| Flock House virus | |
|---|---|
|
Virus classification
| |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Kitrinoviricota |
| Class: | Magsaviricetes |
| Order: | Nodamuvirales |
| Family: | Nodaviridae |
| Genus: | Alphanodavirus |
| Species: | Flock House virus
|
Flock House virus (FHV) is in the Alphanodavirus genus of the Nodaviridae family of viruses. Flock House virus was isolated from a grass grub ( Costelytra zealandica) at the Flock House research station in Bulls, New Zealand. FHV is an extensively studied virus and is considered a model system for the study of other non-enveloped RNA viruses owing to its small size and genetic tractability, particularly to study the role of the transiently exposed hydrophobic gamma peptide and the metastability of the viral capsid. [1] [2] FHV can be engineered in insect cell culture allowing for the tailored production of native or mutant authentic virions or virus-like-particles. FHV is a platform for nanotechnology and nanomedicine, for example, for epitope display and vaccine development. [3] Viral entry into host cells occurs via receptor-mediated endocytosis. [4] Receptor binding initiates a sequence of events during which the virus exploits the host environment in order to deliver the viral cargo in to the host cytosol. Receptor binding prompts the meta-stability of the capsid–proteins, the coordinated rearrangements of which are crucial for subsequent steps in the infection pathway. In addition, the transient exposure of a covalently-independent hydrophobic γ-peptide is responsible for breaching cellular membranes and is thus essential for the viral entry of FHV into host cells. [5]
Genome
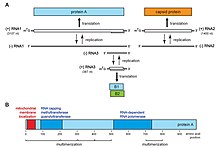
Flock House virus is a small, non-enveloped, icosahedral T=3 insect virus containing a bipartite positive sense ssRNA genome comprising two genes: RNA1 (3.1kb) an RNA2 (1.4kb). RNA1 encodes the RNA-dependent RNA polymerase and also contains a frame-shifted subgenomic RNA 3 (369 nts) that encodes protein B2, responsible for inhibition of RNAi pathways. [6] RNA2 encodes the capsid precursor, alpha, of which 180 copies form the viral capsid of FHV. Upon maturation, alpha undergoes an autocatalytic cleavage in its C-terminus to form beta, forming the main structural capsid component, and gamma, a short hydrophobic peptide required for endosome penetration that remains associated with the viral capsid. Virus-Like-Particles (VLPs) of FHV spontaneously form in S. frugiperda cell lines (e.g. Sf21) when RNA2 is expressed from a baculovirus vector and package cellular RNAs. [7] [8]
Range
FHV was originally isolated from New Zealand grass grubs ( Costelytra zealandica) in the former Flock House agricultural facility in Bulls, Ragnitikei, New Zealand. [9] Isolates were passaged in Drosophila cells in culture, which were subsequently shown to exhibit cell-death (cytopathic effect). FHV can also infect live flies. [10] FHV has been shown to infect medically important genera of insects: mosquitos, e.g. Anopheles gambiae; the tsetse fly; and the Chagas vector, Rhodnius prolixus Stal. [11] [12] Infection of these organisms by FHV has been demonstrated to have similar characteristics in terms of viral titre, virus dissemination and mortality as has been shown for fruit fly infections.[ citation needed]
Capsid structure
The structure and biophysical properties of authentic virions of FHV and of virus-like-particles (VLPs) have been extensively studied.[ citation needed]
Other studies
FHV has provided a model system for the study of the emergence and evolution of defective-interfering RNAs (DI-RNAs).[ citation needed]
References
-
^ Odegard, A; Banerjee, M; Johnson, JE (2010). Flock house virus: a model system for understanding non-enveloped virus entry and membrane penetration. Vol. 343. pp. 1–22.
doi:
10.1007/82_2010_35.
ISBN
978-3-642-13331-2.
PMID
20407886.
{{ cite book}}:|journal=ignored ( help) - ^ Banerjee, M; Johnson, JE (February 2008). "Activation, exposure and penetration of virally encoded, membrane-active polypeptides during non-enveloped virus entry". Current Protein & Peptide Science. 9 (1): 16–27. doi: 10.2174/138920308783565732. PMID 18336320.
- ^ Destito, G; Schneemann, A; Manchester, M (2009). "Biomedical Nanotechnology Using Virus-Based Nanoparticles". Viruses and Nanotechnology. Current Topics in Microbiology and Immunology. Vol. 327. pp. 95–122. doi: 10.1007/978-3-540-69379-6_5. ISBN 978-3-540-69376-5. PMID 19198572.
- ^ Odegard, AL; Kwan, MH; Walukiewicz, HE; Banerjee, M; Schneemann, A; Johnson, JE (September 2009). "Low endocytic pH and capsid protein autocleavage are critical components of Flock House virus cell entry". Journal of Virology. 83 (17): 8628–37. doi: 10.1128/JVI.00873-09. PMC 2738175. PMID 19553341.
- ^ Schneemann, A; Zhong, W; Gallagher, T. M; Rueckert, R. R (1992). "Maturation cleavage required for infectivity of a nodavirus". Journal of Virology. 66 (11): 6728–34. doi: 10.1128/JVI.66.11.6728-6734.1992. PMC 240169. PMID 1404613.
- ^ Chao, JA; Lee, JH; Chapados, BR; Debler, EW; Schneemann, A; Williamson, JR (November 2005). "Dual modes of RNA-silencing suppression by Flock House virus protein B2". Nature Structural & Molecular Biology. 12 (11): 952–7. doi: 10.1038/nsmb1005. PMID 16228003. S2CID 37878393.
- ^ Schneemann, A; Dasgupta, R; Johnson, JE; Rueckert, RR (May 1993). "Use of recombinant baculoviruses in synthesis of morphologically distinct viruslike particles of flock house virus, a nodavirus". Journal of Virology. 67 (5): 2756–63. doi: 10.1128/JVI.67.5.2756-2763.1993. PMC 237599. PMID 8474173.
- ^ Routh, A; Domitrovic, T; Johnson, JE (7 February 2012). "Host RNAs, including transposons, are encapsidated by a eukaryotic single-stranded RNA virus". Proceedings of the National Academy of Sciences of the United States of America. 109 (6): 1907–12. Bibcode: 2012PNAS..109.1907R. doi: 10.1073/pnas.1116168109. PMC 3277583. PMID 22308402.
- ^ Scotti, PD; Dearing, S; Mossop, DW (1983). "Flock House virus: a nodavirus isolated from Costelytra zealandica (White) (Coleoptera: Scarabaeidae)". Archives of Virology. 75 (3): 181–9. doi: 10.1007/BF01315272. PMID 6188442. S2CID 9853542.
- ^ Goic, B; Vodovar, N; Mondotte, JA; Monot, C; Frangeul, L; Blanc, H; Gausson, V; Vera-Otarola, J; Cristofari, G; Saleh, MC (April 2013). "RNA-mediated interference and reverse transcription control the persistence of RNA viruses in the insect model Drosophila" (PDF). Nature Immunology. 14 (4): 396–403. doi: 10.1038/ni.2542. PMID 23435119. S2CID 6690552.
- ^ Dasgupta, R; Cheng, LL; Bartholomay, LC; Christensen, BM (July 2003). "Flock house virus replicates and expresses green fluorescent protein in mosquitoes". The Journal of General Virology. 84 (Pt 7): 1789–97. doi: 10.1099/vir.0.18938-0. PMID 12810873.
- ^ Dasgupta, R; Free, HM; Zietlow, SL; Paskewitz, SM; Aksoy, S; Shi, L; Fuchs, J; Hu, C; Christensen, BM (January 2007). "Replication of flock house virus in three genera of medically important insects". Journal of Medical Entomology. 44 (1): 102–10. doi: 10.1603/0022-2585(2007)44[102:rofhvi]2.0.co;2. PMID 17294927.