
Size of this PNG preview of this SVG file:
472 × 203 pixels. Other resolutions:
320 × 138 pixels |
640 × 275 pixels |
1,024 × 440 pixels |
1,280 × 551 pixels |
2,560 × 1,101 pixels.
Original file (SVG file, nominally 472 × 203 pixels, file size: 15 KB)
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 17:41, 12 April 2015 |
 | 472 × 203 (15 KB) | Krishnavedala | align first molecule with the rest |
| 16:55, 12 April 2015 |
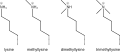
| 430 × 174 (15 KB) | Krishnavedala | {{Information |Description ={{en|1=Methyl lysine}} |Source ={{own}} |Author = Krishnavedala |Date =2015-04-12 |Permission ={{PD-chem}} |other_versions = }} {{Igen|O|v|n={{w|LaTeX}}}} [[Category:Lys... |
File usage
The following pages on the English Wikipedia use this file (pages on other projects are not listed):
Global file usage
The following other wikis use this file:
- Usage on el.wikipedia.org
- Usage on fr.wikipedia.org
- Usage on ms.wikipedia.org
- Usage on uk.wikipedia.org
