
Size of this preview:
633 × 599 pixels. Other resolutions:
254 × 240 pixels |
507 × 480 pixels |
811 × 768 pixels |
1,082 × 1,024 pixels |
2,164 × 2,048 pixels |
3,160 × 2,991 pixels.
Original file (3,160 × 2,991 pixels, file size: 654 KB, MIME type: image/jpeg)
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 15:26, 5 December 2021 |
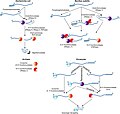 | 3,160 × 2,991 (654 KB) | Guest2625 | Uploaded a work by Katarzyna J. Bandyra, Marie Bouvier, Agamemnon J. Carpousis, and Ben F. Luisi from Katarzyna J. Bandyra, Marie Bouvier, Agamemnon J. Carpousis, Ben F. Luisi, The social fabric of the RNA degradosome, Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms, Volume 1829, Issues 6–7, 2013, Pages 514-522, ISSN 1874-9399, https://doi.org/10.1016/j.bbagrm.2013.02.011 with UploadWizard |
File usage
The following pages on the English Wikipedia use this file (pages on other projects are not listed):
Global file usage
The following other wikis use this file:
- Usage on ja.wikipedia.org
- Usage on ms.wikipedia.org