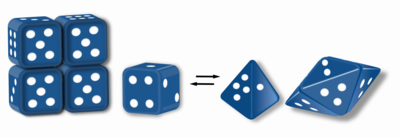
Morpheeins are proteins that can form two or more different homo- oligomers (morpheein forms), but must come apart and change shape to convert between forms. The alternate shape may reassemble to a different oligomer. The shape of the subunit dictates which oligomer is formed. [1] [2] Each oligomer has a finite number of subunits ( stoichiometry). Morpheeins can interconvert between forms under physiological conditions and can exist as an equilibrium of different oligomers. These oligomers are physiologically relevant and are not misfolded protein; this distinguishes morpheeins from prions and amyloid. The different oligomers have distinct functionality. Interconversion of morpheein forms can be a structural basis for allosteric regulation, an idea noted many years ago, [3] [4] and later revived. [1] [2] [5] [6] A mutation that shifts the normal equilibrium of morpheein forms can serve as the basis for a conformational disease. [7] Features of morpheeins can be exploited for drug discovery. [1] [5] [8] The dice image (Fig 1) represents a morpheein equilibrium containing two different monomeric shapes that dictate assembly to a tetramer or a pentamer. The one protein that is established to function as a morpheein is porphobilinogen synthase, [2] [9] [10] though there are suggestions throughout the literature that other proteins may function as morpheeins (for more information see "Table of Putative Morpheeins" below).
Implications for drug discovery
Conformational differences between subunits of different oligomers and related functional differences of a morpheein provide a starting point for drug discovery. Protein function is dependent on the oligomeric form; therefore, the protein's function can be regulated by shifting the equilibrium of forms. A small molecule compound can shift the equilibrium either by blocking or favoring formation of one of the oligomers. The equilibrium can be shifted using a small molecule that has a preferential binding affinity for only one of the alternate morpheein forms. An inhibitor of porphobilinogen synthase with this mechanism of action has been documented. [5]
Implications for allosteric regulation
The morpheein model of allosteric regulation has similarities to and differences from other models. [1] [6] [11] The concerted model (the Monod, Wyman and Changeux (MWC) model) of allosteric regulation requires all subunits to be in the same conformation or state within an oligomer like the morpheein model. [12] [13] However, neither this model nor the sequential model (Koshland, Nemethy, and Filmer model) takes into account that the protein may dissociate to interconvert between oligomers. [12] [13] [14] [15] Nonetheless, shortly after these theories were described, two groups of workers [3] [4] proposed what is now called the morpheein model and showed that it accounted for the regulatory behavior of glutamate dehydrogenase. [16] Kurganov and Friedrich discussed models of this kind extensively in their books. [17] [18]
Implications for teaching about protein structure-function relationships
It is generally taught [ citation needed] that a given amino acid sequence will have only one physiologically relevant (native) quaternary structure; morpheeins challenge this concept. The morpheein model does not require gross changes in the basic protein fold. [1] The conformational differences that accompany conversion between oligomers may be similar to the protein motions necessary for function of some proteins. [19] The morpheein model highlights the importance of conformational flexibility for protein functionality and offers a potential explanation for proteins showing non- Michaelis-Menten kinetics, hysteresis, and/or protein concentration dependent specific activity. [11]
Implications for understanding the structural basis for disease
The term " conformational disease" generally encompasses mutations that result in misfolded proteins that aggregate, such as Alzheimer's and Creutzfeldt–Jakob diseases. [20] In light of the discovery of morpheeins, however, this definition could be expanded to include mutations that shift an equilibrium of alternate oligomeric forms of a protein. An example of such a conformational disease is ALAD porphyria, which results from a mutation of porphobilinogen synthase that causes a shift in its morpheein equilibrium. [7]
Table of proteins whose published behavior is consistent with that of a morpheein
| Protein | Example species | EC number | CAS number | Alternate oligomers | Evidence |
|---|---|---|---|---|---|
| Acetyl-CoA carboxylase-1 | Gallus domesticus | EC 6.4.1.2 | 9023-93-2 | inactive dimer, active dimer, larger [21] | Effector molecules impact multimerization, [22] Multiple/ protein moonlighting functions [21] |
| α-Acetylgalactosaminidase | Bos taurus | EC 4.3.2.2 | 9027-81-0 | inactive monomer, active tetramer [23] | Substrate binding/turnover impacts multimerization, [23] Protein concentration dependent specific activity, [24] Different assemblies have different activities, [24] Conformationally distinct oligomeric forms. [23] [24] |
| Adenylosuccinate lyase | Bacillus subtilis | EC 4.3.2.2 | 9027-81-0 | monomer, dimer, trimer, tetramer [25] | Mutations shift the equilibrium of oligomers, [26] Oligomer-dependent kinetic parameters, [26] Protein concentration dependent molecular weight [26] |
| Aristolochene synthase | Penicillium roqueforti | EC 4.2.3.9 | 94185-89-4 | monomer, higher order [27] | Protein concentration dependent specific activity [28] |
| L- Asparaginase | Leptosphaeria michotii | EC 3.5.1.1 | 9015-68-3 | dimer, tetramer, inactive octamer [29] | Substrate binding/turnover impacts multimerization [30] |
| Aspartokinase | Escherichia coli | EC 2.7.2.4 & EC 1.1.1.3 | 9012-50-4 | monomer, dimer, tetramer [31] [32] | Multiple/ protein moonlighting functions, [33] Conformationally distinct oligomeric forms [32] |
| ATPase of the ABCA1 transporter | Homo sapiens | dimer, tetramer [34] | Substrate binding/turnover impacts multimerization [34] | ||
| Biotin—(acetyl-CoA-carboxylase) ligase holoenzyme synthetase | Escherichia coli | EC 6.3.4.15 | 37340-95-7 | monomer, dimer [35] | Multiple/ protein moonlighting functions, [35] Different assemblies have different activities [36] |
| Chorismate mutase | Escherichia coli | EC 5.4.99.5 | 9068-30-8 | dimer, trimer, hexamer | Conformationally distinct oligomeric forms [37] |
| Citrate synthase | Escherichia coli | EC 2.3.3.1 | 9027-96-7 | monomer, dimer, trimer, tetramer, pentamer, hexamer, dodecamer [38] | Substrate binding/turnover impacts multimerization, [38] Characterized equilibrium of oligomers, [38] Protein concentration dependent specific activity, [38] pH-dependent oligomeric equilibrium [38] |
| Cyanovirin-N | Nostoc ellipsosporum | 918555-82-5 | monomer and domain-swapped dimer [39] [40] | Characterized equilibrium of oligomers, [41] [42] Conformationally distinct oligomeric forms [41] [42] | |
| 3-oxoacid CoA-transferase | Sus scrofa domestica | EC 2.8.3.5 | 9027-43-4 | dimer, tetramer [43] | Chromatographically separable oligomers, [43] Substrate might preferentially stabilize one form [43] |
| Cystathionine β-synthase | Homo sapiens | EC 4.2.1.22 | 9023-99-8 | multiple - ranges from dimer to 16-mer [44] | Effector molecules impact multimerization, [45] Mutations shift the equilibrium of oligomers, [46] Different assemblies have different activities, [45] disease-causing mutations at sites distant from active site [47] |
| D-amino acid oxidase | EC 1.4.3.3 | 9000-88-8 | monomers, dimers, higher-order oligomers [48] [49] | Oligomer-dependent kinetic parameters [48] [49] | |
| Dihydrolipoamide dehydrogenase | Sus scrofa domestica | EC 1.8.1.4 | 9001-18-7 | monomer, two different dimer forms, tetramer [50] | Multiple/ protein moonlighting functions, [50] Different assemblies have different activities, [50] pH-dependent oligomeric equilibrium, [50] Conformationally distinct oligomeric forms [51] [52] [53] |
| Dopamine β-monooxygenase | Bos taurus | EC 1.14.17.1 | 9013-38-1 | dimers, tetramers [54] [55] [56] | Effector molecules impact multimerization, [54] [55] [56] Characterized equilibrium of oligomers, [54] [55] [56] Oligomer-dependent kinetic parameters [54] [55] [56] |
| Geranylgeranyl pyrophosphate synthase / Farnesyltranstransferase | Homo sapiens | EC 2.5.1.29 | 9032-58-0 | hexamer, octamer [57] [58] [59] | Effector molecules impact multimerization [58] |
| GDP-mannose 6-dehydrogenase | Pseudomonas aeruginosa | EC 1.1.1.132 | 37250-63-8 | trimer, 2 tetramers, and hexamer [60] [61] | Protein concentration dependent specific activity, [62] Kinetic hysteresis [62] |
| Glutamate dehydrogenase | Bos taurus | EC 1.4.1.2 | 9001-46-1 | active & inactive hexamers, higher order [63] | Effector molecules impact multimerization, [64] Characterized equilibrium of oligomers [63] |
| Glutamate racemase | Mycobacterium tuberculosis, Escherichia coli, Bacillus subtilis, Aquifex pyrophilus | EC 5.1.1.3 | 9024-08-02 | monomer, 2 dimers, tetramer [65] [66] [67] [68] [69] | Multiple/ protein moonlighting functions, [70] [71] [72] Characterized equilibrium of oligomers, [68] [69] Conformationally distinct oligomeric forms [65] [66] [67] |
| Glyceraldehyde-3-phosphate dehydrogenase | Oryctolagus cuniculas, Sus scrofa domestica | EC 1.2.1.12 1.2.1.12 | 9001-50-7 | monomer, dimer, tetramer [73] Characterized equilibrium of oligomers, [74] Different assemblies have different activities [75] | |
| Glycerol kinase | Escherichia coli | EC 2.7.1.30 | 9030-66-4 | monomer and 2 tetramers [76] [77] [78] | Characterized equilibrium of oligomers, [76] [77] [78] [79] Conformationally distinct oligomeric forms, [79] [80] Effector functions by preventing domain motion [80] |
| HIV- Integrase | Human immunodeficiency virus-1 | EC 2.7.7.- | monomer, dimer, tetramer, higher order [81] [82] [83] | Effector molecules impact multimerization, [84] Multiple/ protein moonlighting functions, [81] [82] [83] Different assemblies have different activities [83] [84] | |
| HPr-Kinase/phosphatase | Bacillus subtilis, Lactobacillus casei, Mycoplasma pneumoniae, Staphylococcus xylosus | EC 2.7.1.-/ EC 3.1.3.- | 9026-43-1 | monomers, dimers, trimers, hexamers [85] [86] [87] [88] [89] [90] | Effector molecules impact multimerization, [89] Multiple/ protein moonlighting functions, [89] Different assemblies have different activities, [89] pH-dependent oligomeric equilibrium [89] |
| Lactate dehydrogenase | Bacillus stearothermophilus | EC 1.1.1.27 | 9001-60-9 | 2 dimers, tetramer [91] [92] | Effector molecules impact multimerization, [92] Characterized equilibrium of oligomers, [92] Protein concentration dependent specific activity, [92] Mutations shift the equilibrium of oligomers, [93] Oligomer-dependent kinetic parameters, [92] Conformationally distinct oligomeric forms [94] |
| Lon protease | Escherichia coli, Mycobacterium smegmatis | EC 3.4.21.53 | 79818-35-2 | monomer, dimer, trimer, tetramer [95] [96] | Effector molecules impact multimerization, [95] [96] Substrate binding/turnover impacts multimerization, [95] [96] Protein concentration dependent specific activity, [97] Kinetic hysteresis [97] |
| Mitochondrial NAD(P)+ Malic enzyme / [[malate dehydrogenase (oxaloacetate-decarboxylating) (NADP+)]] | Homo sapiens | EC 1.1.1.40 | 9028-47-1 | monomer, 2 dimers, tetramer [98] [99] | Effector molecules impact multimerization, [98] Mutations shift the equilibrium of oligomers, [100] Kinetic hysteresis, [99] |
| Peroxiredoxins | Salmonella typhimurium | EC 1.6.4.- & EC 1.11.1.15 | 207137-51-7 | 2 dimers, decamer | Conformationally distinct oligomeric forms, [101] Different assemblies have different activities [102] |
| Phenylalanine hydroxylase | Homo sapiens | EC 1.14.16.1 | 9029-73-6 | high activity tetramer, low activity tetramer [103] | Substrate binding/turnover impacts multimerization, [104] [105] Conformationally distinct oligomeric forms [106] [107] |
| Phosphoenolpyruvate carboxylase | Escherichia coli, Zea mays | EC [1] | 9067-77-0 | inactive dimer, active tetramer [108] | Effector molecules impact multimerization, Characterized equilibrium of oligomers, [108] Kinetic hysteresis, [108] Conformationally distinct oligomeric forms [109] |
| Phosphofructokinase | Bacillus stearothermophilus, Thermus thermophilus | EC 2.7.1.11 | 9001-80-3 | inactive dimer, active tetramer [108] [110] | Effector molecules impact multimerization, [108] [110] Characterized equilibrium of oligomers [108] [110] |
| Polyphenol oxidase | Agaricus bisporus, Malus domestica, Lactuca sativa L. | EC 1.10.3.1 | 9002-10-2 | monomer, trimer, tetramer, octamer, dodecamer [111] [112] | Multiple/ protein moonlighting functions, [113] Substrate binding/turnover impacts multimerization, [114] Different assemblies have different activities, [115] Kinetic hysteresis [114] |
| Porphobilinogen synthase | Drosophila melanogaster, Danio rerio | EC 4.2.1.24 | 9036-37-7 | dimer, hexamer, octamer [116] [117] | PBGS is the prototype morpheein. [116] |
| Pyruvate kinase | Homo sapiens | EC 2.7.1.40 | 9001-59-6 | active and inactive dimers, active tetramer, monomer, trimer, pentamer [118] [119] | Conformationally distinct oligomeric forms [118] [119] |
| Ribonuclease A | Bos taurus | EC 3.1.27.5 3.1.27.5 | 9901-99-4 | monomer, dimer, trimer, tetramer, hexamer, pentamer, higher order [120] [121] [122] [123] [124] | Multiple/ protein moonlighting functions, [125] [126] [127] Different assemblies have different activities, [125] [126] [127] Conformationally distinct oligomeric forms [121] [123] [124] |
| Ribonucleotide reductase | Mus musculus | EC 1.17.4.1 | 9047-64-7 | tetramer, hexamer [128] [129] [130] [131] | Effector molecules impact multimerization [131] |
| S-adenosyl-L-homocysteine hydrolase | Dictyostelium discoideum | EC 3.3.1.1 | 9025-54-1 | tetramer and other [132] [133] [134] | Effector molecules impact multimerization [132] |
| Biodegrative threonine dehydratase / threonine ammonia-lyase | Escherichia coli | EC 4.3.1.19 4.3.1.19 | 774231-81-1 | 2 monomers, 2 tetramers [135] [136] [137] | Effector molecules impact multimerization, [137] Characterized equilibrium of oligomers, [135] [136] Different assemblies have different activities [135] [136] [137] |
| β- Tryptase | Homo sapiens | EC 3.4.21.59 | 97501-93-4 | active and inactive monomers, active and inactive tetramers [138] [139] [140] [141] [142] [143] [144] [145] [146] [147] | Protein concentration dependent specific activity, [148] Characterized equilibrium of oligomers [148] |
| Tumor necrosis factor-α | Homo sapiens | 94948-61-5 | monomer, dimer, trimer [149] [150] | Different assemblies have different activities [151] | |
| Uracil phosphoribosyltransferase | Escherichia coli | EC 2.4.2.9 | 9030-24-4 | trimer, pentamer [152] | Effector molecules impact multimerization, [152] Substrate binding/turnover impacts multimerization, [152] Different assemblies have different activities [152] |
References
- ^ a b c d e Jaffe, Eileen K. (2005). "Morpheeins – a new structural paradigm for allosteric regulation". Trends in Biochemical Sciences. 30 (9): 490–7. doi: 10.1016/j.tibs.2005.07.003. PMID 16023348.
- ^ a b c Breinig, Sabine; Kervinen, Jukka; Stith, Linda; Wasson, Andrew S; Fairman, Robert; Wlodawer, Alexander; Zdanov, Alexander; Jaffe, Eileen K (2003). "Control of tetrapyrrole biosynthesis by alternate quaternary forms of porphobilinogen synthase". Nature Structural Biology. 10 (9): 757–63. doi: 10.1038/nsb963. PMID 12897770. S2CID 24188785.
- ^ a b Frieden, C (1967). "Treatment of enzyme kinetic data. II. The multisite case: comparison of allosteric models and a possible new mechanism". J. Biol. Chem. 242 (18): 4045–4052. doi: 10.1016/S0021-9258(18)95776-5. PMID 6061697.
- ^ a b Nichol, L W; Jackson, W J H; Winzor, D J (1967). "A theoretical study of the binding of small molecules to a polymerizing protein system: a model for allosteric effects". Biochemistry. 6 (8): 2449–2456. doi: 10.1021/bi00860a022. PMID 6049469.
- ^ a b c Lawrence, Sarah H.; Ramirez, Ursula D.; Tang, Lei; Fazliyez, Farit; Kundrat, Lenka; Markham, George D.; Jaffe, Eileen K. (2008). "Shape Shifting Leads to Small-Molecule Allosteric Drug Discovery". Chemistry & Biology. 15 (6): 586–96. doi: 10.1016/j.chembiol.2008.04.012. PMC 2703447. PMID 18559269.
- ^ a b c Selwood, Trevor; Jaffe, Eileen K. (2012). "Dynamic dissociating homo-oligomers and the control of protein function". Archives of Biochemistry and Biophysics. 519 (2): 131–43. doi: 10.1016/j.abb.2011.11.020. PMC 3298769. PMID 22182754.
- ^ a b Jaffe, Eileen K.; Stith, Linda (2007). "ALAD Porphyria is a Conformational Disease". The American Journal of Human Genetics. 80 (2): 329–37. doi: 10.1086/511444. PMC 1785348. PMID 17236137.
-
^ Jaffe, Eileen K. (2010).
"Morpheeins – A New Pathway For Allosteric Drug Discovery". The Open Conference Proceedings Journal. 1: 1–6.
doi:
10.2174/2210289201001010001 (inactive 2024-03-28).
PMC
3107518.
PMID
21643557.
{{ cite journal}}: CS1 maint: DOI inactive as of March 2024 ( link) - ^ Tang, L.; Stith, L; Jaffe, EK (2005). "Substrate-induced Interconversion of Protein Quaternary Structure Isoforms". Journal of Biological Chemistry. 280 (16): 15786–93. doi: 10.1074/jbc.M500218200. PMID 15710608.
- ^ Jaffe, Eileen K.; Lawrence, Sarah H. (2012). "Allostery and the dynamic oligomerization of porphobilinogen synthase". Archives of Biochemistry and Biophysics. 519 (2): 144–53. doi: 10.1016/j.abb.2011.10.010. PMC 3291741. PMID 22037356.
- ^ a b Lawrence, Sarah H.; Jaffe, Eileen K. (2008). "Expanding the concepts in protein structure-function relationships and enzyme kinetics: Teaching using morpheeins". Biochemistry and Molecular Biology Education. 36 (4): 274–283. doi: 10.1002/bmb.20211. PMC 2575429. PMID 19578473.
- ^ a b Monod, Jacques; Changeux, Jean-Pierre; Jacob, François (1963). "Allosteric proteins and cellular control systems". Journal of Molecular Biology. 6 (4): 306–29. doi: 10.1016/S0022-2836(63)80091-1. PMID 13936070.
- ^ a b Monod, Jacques; Wyman, Jeffries; Changeux, Jean-Pierre (1965). "On the nature of allosteric transitions: A plausible model". Journal of Molecular Biology. 12: 88–118. doi: 10.1016/S0022-2836(65)80285-6. PMID 14343300.
- ^ Koshland, D.E. (1970). "7 The Molecular Basis for Enzyme Regulation". The Enzymes Volume 1. Vol. 1. pp. 341–396. doi: 10.1016/S1874-6047(08)60170-5. ISBN 978-0-12-122701-2.
- ^ Koshland, D. E.; Nemethy, G.; Filmer, D. (1966). "Comparison of Experimental Binding Data and Theoretical Models in Proteins Containing Subunits". Biochemistry. 5 (1): 365–85. doi: 10.1021/bi00865a047. PMID 5938952.
- ^ Frieden, C; Colman, R F (1967). "Glutamate dehydrogenase concentration as a determinant in the effect of purine nucleotides on the enzymatiuc activity". J. Biol. Chem. 242: 1705–1715. doi: 10.1016/S0021-9258(18)96059-X.
- ^ Kurganov, B I (1982). Allosteric Enzymes: Kinetic Behaviour. Chichester: Wiley–Interscience. pp. 151–248. ISBN 978-0471101956.
- ^ Friedrich, P (1984). Supramolecular Enzyme Organization: Quaternary Structure and Beyond. Oxford: Pergamon Press. pp. 66–71. ISBN 0-08-026376-3.
- ^ Gerstein, Mark; Echols, Nathaniel (2004). "Exploring the range of protein flexibility, from a structural proteomics perspective". Current Opinion in Chemical Biology. 8 (1): 14–9. doi: 10.1016/j.cbpa.2003.12.006. PMID 15036151.
- ^ Carrell, Robin W; Lomas, David A (1997). "Conformational disease". The Lancet. 350 (9071): 134–8. doi: 10.1016/S0140-6736(97)02073-4. PMID 9228977. S2CID 39124185.
- ^ a b Boone, A.N.; Brownsey, R.W.; Elliott, J.E.; Kulpa, J.E.; Lee, W.M. (2006). "Regulation of acetyl-CoA carboxylase". Biochemical Society Transactions. 34 (2): 223–7. doi: 10.1042/BST20060223. PMID 16545081.
- ^ Shen, Yang; Volrath, Sandra L.; Weatherly, Stephanie C.; Elich, Tedd D.; Tong, Liang (2004). "A Mechanism for the Potent Inhibition of Eukaryotic Acetyl-Coenzyme a Carboxylase by Soraphen A, a Macrocyclic Polyketide Natural Product". Molecular Cell. 16 (6): 881–91. doi: 10.1016/j.molcel.2004.11.034. PMID 15610732.
- ^ a b c Weissmann, Bernard; Wang, Ching-Te (1971). "Association-dissociation and abnormal kinetics of bovine .alpha.-acetylgalactosaminidase". Biochemistry. 10 (6): 1067–72. doi: 10.1021/bi00782a021. PMID 5550813.
- ^ a b c Weissmann, Bernard; Hinrichsen, Dorotea F. (1969). "Mammalian α-acetylgalactosaminidase. Occurrence, partial purification, and action on linkages in submaxillary mucins". Biochemistry. 8 (5): 2034–43. doi: 10.1021/bi00833a038. PMID 5785223.
- ^ De Zoysa Ariyananda, Lushanti; Colman, Roberta F. (2008). "Evaluation of Types of Interactions in Subunit Association in Bacillus subtilis Adenylosuccinate Lyase". Biochemistry. 47 (9): 2923–34. doi: 10.1021/bi701400c. PMID 18237141.
- ^ a b c Palenchar, Jennifer Brosius; Colman, Roberta F. (2003). "Characterization of a Mutant Bacillus subtilis Adenylosuccinate Lyase Equivalent to a Mutant Enzyme Found in Human Adenylosuccinate Lyase Deficiency: Asparagine 276 Plays an Important Structural Role". Biochemistry. 42 (7): 1831–41. doi: 10.1021/bi020640+. PMID 12590570.
- ^ Hohn, Thomas M.; Plattner, Ronald D. (1989). "Purification and characterization of the sesquiterpene cyclase aristolochene synthase from Penicillium roqueforti". Archives of Biochemistry and Biophysics. 272 (1): 137–43. doi: 10.1016/0003-9861(89)90204-X. PMID 2544140.
- ^ Caruthers, J. M.; Kang, I; Rynkiewicz, MJ; Cane, DE; Christianson, DW (2000). "Crystal Structure Determination of Aristolochene Synthase from the Blue Cheese Mold, Penicillium roqueforti". Journal of Biological Chemistry. 275 (33): 25533–9. doi: 10.1074/jbc.M000433200. PMID 10825154.
- ^ Jerebzoff-Quintin, Simonne; Jerebzoff, Stephan (1985). "L-Asparaginase activity in Leptosphaeria michotii. Isolation and properties of two forms of the enzyme". Physiologia Plantarum. 64: 74–80. doi: 10.1111/j.1399-3054.1985.tb01215.x.
- ^ Yun, Mi-Kyung; Nourse, Amanda; White, Stephen W.; Rock, Charles O.; Heath, Richard J. (2007). "Crystal Structure and Allosteric Regulation of the Cytoplasmic Escherichia coli l-Asparaginase I". Journal of Molecular Biology. 369 (3): 794–811. doi: 10.1016/j.jmb.2007.03.061. PMC 1991333. PMID 17451745.
- ^ Garel, J.-R. (1980). "Sequential Folding of a Bifunctional Allosteric Protein". Proceedings of the National Academy of Sciences. 77 (6): 3379–3383. Bibcode: 1980PNAS...77.3379G. doi: 10.1073/pnas.77.6.3379. JSTOR 8892. PMC 349619. PMID 6774337.
- ^ a b Kotaka, M.; Ren, J.; Lockyer, M.; Hawkins, A. R.; Stammers, D. K. (2006). "Structures of R- and T-state Escherichia coli Aspartokinase III: MECHANISMS OF THE ALLOSTERIC TRANSITION AND INHIBITION BY LYSINE". Journal of Biological Chemistry. 281 (42): 31544–52. doi: 10.1074/jbc.M605886200. PMID 16905770.
- ^ Ogilvie, JW; Vickers, LP; Clark, RB; Jones, MM (1975). "Aspartokinase I-homoserine dehydrogenase I of Escherichia coli K12 (lambda). Activation by monovalent cations and an analysis of the effect of the adenosine triphosphate-magnesium ion complex on this activation process". The Journal of Biological Chemistry. 250 (4): 1242–50. doi: 10.1016/S0021-9258(19)41805-X. PMID 163250.
- ^ a b Trompier, D.; Alibert, M; Davanture, S; Hamon, Y; Pierres, M; Chimini, G (2006). "Transition from Dimers to Higher Oligomeric Forms Occurs during the ATPase Cycle of the ABCA1 Transporter". Journal of Biological Chemistry. 281 (29): 20283–90. doi: 10.1074/jbc.M601072200. PMID 16709568.
- ^ a b Eisenstein, Edward; Beckett, Dorothy (1999). "Dimerization of theEscherichiacoliBiotin Repressor: Corepressor Function in Protein Assembly". Biochemistry. 38 (40): 13077–84. doi: 10.1021/bi991241q. PMID 10529178.
- ^ Streaker, Emily D.; Beckett, Dorothy (1998). "Coupling of Site-Specific DNA Binding to Protein Dimerization in Assembly of the Biotin Repressor−Biotin Operator Complex". Biochemistry. 37 (9): 3210–9. doi: 10.1021/bi9715019. PMID 9485476.
- ^ Vamvaca, Katherina; Butz, Maren; Walter, Kai U.; Taylor, Sean V.; Hilvert, Donald (2005). "Simultaneous optimization of enzyme activity and quaternary structure by directed evolution". Protein Science. 14 (8): 2103–14. doi: 10.1110/ps.051431605. PMC 2279322. PMID 15987889.
- ^ a b c d e Tong, E. K.; Duckworth, Harry W. (1975). "Quaternary structure of citrate synthase from Escherichia coli K 12". Biochemistry. 14 (2): 235–41. doi: 10.1021/bi00673a007. PMID 1091285.
- ^ Bewley, Carole A.; Gustafson, Kirk R.; Boyd, Michael R.; Covell, David G.; Bax, Ad; Clore, G. Marius; Gronenborn, Angela M. (1998). "Solution structure of cyanovirin-N, a potent HIV-inactivating protein". Nature Structural Biology. 5 (7): 571–8. doi: 10.1038/828. PMID 9665171. S2CID 11367037.
- ^ Yang, Fan; Bewley, Carole A; Louis, John M; Gustafson, Kirk R; Boyd, Michael R; Gronenborn, Angela M; Clore, G.Marius; Wlodawer, Alexander (1999). "Crystal structure of cyanovirin-N, a potent HIV-inactivating protein, shows unexpected domain swapping". Journal of Molecular Biology. 288 (3): 403–12. doi: 10.1006/jmbi.1999.2693. PMID 10329150. S2CID 308708.
- ^ a b Barrientos, LG; Gronenborn, AM (2005). "The highly specific carbohydrate-binding protein cyanovirin-N: Structure, anti-HIV/Ebola activity and possibilities for therapy". Mini Reviews in Medicinal Chemistry. 5 (1): 21–31. doi: 10.2174/1389557053402783. PMID 15638789.
- ^ a b Barrientos, LG; Louis, JM; Botos, I; Mori, T; Han, Z; O'Keefe, BR; Boyd, MR; Wlodawer, A; et al. (2002). "The domain-swapped dimer of cyanovirin-N is in a metastable folded state: Reconciliation of X-ray and NMR structures". Structure. 10 (5): 673–86. doi: 10.1016/S0969-2126(02)00758-X. PMID 12015150.
- ^ a b c Rochet, Jean-Christophe; Brownie, Edward R.; Oikawa, Kim; Hicks, Leslie D.; Fraser, Marie E.; James, Michael N. G.; Kay, Cyril M.; Bridger, William A.; et al. (2000). "Pig Heart CoA Transferase Exists as Two Oligomeric Forms Separated by a Large Kinetic Barrier". Biochemistry. 39 (37): 11291–302. doi: 10.1021/bi0003184. PMID 10985774.
- ^ Frank, Nina; Kery, Vladimir; MacLean, Kenneth N.; Kraus, Jan P. (2006). "Solvent-Accessible Cysteines in Human Cystathionine β-Synthase: Crucial Role of Cysteine 431 inS-Adenosyl-l-methionine Binding". Biochemistry. 45 (36): 11021–9. doi: 10.1021/bi060737m. PMID 16953589.
- ^ a b Sen, Suvajit; Banerjee, Ruma (2007). "A Pathogenic Linked Mutation in the Catalytic Core of Human Cystathionine β-Synthase Disrupts Allosteric Regulation and Allows Kinetic Characterization of a Full-Length Dimer". Biochemistry. 46 (13): 4110–6. doi: 10.1021/bi602617f. PMC 3204387. PMID 17352495.
- ^ Kery, Vladimir; Poneleit, Loelle; Kraus, Jan P. (1998). "Trypsin Cleavage of Human Cystathionine β-Synthase into an Evolutionarily Conserved Active Core: Structural and Functional Consequences". Archives of Biochemistry and Biophysics. 355 (2): 222–32. doi: 10.1006/abbi.1998.0723. PMID 9675031.
- ^ Shan, Xiaoyin; Kruger, Warren D. (1998). "Correction of disease-causing CBS mutations in yeast". Nature Genetics. 19 (1): 91–3. doi: 10.1038/ng0598-91. PMID 9590298. S2CID 47102642.
- ^ a b Antonini, E; Brunori, M; Bruzzesi, R; Chiancone, E; Massey, V (1966). "Association-dissociation phenomena of D-amino acid oxidase". The Journal of Biological Chemistry. 241 (10): 2358–66. doi: 10.1016/S0021-9258(18)96629-9. PMID 4380380.
- ^ a b Massey, V; Curti, B; Ganther, H (1966). "A temperature-dependent conformational change in D-amino acid oxidase and its effect on catalysis". The Journal of Biological Chemistry. 241 (10): 2347–57. doi: 10.1016/S0021-9258(18)96628-7. PMID 5911617.
- ^ a b c d Babady, N. E.; Pang, Y.-P.; Elpeleg, O.; Isaya, G. (2007). "Cryptic proteolytic activity of dihydrolipoamide dehydrogenase". Proceedings of the National Academy of Sciences. 104 (15): 6158–63. Bibcode: 2007PNAS..104.6158B. doi: 10.1073/pnas.0610618104. PMC 1851069. PMID 17404228.
- ^ Muiswinkel-Voetberg, H.; Visser, Jaap; Veeger, Cornelis (1973). "Conformational Studies on Lipoamide Dehydrogenase from Pig Heart. 1. Interconversion of Dissociable and Non-Dissociable Forms". European Journal of Biochemistry. 33 (2): 265–70. doi: 10.1111/j.1432-1033.1973.tb02679.x. PMID 4348439.
- ^ Klyachko, N. L.; Shchedrina, VA; Efimov, AV; Kazakov, SV; Gazaryan, IG; Kristal, BS; Brown, AM (2005). "PH-dependent Substrate Preference of Pig Heart Lipoamide Dehydrogenase Varies with Oligomeric State: RESPONSE TO MITOCHONDRIAL MATRIX ACIDIFICATION". Journal of Biological Chemistry. 280 (16): 16106–14. doi: 10.1074/jbc.M414285200. PMID 15710613.
- ^ Muiswinkel-Voetberg, H.; Veeger, Cornelis (1973). "Conformational Studies on Lipoamide Dehydrogenase from Pig Heart. 2. Spectroscopic Studies on the Apoenzyme and the Monomeric and Dimeric Forms". European Journal of Biochemistry. 33 (2): 271–8. doi: 10.1111/j.1432-1033.1973.tb02680.x. PMID 4348440.
- ^ a b c d Saxena, Ashima; Hensley, Preston; Osborne, James C.; Fleming, Patrick J. (1985). "The pH-dependent Subunit Dissociation and Catalytic Activity of Bovine Dopamine β-Hydroxylase". Journal of Biological Chemistry. 260 (6): 3386–92. doi: 10.1016/S0021-9258(19)83633-5. PMID 3972830.
- ^ a b c d Dhawan, S; Hensley, P; Osborne Jr, JC; Fleming, PJ (1986). "Adenosine 5'-diphosphate-dependent subunit dissociation of bovine dopamine beta-hydroxylase". The Journal of Biological Chemistry. 261 (17): 7680–4. doi: 10.1016/S0021-9258(19)57453-1. PMID 3711102.
- ^ a b c d Stewart, L C; Klinman, J P (1988). "Dopamine Beta-Hydroxylase of Adrenal Chromaffin Granules: Structure and Function". Annual Review of Biochemistry. 57: 551–92. doi: 10.1146/annurev.bi.57.070188.003003. PMID 3052283.
- ^ Kuzuguchi, T.; Morita, Y; Sagami, I; Sagami, H; Ogura, K (1999). "Human Geranylgeranyl Diphosphate Synthase. CDNA CLONING AND EXPRESSION". Journal of Biological Chemistry. 274 (9): 5888–94. doi: 10.1074/jbc.274.9.5888. PMID 10026212.
- ^ a b Kavanagh, K. L.; Dunford, JE; Bunkoczi, G; Russell, RG; Oppermann, U (2006). "The Crystal Structure of Human Geranylgeranyl Pyrophosphate Synthase Reveals a Novel Hexameric Arrangement and Inhibitory Product Binding" (PDF). Journal of Biological Chemistry. 281 (31): 22004–12. doi: 10.1074/jbc.M602603200. PMID 16698791.
- ^ Miyagi, Y.; Matsumura, Y.; Sagami, H. (2007). "Human Geranylgeranyl Diphosphate Synthase is an Octamer in Solution". Journal of Biochemistry. 142 (3): 377–81. doi: 10.1093/jb/mvm144. PMID 17646172.
- ^ Snook, Christopher F.; Tipton, Peter A.; Beamer, Lesa J. (2003). "Crystal Structure of GDP-Mannose Dehydrogenase: A Key Enzyme of Alginate Biosynthesis inP. Aeruginosa". Biochemistry. 42 (16): 4658–68. doi: 10.1021/bi027328k. PMID 12705829.
- ^ Roychoudhury, S; May, TB; Gill, JF; Singh, SK; Feingold, DS; Chakrabarty, AM (1989). "Purification and characterization of guanosine diphospho-D-mannose dehydrogenase. A key enzyme in the biosynthesis of alginate by Pseudomonas aeruginosa". The Journal of Biological Chemistry. 264 (16): 9380–5. doi: 10.1016/S0021-9258(18)60542-3. PMID 2470755.
- ^ a b Naught, Laura E.; Gilbert, Sunny; Imhoff, Rebecca; Snook, Christopher; Beamer, Lesa; Tipton, Peter (2002). "Allosterism and Cooperativity inPseudomonas aeruginosaGDP-Mannose Dehydrogenase". Biochemistry. 41 (30): 9637–45. doi: 10.1021/bi025862m. PMID 12135385.
- ^ a b Fisher, Harvey F. (2006). "Glutamate Dehydrogenase—ligand Complexes and Their Relationship to the Mechanism of the Reaction". Advances in Enzymology and Related Areas of Molecular Biology. Advances in Enzymology - and Related Areas of Molecular Biology. Vol. 39. pp. 369–417. doi: 10.1002/9780470122846.ch6. ISBN 978-0-470-12284-6. PMID 4147773.
- ^ Huang, CY; Frieden, C (1972). "The mechanism of ligand-induced structural changes in glutamate dehydrogenase. Studies of the rate of depolymerization and isomerization effected by coenzymes and guanine nucleotides". The Journal of Biological Chemistry. 247 (11): 3638–46. doi: 10.1016/S0021-9258(19)45188-0. PMID 4402280.
- ^ a b Kim, Sang Suk; Choi, I.-G.; Kim, Sung-Hou; Yu, Y. G. (1999). "Molecular cloning, expression, and characterization of a thermostable glutamate racemase from a hyperthermophilic bacterium, Aquifex pyrophilus". Extremophiles. 3 (3): 175–83. doi: 10.1007/s007920050114. PMID 10484173. S2CID 709039.
- ^ a b Lundqvist, Tomas; Fisher, Stewart L.; Kern, Gunther; Folmer, Rutger H. A.; Xue, Yafeng; Newton, D. Trevor; Keating, Thomas A.; Alm, Richard A.; et al. (2007). "Exploitation of structural and regulatory diversity in glutamate racemases". Nature. 447 (7146): 817–22. Bibcode: 2007Natur.447..817L. doi: 10.1038/nature05689. PMID 17568739. S2CID 4408683.
- ^ a b May, Melissa; Mehboob, Shahila; Mulhearn, Debbie C.; Wang, Zhiqiang; Yu, Huidong; Thatcher, Gregory R.J.; Santarsiero, Bernard D.; Johnson, Michael E.; et al. (2007). "Structural and Functional Analysis of Two Glutamate Racemase Isozymes from Bacillus anthracis and Implications for Inhibitor Design". Journal of Molecular Biology. 371 (5): 1219–37. doi: 10.1016/j.jmb.2007.05.093. PMC 2736553. PMID 17610893.
- ^ a b Taal, Makie A.; Sedelnikova, Svetlana E.; Ruzheinikov, Sergey N.; Baker, Patrick J.; Rice, David W. (2004). "Expression, purification and preliminary X-ray analysis of crystals ofBacillus subtilisglutamate racemase". Acta Crystallographica Section D. 60 (11): 2031–4. Bibcode: 2004AcCrD..60.2031T. doi: 10.1107/S0907444904021134. PMID 15502318.
- ^ a b Kim, Kook-Han; Bong, Young-Jong; Park, Joon Kyu; Shin, Key-Jung; Hwang, Kwang Yeon; Kim, Eunice Eunkyeong (2007). "Structural Basis for Glutamate Racemase Inhibition". Journal of Molecular Biology. 372 (2): 434–43. doi: 10.1016/j.jmb.2007.05.003. PMID 17658548.
- ^ Ashiuchi, M.; Kuwana, E; Yamamoto, T; Komatsu, K; Soda, K; Misono, H (2002). "Glutamate Racemase is an Endogenous DNA Gyrase Inhibitor". Journal of Biological Chemistry. 277 (42): 39070–3. doi: 10.1074/jbc.C200253200. hdl: 10126/3383. PMID 12213801.
- ^ Ashiuchi, M.; Tani, K.; Soda, K.; Misono, H. (1998). "Properties of Glutamate Racemase from Bacillus subtilis IFO 3336 Producing Poly- -Glutamate". Journal of Biochemistry. 123 (6): 1156–63. doi: 10.1093/oxfordjournals.jbchem.a022055. PMID 9604005.
- ^ Sengupta, S.; Ghosh, S.; Nagaraja, V. (2008). "Moonlighting function of glutamate racemase from Mycobacterium tuberculosis: Racemization and DNA gyrase inhibition are two independent activities of the enzyme". Microbiology. 154 (9): 2796–803. doi: 10.1099/mic.0.2008/020933-0. PMID 18757813.
- ^ Sirover, Michael A (1999). "New insights into an old protein: The functional diversity of mammalian glyceraldehyde-3-phosphate dehydrogenase". Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology. 1432 (2): 159–84. doi: 10.1016/S0167-4838(99)00119-3. PMID 10407139.
- ^ Constantinides, SM; Deal Jr, WC (1969). "Reversible dissociation of tetrameric rabbit muscle glyceraldehyde 3-phosphate dehydrogenase into dimers or monomers by adenosine triphosphate". The Journal of Biological Chemistry. 244 (20): 5695–702. doi: 10.1016/S0021-9258(18)63615-4. PMID 4312250.
- ^ Kumagai, H; Sakai, H (1983). "A porcine brain protein (35 K protein) which bundles microtubules and its identification as glyceraldehyde 3-phosphate dehydrogenase". Journal of Biochemistry. 93 (5): 1259–69. doi: 10.1093/oxfordjournals.jbchem.a134260. PMID 6885722.
- ^ a b De Riel, Jon K.; Paulus, Henry (1978). "Subunit dissociation in the allosteric regulation of glycerol kinase from Escherichia coli. 2. Physical evidence". Biochemistry. 17 (24): 5141–6. doi: 10.1021/bi00617a011. PMID 215195.
- ^ a b De Riel, Jon K.; Paulus, Henry (1978). "Subunit dissociation in the allosteric regulation of glycerol kinase from Escherichia coli. 1. Kinetic evidence". Biochemistry. 17 (24): 5134–40. doi: 10.1021/bi00617a010. PMID 215194.
- ^ a b De Riel, Jon K.; Paulus, Henry (1978). "Subunit dissociation in the allosteric regulation of glycerol kinase from Escherichia coli. 3. Role in desensitization". Biochemistry. 17 (24): 5146–50. doi: 10.1021/bi00617a012. PMID 31903.
- ^ a b Feese, Michael D; Faber, H Rick; Bystrom, Cory E; Pettigrew, Donald W; Remington, S James (1998). "Glycerol kinase from Escherichia coli and an Ala65→Thr mutant: The crystal structures reveal conformational changes with implications for allosteric regulation". Structure. 6 (11): 1407–18. doi: 10.1016/S0969-2126(98)00140-3. PMID 9817843.
- ^ a b Bystrom, Cory E.; Pettigrew, Donald W.; Branchaud, Bruce P.; O'Brien, Patrick; Remington, S. James (1999). "Crystal Structures ofEscherichia coliGlycerol Kinase Variant S58→W in Complex with Nonhydrolyzable ATP Analogues Reveal a Putative Active Conformation of the Enzyme as a Result of Domain Motion". Biochemistry. 38 (12): 3508–18. doi: 10.1021/bi982460z. PMID 10090737.
- ^ a b Deprez, Eric; Tauc, Patrick; Leh, Hervé; Mouscadet, Jean-François; Auclair, Christian; Brochon, Jean-Claude (2000). "Oligomeric States of the HIV-1 Integrase As Measured by Time-Resolved Fluorescence Anisotropy". Biochemistry. 39 (31): 9275–84. doi: 10.1021/bi000397j. PMID 10924120.
- ^ a b Deprez, E.; Tauc, P.; Leh, H.; Mouscadet, J.-F.; Auclair, C.; Hawkins, M. E.; Brochon, J.-C. (2001). "DNA binding induces dissociation of the multimeric form of HIV-1 integrase: A time-resolved fluorescence anisotropy study". Proceedings of the National Academy of Sciences. 98 (18): 10090–5. Bibcode: 2001PNAS...9810090D. doi: 10.1073/pnas.181024498. PMC 56920. PMID 11504911.
- ^ a b c Faure, A. l.; Calmels, C; Desjobert, C; Castroviejo, M; Caumont-Sarcos, A; Tarrago-Litvak, L; Litvak, S; Parissi, V (2005). "HIV-1 integrase crosslinked oligomers are active in vitro". Nucleic Acids Research. 33 (3): 977–86. doi: 10.1093/nar/gki241. PMC 549407. PMID 15718297.
- ^ a b Guiot, E.; Carayon, K; Delelis, O; Simon, F; Tauc, P; Zubin, E; Gottikh, M; Mouscadet, JF; et al. (2006). "Relationship between the Oligomeric Status of HIV-1 Integrase on DNA and Enzymatic Activity". Journal of Biological Chemistry. 281 (32): 22707–19. doi: 10.1074/jbc.M602198200. PMID 16774912.
- ^ Fieulaine, S.; Morera, S; Poncet, S; Monedero, V; Gueguen-Chaignon, V; Galinier, A; Janin, J; Deutscher, J; et al. (2001). "X-ray structure of HPr kinase: A bacterial protein kinase with a P-loop nucleotide-binding domain". The EMBO Journal. 20 (15): 3917–27. doi: 10.1093/emboj/20.15.3917. PMC 149164. PMID 11483495.
- ^ Márquez, José Antonio; Hasenbein, Sonja; Koch, Brigitte; Fieulaine, Sonia; Nessler, Sylvie; Russell, Robert B.; Hengstenberg, Wolfgang; Scheffzek, Klaus (2002). "Structure of the full-length HPr kinase/phosphatase from Staphylococcus xylosus at 1.95 Å resolution: Mimicking the product/substrate of the phospho transfer reactions". Proceedings of the National Academy of Sciences. 99 (6): 3458–63. Bibcode: 2002PNAS...99.3458M. doi: 10.1073/pnas.052461499. JSTOR 3058148. PMC 122545. PMID 11904409.
- ^ Allen, Gregory S.; Steinhauer, Katrin; Hillen, Wolfgang; Stülke, Jörg; Brennan, Richard G. (2003). "Crystal Structure of HPr Kinase/Phosphatase from Mycoplasma pneumoniae". Journal of Molecular Biology. 326 (4): 1203–17. doi: 10.1016/S0022-2836(02)01378-5. PMID 12589763.
- ^ Poncet, Sandrine; Mijakovic, Ivan; Nessler, Sylvie; Gueguen-Chaignon, Virginie; Chaptal, Vincent; Galinier, Anne; Boël, Grégory; Mazé, Alain; et al. (2004). "HPr kinase/phosphorylase, a Walker motif A-containing bifunctional sensor enzyme controlling catabolite repression in Gram-positive bacteria". Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics. 1697 (1–2): 123–35. doi: 10.1016/j.bbapap.2003.11.018. PMID 15023355.
- ^ a b c d e Ramstrom, H.; Sanglier, S; Leize-Wagner, E; Philippe, C; Van Dorsselaer, A; Haiech, J (2002). "Properties and Regulation of the Bifunctional Enzyme HPr Kinase/Phosphatase in Bacillus subtilis". Journal of Biological Chemistry. 278 (2): 1174–85. doi: 10.1074/jbc.M209052200. PMID 12411438.
- ^ Jault, J.-M.; Fieulaine, S; Nessler, S; Gonzalo, P; Di Pietro, A; Deutscher, J; Galinier, A (2000). "The HPr Kinase from Bacillus subtilis is a Homo-oligomeric Enzyme Which Exhibits Strong Positive Cooperativity for Nucleotide and Fructose 1,6-Bisphosphate Binding" (PDF). Journal of Biological Chemistry. 275 (3): 1773–80. doi: 10.1074/jbc.275.3.1773. PMID 10636874.
- ^ Clarke, Anthony R.; Waldman, Adam D.B.; Munro, Ian; Holbrook, J.John (1985). "Changes in the state of subunit association of lactate dehydrogenase from Bacillus stearothermophilus". Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology. 828 (3): 375–9. doi: 10.1016/0167-4838(85)90319-X. PMID 3986214.
- ^ a b c d e Clarke, Anthony R.; Waldman, Adam D.B.; Hart, Keith W.; John Holbrook, J. (1985). "The rates of defined changes in protein structure during the catalytic cycle of lactate dehydrogenase". Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology. 829 (3): 397–407. doi: 10.1016/0167-4838(85)90250-X. PMID 4005269.
- ^ Clarke, Anthony R.; Wigley, Dale B.; Barstow, David A.; Chia, William N.; Atkinson, Tony; Holbrook, J.John (1987). "A single amino acid substitution deregulates a bacterial lactate dehydrogenase and stabilizes its tetrameric structure". Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology. 913 (1): 72–80. doi: 10.1016/0167-4838(87)90234-2. PMID 3580377.
- ^ Cameron, Alexander D.; Roper, David I.; Moreton, Kathleen M.; Muirhead, Hilary; Holbrook, J.John; Wigley, Dale B. (1994). "Allosteric Activation in Bacillus stearothermophilus Lactate Dehydrogenase Investigated by an X-ray Crystallographic Analysis of a Mutant Designed to Prevent Tetramerization of the Enzyme". Journal of Molecular Biology. 238 (4): 615–25. doi: 10.1006/jmbi.1994.1318. PMID 8176749.
- ^ a b c Roudiak, Stanislav G.; Shrader, Thomas E. (1998). "Functional Role of the N-Terminal Region of the Lon Protease fromMycobacterium smegmatis". Biochemistry. 37 (32): 11255–63. doi: 10.1021/bi980945h. PMID 9698372.
- ^ a b c Rudyak, Stanislav G.; Brenowitz, Michael; Shrader, Thomas E. (2001). "Mg2+-Linked Oligomerization Modulates the Catalytic Activity of the Lon (La) Protease from Mycobacterium smegmatis". Biochemistry. 40 (31): 9317–23. doi: 10.1021/bi0102508. PMID 11478899.
- ^ a b Vineyard, Diana; Patterson-Ward, Jessica; Lee, Irene (2006). "Single-Turnover Kinetic Experiments Confirm the Existence of High- and Low-Affinity ATPase Sites inEscherichia coliLon Protease". Biochemistry. 45 (14): 4602–10. doi: 10.1021/bi052377t. PMC 2515378. PMID 16584195.
- ^ a b Yang, Zhiru; Lanks, Charles W.; Tong, Liang (2002). "Molecular Mechanism for the Regulation of Human Mitochondrial NAD(P)+-Dependent Malic Enzyme by ATP and Fumarate". Structure. 10 (7): 951–60. doi: 10.1016/S0969-2126(02)00788-8. PMID 12121650.
- ^ a b Gerald e, Edwards; Carlos s, Andreo (1992). "NADP-malic enzyme from plants". Phytochemistry. 31 (6): 1845–57. Bibcode: 1992PChem..31.1845G. doi: 10.1016/0031-9422(92)80322-6. PMID 1368216.
- ^ Hsieh, J.-Y.; Chen, S.-H.; Hung, H.-C. (2009). "Functional Roles of the Tetramer Organization of Malic Enzyme". Journal of Biological Chemistry. 284 (27): 18096–105. doi: 10.1074/jbc.M109.005082. PMC 2709377. PMID 19416979.
- ^ Poole, Leslie B. (2005). "Bacterial defenses against oxidants: Mechanistic features of cysteine-based peroxidases and their flavoprotein reductases". Archives of Biochemistry and Biophysics. 433 (1): 240–54. doi: 10.1016/j.abb.2004.09.006. PMID 15581580.
- ^ Aran, Martin; Ferrero, Diego S.; Pagano, Eduardo; Wolosiuk, Ricardo A. (2009). "Typical 2-Cys peroxiredoxins - modulation by covalent transformations and noncovalent interactions". FEBS Journal. 276 (9): 2478–93. doi: 10.1111/j.1742-4658.2009.06984.x. hdl: 11336/20656. PMID 19476489. S2CID 1698327.
- ^ Bjørgo, Elisa; De Carvalho, Raquel Margarida Negrão; Flatmark, Torgeir (2001). "A comparison of kinetic and regulatory properties of the tetrameric and dimeric forms of wild-type and Thr427→Pro mutant human phenylalanine hydroxylase". European Journal of Biochemistry. 268 (4): 997–1005. doi: 10.1046/j.1432-1327.2001.01958.x. PMID 11179966.
- ^ Martinez, Aurora; Knappskog, Per M.; Olafsdottir, Sigridur; Døskeland, Anne P.; Eiken, Hans Geir; Svebak, Randi Myrseth; Bozzini, MeriLisa; Apold, Jaran; et al. (1995). "Expression of recombinant human phenylalanine hydroxylase as fusion protein in Escherichia coli circumvents proteolytic degradation by host cell proteases. Isolation and characterization of the wild-type enzyme". The Biochemical Journal. 306 (2): 589–97. doi: 10.1042/bj3060589. PMC 1136558. PMID 7887915.
- ^ Knappskog, Per M.; Flatmark, Torgeir; Aarden, Johanna M.; Haavik, Jan; Martinez, Aurora (1996). "Structure/Function Relationships in Human Phenylalanine Hydroxylase. Effect of Terminal Deletions on the Oligomerization, Activation and Cooperativity of Substrate Binding to the Enzyme". European Journal of Biochemistry. 242 (3): 813–21. doi: 10.1111/j.1432-1033.1996.0813r.x. PMID 9022714.
- ^ Phillips, Robert S.; Parniak, Michael A.; Kaufman, Seymour (1984). "Spectroscopic investigation of ligand interaction with hepatic phenylalanine hydroxylase: Evidence for a conformational change associated with activation". Biochemistry. 23 (17): 3836–42. doi: 10.1021/bi00312a007. PMID 6487579.
- ^ Fusetti, F.; Erlandsen, H; Flatmark, T; Stevens, RC (1998). "Structure of Tetrameric Human Phenylalanine Hydroxylase and Its Implications for Phenylketonuria". Journal of Biological Chemistry. 273 (27): 16962–7. doi: 10.1074/jbc.273.27.16962. PMID 9642259.
- ^ a b c d e f Wohl, RC; Markus, G (1972). "Phosphoenolpyruvate carboxylase of Escherichia coli. Purification and some properties". The Journal of Biological Chemistry. 247 (18): 5785–92. doi: 10.1016/S0021-9258(19)44827-8. PMID 4560418.
- ^ Kai, Yasushi; Matsumura, Hiroyoshi; Izui, Katsura (2003). "Phosphoenolpyruvate carboxylase: Three-dimensional structure and molecular mechanisms". Archives of Biochemistry and Biophysics. 414 (2): 170–9. doi: 10.1016/S0003-9861(03)00170-X. PMID 12781768.
- ^ a b c Xu, Jing; Oshima, Tairo; Yoshida, Masasuke (1990). "Tetramer-dimer conversion of phosphofructokinase from Thermus thermophilus induced by its allosteric effectors". Journal of Molecular Biology. 215 (4): 597–606. doi: 10.1016/S0022-2836(05)80171-8. PMID 2146397.
- ^ Jolley Jr, RL; Mason, HS (1965). "The Multiple Forms of Mushroom Tyrosinase. Interconversion". The Journal of Biological Chemistry. 240: PC1489–91. doi: 10.1016/S0021-9258(18)97603-9. PMID 14284774.
- ^ Jolley Jr, RL; Robb, DA; Mason, HS (1969). "The multiple forms of mushroom tyrosinase. Association-dissociation phenomena". The Journal of Biological Chemistry. 244 (6): 1593–9. doi: 10.1016/S0021-9258(18)91800-4. PMID 4975157.
- ^ Mallette, MF; Dawson, CR (1949). "On the nature of highly purified mushroom tyrosinase preparations". Archives of Biochemistry. 23 (1): 29–44. PMID 18135760.
- ^ a b Chazarra, Soledad; García-Carmona, Francisco; Cabanes, Juana (2001). "Hysteresis and Positive Cooperativity of Iceberg Lettuce Polyphenol Oxidase". Biochemical and Biophysical Research Communications. 289 (3): 769–75. doi: 10.1006/bbrc.2001.6014. PMID 11726215.
- ^ Harel, E.; Mayer, A.M. (1968). "Interconversion of sub-units of catechol oxidase from apple chloroplasts". Phytochemistry. 7 (2): 199–204. Bibcode: 1968PChem...7..199H. doi: 10.1016/S0031-9422(00)86315-3.
- ^ a b Jaffe EK, Lawrence SH (March 2012). "Allostery and the dynamic oligomerization of porphobilinogen synthase". Arch. Biochem. Biophys. 519 (2): 144–53. doi: 10.1016/j.abb.2011.10.010. PMC 3291741. PMID 22037356.
- ^ Breinig S, Kervinen J, Stith L, Wasson AS, Fairman R, Wlodawer A, Zdanov A, Jaffe EK (September 2003). "Control of tetrapyrrole biosynthesis by alternate quaternary forms of porphobilinogen synthase". Nat. Struct. Biol. 10 (9): 757–63. doi: 10.1038/nsb963. PMID 12897770. S2CID 24188785.
- ^ a b Schulz, Ju¨Rgen; Sparmann, Gisela; Hofmann, Eberhard (1975). "Alanine-mediated reversible inactivation of tumour pyruvate kinase caused by a tetramer-dimer transition". FEBS Letters. 50 (3): 346–50. doi: 10.1016/0014-5793(75)90064-2. PMID 1116605. S2CID 5665440.
- ^ a b Ibsen, KH; Schiller, KW; Haas, TA (1971). "Interconvertible kinetic and physical forms of human erythrocyte pyruvate kinase". The Journal of Biological Chemistry. 246 (5): 1233–40. doi: 10.1016/S0021-9258(19)76963-4. PMID 5545066.
- ^ Liu, Yanshun; Gotte, Giovanni; Libonati, Massimo; Eisenberg, David (2009). "Structures of the two 3D domain-swapped RNase a trimers". Protein Science. 11 (2): 371–80. doi: 10.1110/ps.36602. PMC 2373430. PMID 11790847.
- ^ a b Gotte, Giovanni; Bertoldi, Mariarita; Libonati, Massimo (1999). "Structural versatility of bovine ribonuclease A. Distinct conformers of trimeric and tetrameric aggregates of the enzyme". European Journal of Biochemistry. 265 (2): 680–7. doi: 10.1046/j.1432-1327.1999.00761.x. PMID 10504400.
- ^ Gotte, Giovanni; Laurents, Douglas V.; Libonati, Massimo (2006). "Three-dimensional domain-swapped oligomers of ribonuclease A: Identification of a fifth tetramer, pentamers and hexamers, and detection of trace heptameric, octameric and nonameric species". Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics. 1764 (1): 44–54. doi: 10.1016/j.bbapap.2005.10.011. PMID 16310422.
- ^ a b Gotte, Giovanni; Libonati, Massimo (1998). "Two different forms of aggregated dimers of ribonuclease A". Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology. 1386 (1): 106–112. doi: 10.1016/S0167-4838(98)00087-9. PMID 9675255.
- ^ a b Libonati, Massimo; Gotte, Giovanni (2004). "Oligomerization of bovine ribonuclease A: Structural and functional features of its multimers". Biochemical Journal. 380 (2): 311–27. doi: 10.1042/BJ20031922. PMC 1224197. PMID 15104538.
- ^ a b Libonati, M. (2004). "Biological actions of the oligomers of ribonuclease A". Cellular and Molecular Life Sciences. 61 (19–20): 2431–6. doi: 10.1007/s00018-004-4302-x. PMID 15526151. S2CID 8769502.
- ^ a b Libonati, M; Bertoldi, M; Sorrentino, S (1996). "The activity on double-stranded RNA of aggregates of ribonuclease a higher than dimers increases as a function of the size of the aggregates". The Biochemical Journal. 318 (1): 287–90. doi: 10.1042/bj3180287. PMC 1217620. PMID 8761484.
- ^ a b Libonati, M.; Gotte, G.; Vottariello, F. (2008). "A Novel Biological Actions Acquired by Ribonuclease Through Oligomerization". Current Pharmaceutical Biotechnology. 9 (3): 200–9. doi: 10.2174/138920108784567308. PMID 18673285.
- ^ Kashlan, Ossama B.; Cooperman, Barry S. (2003). "Comprehensive Model for Allosteric Regulation of Mammalian Ribonucleotide Reductase: Refinements and Consequences†". Biochemistry. 42 (6): 1696–706. doi: 10.1021/bi020634d. PMID 12578384.
- ^ Kashlan, Ossama B.; Scott, Charles P.; Lear, James D.; Cooperman, Barry S. (2002). "A Comprehensive Model for the Allosteric Regulation of Mammalian Ribonucleotide Reductase. Functional Consequences of ATP- and dATP-Induced Oligomerization of the Large Subunit†". Biochemistry. 41 (2): 462–74. doi: 10.1021/bi011653a. PMID 11781084.
- ^ Eriksson, Mathias; Uhlin, Ulla; Ramaswamy, S; Ekberg, Monica; Regnström, Karin; Sjöberg, Britt-Marie; Eklund, Hans (1997). "Binding of allosteric effectors to ribonucleotide reductase protein R1: Reduction of active-site cysteines promotes substrate binding". Structure. 5 (8): 1077–92. doi: 10.1016/S0969-2126(97)00259-1. PMID 9309223.
- ^ a b Fairman, James Wesley; Wijerathna, Sanath Ranjan; Ahmad, Md Faiz; Xu, Hai; Nakano, Ryo; Jha, Shalini; Prendergast, Jay; Welin, R Martin; et al. (2011). "Structural basis for allosteric regulation of human ribonucleotide reductase by nucleotide-induced oligomerization". Nature Structural & Molecular Biology. 18 (3): 316–22. doi: 10.1038/nsmb.2007. PMC 3101628. PMID 21336276.
- ^ a b Hohman, R.J.; Guitton, M.C.; Véron, M. (1984). "Purification of S-adenosyl-l-homocysteine hydrolase from Dictyostelium discoideum: Reversible inactivation by cAMP and 2′-deoxyadenosine". Archives of Biochemistry and Biophysics. 233 (2): 785–95. doi: 10.1016/0003-9861(84)90507-1. PMID 6091559.
- ^ Guranowski, Andrzej; Pawelkiewicz, Jerzy (1977). "Adenosylhomocysteinase from Yellow Lupin Seeds. Purification and Properties". European Journal of Biochemistry. 80 (2): 517–23. doi: 10.1111/j.1432-1033.1977.tb11907.x. PMID 923592.
- ^ Kajander, EO; Raina, AM (1981). "Affinity-chromatographic purification of S-adenosyl-L-homocysteine hydrolase. Some properties of the enzyme from rat liver". The Biochemical Journal. 193 (2): 503–12. doi: 10.1042/bj1930503. PMC 1162632. PMID 7305945.
- ^ a b c Saeki, Y; Ito, S; Shizuta, Y; Hayaishi, O; Kagamiyama, H; Wada, H (1977). "Subunit structure of biodegradative threonine deaminase". The Journal of Biological Chemistry. 252 (7): 2206–8. doi: 10.1016/S0021-9258(17)40542-4. PMID 321452.
- ^ a b c Phillips, A.T.; Wood, W.A. (1964). "Basis for AMP activation of "Biodegradative" threonine dehydrase from". Biochemical and Biophysical Research Communications. 15 (6): 530–535. doi: 10.1016/0006-291X(64)90499-1.
- ^ a b c Gerlt, JA; Rabinowitz, KW; Dunne, CP; Wood, WA (1973). "The mechanism of action of 5'-adenylic acid-activated threonine dehydrase. V. Relation between ligand-induced allosteric activation and the protomeroligomer interconversion". The Journal of Biological Chemistry. 248 (23): 8200–6. doi: 10.1016/S0021-9258(19)43214-6. PMID 4584826.
- ^ Addington, Adele K.; Johnson, David A. (1996). "Inactivation of Human Lung Tryptase: Evidence for a Re-Activatable Tetrameric Intermediate and Active Monomers". Biochemistry. 35 (42): 13511–8. doi: 10.1021/bi960042t. PMID 8885830.
- ^ Fajardo, Ignacio; Pejler, Gunnar (2003). "Formation of active monomers from tetrameric human β-tryptase". Biochemical Journal. 369 (3): 603–10. doi: 10.1042/BJ20021418. PMC 1223112. PMID 12387726.
- ^ Fukuoka, Yoshihiro; Schwartz, Lawrence B. (2004). "Human β-Tryptase: Detection and Characterization of the Active Monomer and Prevention of Tetramer Reconstitution by Protease Inhibitors". Biochemistry. 43 (33): 10757–64. doi: 10.1021/bi049486c. PMID 15311937.
- ^ Fukuoka, Y; Schwartz, LB (2006). "The B12 anti-tryptase monoclonal antibody disrupts the tetrameric structure of heparin-stabilized beta-tryptase to form monomers that are inactive at neutral pH and active at acidic pH". Journal of Immunology. 176 (5): 3165–72. doi: 10.4049/jimmunol.176.5.3165. PMC 1810230. PMID 16493076.
- ^ Fukuoka, Yoshihiro; Schwartz, Lawrence B. (2007). "Active monomers of human β-tryptase have expanded substrate specificities". International Immunopharmacology. 7 (14): 1900–8. doi: 10.1016/j.intimp.2007.07.007. PMC 2278033. PMID 18039527.
- ^ Hallgren, J.; Spillmann, D; Pejler, G (2001). "Structural Requirements and Mechanism for Heparin-induced Activation of a Recombinant Mouse Mast Cell Tryptase, Mouse Mast Cell Protease-6. FORMATION OF ACTIVE TRYPTASE MONOMERS IN THE PRESENCE OF LOW MOLECULAR WEIGHT HEPARIN". Journal of Biological Chemistry. 276 (46): 42774–81. doi: 10.1074/jbc.M105531200. PMID 11533057.
- ^ Schechter, Norman M.; Choi, Eun-Jung; Selwood, Trevor; McCaslin, Darrell R. (2007). "Characterization of Three Distinct Catalytic Forms of Human Tryptase-β: Their Interrelationships and Relevance". Biochemistry. 46 (33): 9615–29. doi: 10.1021/bi7004625. PMID 17655281.
- ^ Schechter, Norman M.; Eng, Grace Y.; Selwood, Trevor; McCaslin, Darrell R. (1995). "Structural Changes Associated with the Spontaneous Inactivation of the Serine Proteinase Human Tryptase". Biochemistry. 34 (33): 10628–38. doi: 10.1021/bi00033a038. PMID 7654717.
- ^ Schwartz, Lawrence B. (1994). "[6] Tryptase: A mast cell serine protease". Proteolytic Enzymes: Serine and Cysteine Peptidases. Methods in Enzymology. Vol. 244. pp. 88–100. doi: 10.1016/0076-6879(94)44008-5. ISBN 978-0-12-182145-6. PMID 7845247.
- ^ Strik, Merel C. M.; Wolbink, Angela; Wouters, Dorine; Bladergroen, Bellinda A.; Verlaan, Angelique R.; van Houdt, Inge S.; Hijlkema, Sanne; Hack, C. Erik; et al. (2004). "Intracellular serpin SERPINB6 (PI6) is abundantly expressed by human mast cells and forms complexes with β-tryptase monomers". Blood. 103 (7): 2710–7. doi: 10.1182/blood-2003-08-2981. PMID 14670919.
- ^ a b Kozik, Andrzej; Potempa, Jan; Travis, James (1998). "Spontaneous inactivation of human lung tryptase as probed by size-exclusion chromatography and chemical cross-linking: Dissociation of active tetrameric enzyme into inactive monomers is the primary event of the entire process". Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology. 1385 (1): 139–48. doi: 10.1016/S0167-4838(98)00053-3. PMID 9630576.
- ^ Alzani, R.; Cozzi, E.; Corti, A.; Temponi, M.; Trizio, D.; Gigli, M.; Rizzo, V. (1995). "Mechanism of suramin-induced deoligomerization of tumor necrosis factor .alpha". Biochemistry. 34 (19): 6344–50. doi: 10.1021/bi00019a012. PMID 7756262.
- ^ Corti, A; Fassina, G; Marcucci, F; Barbanti, E; Cassani, G (1992). "Oligomeric tumour necrosis factor alpha slowly converts into inactive forms at bioactive levels". The Biochemical Journal. 284 (3): 905–10. doi: 10.1042/bj2840905. PMC 1132625. PMID 1622406.
- ^ Hlodan, Roman; Pain, Roger H. (1995). "The Folding and Assembly Pathway of Tumour Necrosis Factor TNFalpha, a Globular Trimeric Protein". European Journal of Biochemistry. 231 (2): 381–7. doi: 10.1111/j.1432-1033.1995.0381e.x. PMID 7635149.
- ^ a b c d Jensen, Kaj Frank; Mygind, Bente (1996). "Different Oligomeric States are Involved in the Allosteric Behavior of Uracil Phosphoribosyltransferase from Escherichia Coli". European Journal of Biochemistry. 240 (3): 637–45. doi: 10.1111/j.1432-1033.1996.0637h.x. PMID 8856065.