23] cytostatics, [24] anticholesteremic, [25] antiparasitics, [23] coccidiostats, animal growth promoters and natural insecticides [26] are in commercial use.
Medicinal
There are more than 10,000 known polyketides, 1% of which are known to have potential for drug activity. [27] Polyketides comprise 20% of the top-selling pharmaceuticals with combined worldwide revenues of over USD 18 billion per year. [28]
|
|

|
|
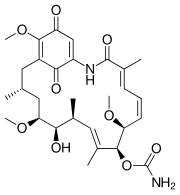
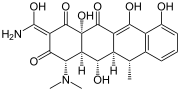
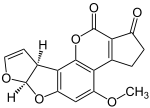
| Geldanamycin, an antibiotic. | Doxycycline, an antibiotic. | Erythromycin, an antibiotic. | Aflatoxin B1 known carcinogenic compound. |
Examples
-
Macrolides
- Pikromycin, the first isolated macrolide (1951 [29])
- The antibiotics erythromycin A, clarithromycin, and azithromycin
- The antihelminthics ivermectin
-
Ansamycins
- The antitumor agents geldanamycin and macbecin,
- The antibiotic rifamycin
-
Polyenes
- The antifungals amphotericin, nystatin and pimaricin
-
Polyethers
- The antibiotic monensin
-
Tetracyclines
- The antibiotic agent doxycycline
- Acetogenins
- Others
- The immunosuppressants tacrolimus (FK506) (a calcineurin inhibitor) and sirolimus (rapamycin) (a mTOR inhibitor)
- Radicicol and the pochonin family ( HSP90 inhibitors)
- The cholesterol lowering agent lovastatin
- Discodermolide
- Aflatoxin
- Usnic acid
- Anthracimycin
- Anthramycin
- Olivetolic acid (intermediate in cannabinoid pathways) [30]
Agricultural
Polyketides can be used for crop protection as pesticides. [31]
Examples
- Pesticides
- spinosad or spinosyn (an insecticide)
- avermectin
- polynactins
- tetramycin
Industrial
Polyketides can be used for industrial purposes, such as pigmentation [32] and dietary flavonoids. [33]
Examples
- Pigments
- azaphilones
- hydroxyanthraquinones
- naphthoquinones
- Flavonoids
Biotechnology
Protein engineering has opened avenues for creating polyketides not found in nature. For example, the modular nature of PKSs allows for domains to be replaced, added or deleted. Introducing diversity in assembly lines enables the discovery of new polyketides with increased bioactivity or new bioactivity. [21]
Furthermore, the use of genome mining allows for discovery of new natural polyketides and their assembly lines. [9]
See also
References
- ^ IUPAC, Compendium of Chemical Terminology, 2nd ed. (the "Gold Book") (1997). Online corrected version: (2006–) " Polyketides". doi: 10.1351/goldbook.P04734
- ^ Collie N, Myers WS (1893). "VII.—The formation of orcinol and other condensation products from dehydracetic acid". Journal of the Chemical Society, Transactions. 63: 122–128. doi: 10.1039/CT8936300122. ISSN 0368-1645.
- ^ Collie JN (1907). "CLXXI.—Derivatives of the multiple keten group". Journal of the Chemical Society, Transactions. 91: 1806–1813. doi: 10.1039/CT9079101806. ISSN 0368-1645.
- ^ a b c Smith S, Tsai SC (October 2007). "The type I fatty acid and polyketide synthases: a tale of two megasynthases". Natural Product Reports. 24 (5): 1041–1072. doi: 10.1039/B603600G. PMC 2263081. PMID 17898897.
- ^ Birch AJ, Massy-Westropp RA, Moye CJ (1955). "Studies in relation to biosynthesis. VII. 2-Hydroxy-6-methylbenzoic acid in Penicillium griseofulvum Dierckx". Australian Journal of Chemistry. 8 (4): 539–544. doi: 10.1071/ch9550539. ISSN 1445-0038.
- ^ Lane AL, Moore BS (February 2011). "A sea of biosynthesis: marine natural products meet the molecular age". Natural Product Reports. 28 (2): 411–428. doi: 10.1039/C0NP90032J. PMC 3101795. PMID 21170424.
- ^ Johnston C, Ibrahim A, Magarvey N (2012-08-01). "Informatic strategies for the discovery of polyketides and nonribosomal peptides". MedChemComm. 3 (8): 932–937. doi: 10.1039/C2MD20120H. ISSN 2040-2511.
- ^ Pfeifer BA, Khosla C (March 2001). "Biosynthesis of polyketides in heterologous hosts". Microbiology and Molecular Biology Reviews. 65 (1): 106–118. doi: 10.1128/MMBR.65.1.106-118.2001. PMC 99020. PMID 11238987.
- ^ a b Gomes ES, Schuch V, de Macedo Lemos EG (December 2013). "Biotechnology of polyketides: new breath of life for the novel antibiotic genetic pathways discovery through metagenomics". Brazilian Journal of Microbiology. 44 (4): 1007–1034. doi: 10.1590/s1517-83822013000400002. PMC 3958165. PMID 24688489.
- ^ a b Voet D, Voet JG, Pratt CW (2013). Fundamentals of Biochemistry: Life at the Molecular Level (4th ed.). John Wiley & Sons. p. 688. ISBN 9780470547847.
- ^ Staunton J, Weissman KJ (August 2001). "Polyketide biosynthesis: a millennium review". Natural Product Reports. 18 (4): 380–416. doi: 10.1039/a909079g. PMID 11548049.
- ^ Moore BS, Hertweck C (February 2002). "Biosynthesis and attachment of novel bacterial polyketide synthase starter units". Natural Product Reports. 19 (1): 70–99. doi: 10.1039/B003939J. PMID 11902441.
- ^ Wang J, Zhang R, Chen X, et al. (May 2020). "Biosynthesis of aromatic polyketides in microorganisms using type II polyketide synthases". Microbial Cell Factories. 19 (1): 110. doi: 10.1186/s12934-020-01367-4. PMC 7247197. PMID 32448179.
- ^ Moretto L, Heylen R, Holroyd N, et al. (February 2019). "Modular type I polyketide synthase acyl carrier protein domains share a common N-terminally extended fold". Scientific Reports. 9 (1): 2325. Bibcode: 2019NatSR...9.2325M. doi: 10.1038/s41598-019-38747-9. PMC 6382882. PMID 30787330.
- ^ a b Walsh C, Tang Y (2017). Natural product biosynthesis. Royal Society of Chemistry. ISBN 978-1-78801-131-0. OCLC 985609285.
- ^ Risdian C, Mozef T, Wink J (May 2019). "Biosynthesis of Polyketides in Streptomyces". Microorganisms. 7 (5): 124. doi: 10.3390/microorganisms7050124. PMC 6560455. PMID 31064143.
- ^ Robinson JA (May 1991). "Polyketide synthase complexes: their structure and function in antibiotic biosynthesis". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 332 (1263): 107–114. Bibcode: 1991RSPTB.332..107R. doi: 10.1098/rstb.1991.0038. PMID 1678529.
- ^ Noar RD, Daub ME (2016-07-07). "Bioinformatics Prediction of Polyketide Synthase Gene Clusters from Mycosphaerella fijiensis". PLOS ONE. 11 (7): e0158471. Bibcode: 2016PLoSO..1158471N. doi: 10.1371/journal.pone.0158471. PMC 4936691. PMID 27388157.
- ^ Katz L (November 1997). "Manipulation of Modular Polyketide Synthases". Chemical Reviews. 97 (7): 2557–2576. doi: 10.1021/cr960025+. PMID 11851471.
- ^ Shen B (April 2003). "Polyketide biosynthesis beyond the type I, II and III polyketide synthase paradigms". Current Opinion in Chemical Biology. 7 (2): 285–295. doi: 10.1016/S1367-5931(03)00020-6. PMID 12714063.
- ^ a b Nivina A, Yuet KP, Hsu J, Khosla C (December 2019). "Evolution and Diversity of Assembly-Line Polyketide Synthases". Chemical Reviews. 119 (24): 12524–12547. doi: 10.1021/acs.chemrev.9b00525. PMC 6935866. PMID 31838842.
- ^ "5.13E: Polyketide Antibiotics". Biology LibreTexts. 2017-05-09. Retrieved 2021-07-05.
- ^ a b Ross C, Opel V, Scherlach K, Hertweck C (December 2014). "Biosynthesis of antifungal and antibacterial polyketides by Burkholderia gladioli in coculture with Rhizopus microsporus". Mycoses. 57 (Suppl 3): 48–55. doi: 10.1111/myc.12246. PMID 25250879.
- ^ Jiang L, Pu H, Xiang J, et al. (2018). "Huanglongmycin A-C, Cytotoxic Polyketides Biosynthesized by a Putative Type II Polyketide Synthase From Streptomyces sp. CB09001". Frontiers in Chemistry. 6: 254. Bibcode: 2018FrCh....6..254J. doi: 10.3389/fchem.2018.00254. PMC 6036704. PMID 30013965.
- ^ Chan YA, Podevels AM, Kevany BM, Thomas MG (January 2009). "Biosynthesis of polyketide synthase extender units". Natural Product Reports. 26 (1): 90–114. doi: 10.1039/b801658p. PMC 2766543. PMID 19374124.
- ^ Kim HJ, Choi SH, Jeon BS, et al. (December 2014). "Chemoenzymatic synthesis of spinosyn A". Angewandte Chemie. 53 (49): 13553–13557. doi: 10.1002/anie.201407806. PMC 4266379. PMID 25287333.
- ^ Baerson SR, Rimando AM (2007-01-11). "A Plethora of Polyketides: Structures, Biological Activities, and Enzymes". In Rimando AM, Baerson SR (eds.). Polyketides: Biosynthesis, Biological Activity, and Genetic Engineering. ACS Symposium Series. Vol. 955. Washington, DC: American Chemical Society. pp. 2–14. doi: 10.1021/bk-2007-0955.ch001. ISBN 978-0-8412-3978-4.
- ^ Weissman K, Leadlay B (2005). "Combinatorial biosynthesis of reduced polyketides". Nature Reviews Microbiology. 3 (12): 925–936. doi: 10.1038/nrmicro1287. PMID 16322741. S2CID 205496204.
- ^ Brockmann H, Henkel W (1951). "Pikromycin, ein bitter schmeckendes Antibioticum aus Actinomyceten" [Pikromycin, a bitter tasting antibiotic from an actinomycete]. Chem. Ber. (in German). 84 (3): 284–288. doi: 10.1002/cber.19510840306.
- ^ Gagne SJ, Stout JM, Liu E, et al. (July 2012). "Identification of olivetolic acid cyclase from Cannabis sativa reveals a unique catalytic route to plant polyketides". Proceedings of the National Academy of Sciences of the United States of America. 109 (31): 12811–12816. Bibcode: 2012PNAS..10912811G. doi: 10.1073/pnas.1200330109. PMC 3411943. PMID 22802619.
- ^ Li S, Yang B, Tan GY, et al. (June 2021). "Polyketide pesticides from actinomycetes". Current Opinion in Biotechnology. Chemical Biotechnology ● Pharmaceutical Biotechnology. 69: 299–307. doi: 10.1016/j.copbio.2021.05.006. PMID 34102376. S2CID 235378697.
- ^ Caro Y, Venkatachalam M, Lebeau J, et al. (2016). "Pigments and Colorants from Filamentous Fungi". In Merillon JM, Ramawat KG (eds.). Fungal Metabolites. Reference Series in Phytochemistry. Cham: Springer International Publishing. pp. 1–70. doi: 10.1007/978-3-319-19456-1_26-1. ISBN 978-3-319-19456-1.
- ^ Tauchen J, Huml L, Rimpelova S, Jurášek M (August 2020). "Flavonoids and Related Members of the Aromatic Polyketide Group in Human Health and Disease: Do They Really Work?". Molecules. 25 (17): 3846. doi: 10.3390/molecules25173846. PMC 7504053. PMID 32847100.