The Golgi matrix is a collection of proteins involved in the structure and function of the Golgi apparatus. [1] [2] [3] The matrix was first isolated in 1994 as an amorphous collection of 12 proteins that remained associated together in the presence of detergent (which removed Golgi membranes) and 150 m M NaCl (which removed weakly associated proteins). [4] Treatment with a protease enzyme removed the matrix, which confirmed the importance of proteins for the matrix structure. [4] Modern freeze etch [5] electron microscopy (EM) clearly shows a mesh connecting Golgi cisternae and associated vesicles. [6] [7] Further support for the existence of a matrix comes from EM images showing that ribosomes are excluded from regions between and near Golgi cisternae. [8] [9] [10] [11] [12] [13]


Structure and function

The first individual protein component of the matrix was identified in 1995 as Golgin A2 (then called GM130). [14] Since then, many other golgin family proteins have been found to be in the Golgi matrix [2] and are associated with the Golgi membranes in a variety of ways. [15] [1] For example, GMAP210 (Golgi Microtubule Associated Protein 210) has an ALPS (Amphipathic Lipid-Packing Sensor) motif in the N-termal 38 amino acids and an ARF1-binding domain called GRAB (Grip-Related Arf-Binding) at the C-terminus. [16] Thus, the GRAB-domain can bind indirectly to Golgi cisternae and its ALPS motif can tether vesicles. [17] Golgins have coiled-coil domains and are thus predicted to have elongated structures [2] up to 200 nm in length. [18] Most are peripheral membrane proteins attached at one end to Golgi membranes. [2] They have flexible regions between the coiled-coil domains, which make them ideal candidates for mediating the dynamic vesicle docking to Golgi cisternae and dynamic structure of the Golgi itself. [2]
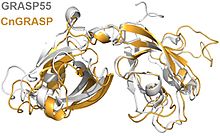
Golgi reassembly-stacking proteins are an evolutionarily conserved family of proteins in the Golgi matrix. [2] GRASP65 and GRASP55 are the 2 human GRASPs. These proteins were named from their requirement for accurate Golgi reassembly during an in vitro assay, [2] but they have also been shown to function in vivo, as shown in the accompanying figure. [19] GRASPs associate with lipid bilayers because they are myristoylated and their myristic acid residue intercalates into the lipid layer. [7] Their trans oligomerization is controlled by phosphorylation [6] and is thought to explain the fragmentation of the Golgi as required during mitosis. [7]
Components
Disease associations
- CG-NAP hereditary Long QT syndrome LQT11 [20]
- COH1 Cohen Syndrome [21]
- GMAP210 Achondrogenesis type IA [22]
- Golgin A2 a complex, neuromuscular disorder [23]
- GORAB Gerodermia osteodysplastica [24]
References
- ^ a b Short B, Haas A, Barr FA (2005). "Golgins and GTPases, giving identity and structure to the Golgi apparatus". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research. 1744 (3): 383–95. doi: 10.1016/j.bbamcr.2005.02.001. PMID 15979508.
- ^ a b c d e f g Xiang Y, Wang Y (2011). "New components of the Golgi matrix". Cell and Tissue Research. 344 (3): 365–79. doi: 10.1007/s00441-011-1166-x. PMC 3278855. PMID 21494806.
- ^ Lowe, M (2011). "Structural organization of the Golgi apparatus". Current Opinion in Cell Biology. 23 (1): 85–93. doi: 10.1016/j.ceb.2010.10.004. PMID 21071196.
- ^ a b Slusarewicz P, Nilsson T, Hui N, Watson R, Warren G (1994). "Isolation of a matrix that binds medial Golgi enzymes". The Journal of Cell Biology. 124 (4): 405–13. doi: 10.1083/jcb.124.4.405. PMC 2119912. PMID 8106542.
- ^ Heuser JE (2011). "The origins and evolution of freeze-etch electron microscopy". Journal of Electron Microscopy. 60 (Suppl 1): S3–29. doi: 10.1093/jmicro/dfr044. PMC 3202940. PMID 21844598.
- ^
a
b Zhang, X. and Wang, Y.
"Golgi structure and the role of GRASP65 in Golgi stack formation". Retrieved 27 May 2017.
{{ cite web}}: CS1 maint: multiple names: authors list ( link) - ^
a
b
c Zhang, X. and Wang, Y., Front Cell Dev Biol. 2015; 3: 84. Published online 2016 Jan 6. doi: 10.3389/fcell.2015.00084 (2015).
"GRASPs in Golgi Structure and Function". Frontiers in Cell and Developmental Biology. 3: 84.
doi:
10.3389/fcell.2015.00084.
PMC
4701983.
PMID
26779480.
{{ cite journal}}: CS1 maint: multiple names: authors list ( link) CS1 maint: numeric names: authors list ( link) - ^ Fig. 14 in Mogelsvang S, Gomez-Ospina N, Soderholm J, Glick BS, Staehelin LA (2003). "Tomographic Evidence for Continuous Turnover of Golgi Cisternae in Pichia pastoris". Molecular Biology of the Cell. 14 (6): 2277–91. doi: 10.1091/mbc.e02-10-0697. PMC 260745. PMID 12808029.
- ^ Staehelin LA; Kang BH. "Electron tomographic model of a Golgi stack and its encompassing, ribosome-excluding scaffold (Golgi matrix)". plantphysiol.org. American Society of Plant Biologists. Retrieved 27 May 2017.
- ^ Staehelin LA; Kang BH. "Transfer of COPII vesicles and their scaffolds to the cis-Golgi matrix". plantphysiol.org. American Society of Plant Biologists. Retrieved 27 May 2017.
- ^ Lucocq JM, Pryde JG, Berger EG, Warren G (1987). "A mitotic form of the Golgi apparatus in HeLa cells". The Journal of Cell Biology. 104 (4): 865–74. doi: 10.1083/jcb.104.4.865. PMC 2114436. PMID 3104351.
- ^ Mogelsvang S, Gomez-Ospina N, Soderholm J, Glick BS, Staehelin LA (2003). "Tomographic evidence for continuous turnover of Golgi cisternae in Pichia pastoris". Molecular Biology of the Cell. 14 (6): 2277–91. doi: 10.1091/mbc.E02-10-0697. PMC 260745. PMID 12808029.
- ^ Staehelin LA, Kang BH (2008). "Nanoscale architecture of endoplasmic reticulum export sites and of Golgi membranes as determined by electron tomography". Plant Physiology. 147 (4): 1454–68. doi: 10.1104/pp.108.120618. PMC 2492626. PMID 18678738.
- ^ Nakamura N, Rabouille C, Watson R, Nilsson T, Hui N, Slusarewicz P, Kreis TE, Warren G (1995). "Characterization of a cis-Golgi matrix protein, GM130". The Journal of Cell Biology. 131 (6 Pt 2): 1715–26. doi: 10.1083/jcb.131.6.1715. PMC 2120691. PMID 8557739.
-
^ Benjamin Short, Alexander Haas, Francis A. Barr.
"Golgins associate with Golgi membranes in a variety of ways". ars.els-cdn.com/. Biochimica et Biophysica Acta (BBA) - Molecular Cell Research. Retrieved 31 May 2017.
{{ cite web}}: CS1 maint: multiple names: authors list ( link) - ^ Cardenas J, Rivero S, Goud B, Bornens M, Rios RM (2009). "Golgi localisation of GMAP210 requires two distinct cis-membrane binding mechanisms". BMC Biology. 7: 56. doi: 10.1186/1741-7007-7-56. PMC 2744908. PMID 19715559.
- ^ Doucet CM, Esmery N, de Saint-Jean M, Antonny B (2015). "Membrane Curvature Sensing by Amphipathic Helices Is Modulated by the Surrounding Protein Backbone". PLOS ONE. 10 (9): e0137965. Bibcode: 2015PLoSO..1037965D. doi: 10.1371/journal.pone.0137965. PMC 4569407. PMID 26366573.
- ^ Drin G, Morello V, Casella JF, Gounon P, Antonny B (2008). "Asymmetric tethering of flat and curved lipid membranes by a golgin". Science. 320 (5876): 670–3. Bibcode: 2008Sci...320..670D. doi: 10.1126/science.1155821. PMID 18451304. S2CID 6619427.
- ^ Wang Y, Wei JH, Bisel B, Tang D, Seemann J (2008). "Golgi cisternal unstacking stimulates COPI vesicle budding and protein transport". PLOS ONE. 3 (2): e1647. Bibcode: 2008PLoSO...3.1647W. doi: 10.1371/journal.pone.0001647. PMC 2249924. PMID 18297130.
- ^ Chen L, Marquardt ML, Tester DJ, Sampson KJ, Ackerman MJ, Kass RS (2007). "Mutation of an A-kinase-anchoring protein causes long-QT syndrome". Proceedings of the National Academy of Sciences of the United States of America. 104 (52): 20990–5. Bibcode: 2007PNAS..10420990C. doi: 10.1073/pnas.0710527105. PMC 2409254. PMID 18093912.
- ^ Kolehmainen J, Black GC, Saarinen A, et al. (2003). "Cohen syndrome is caused by mutations in a novel gene, COH1, encoding a transmembrane protein with a presumed role in vesicle-mediated sorting and intracellular protein transport". Am. J. Hum. Genet. 72 (6): 1359–69. doi: 10.1086/375454. PMC 1180298. PMID 12730828.
- ^ Smits P, Bolton AD, Funari V, Hong M, Boyden ED, Lu L, Manning DK, Dwyer ND, Moran JL, Prysak M, Merriman B, Nelson SF, Bonafé L, Superti-Furga A, Ikegawa S, Krakow D, Cohn DH, Kirchhausen T, Warman ML, Beier DR (2010). "Lethal skeletal dysplasia in mice and humans lacking the golgin GMAP-210". The New England Journal of Medicine. 362 (3): 206–16. doi: 10.1056/NEJMoa0900158. PMC 3108191. PMID 20089971.
- ^ Shamseldin HE, Bennett AH, Alfadhel M, Gupta V, Alkuraya FS (2016). "GOLGA2, encoding a master regulator of golgi apparatus, is mutated in a patient with a neuromuscular disorder". Human Genetics. 135 (2): 245–51. doi: 10.1007/s00439-015-1632-8. PMC 4975006. PMID 26742501.
- ^ Hennies HC, Kornak U, Zhang H, et al. (December 2008). "Gerodermia osteodysplastica is caused by mutations in SCL1BP1, a Rab-6 interacting golgin". Nat. Genet. 40 (12): 1410–2. doi: 10.1038/ng.252. PMC 3122266. PMID 18997784.