
Cryogenic electron microscopy (cryo-EM) is a cryomicroscopy technique applied on samples cooled to cryogenic temperatures. For biological specimens, the structure is preserved by embedding in an environment of vitreous ice. An aqueous sample solution is applied to a grid-mesh and plunge-frozen in liquid ethane or a mixture of liquid ethane and propane. [1] While development of the technique began in the 1970s, recent advances in detector technology and software algorithms have allowed for the determination of biomolecular structures at near-atomic resolution. [2] This has attracted wide attention to the approach as an alternative to X-ray crystallography or NMR spectroscopy for macromolecular structure determination without the need for crystallization. [3]
In 2017, the Nobel Prize in Chemistry was awarded to Jacques Dubochet, Joachim Frank, and Richard Henderson "for developing cryo-electron microscopy for the high-resolution structure determination of biomolecules in solution." [4] Nature Methods also named cryo-EM as the "Method of the Year" in 2015. [5]
History
Early development
In the 1960s, the use of transmission electron microscopy for structure determination methods was limited because of the radiation damage due to high energy electron beams. Scientists hypothesized that examining specimens at low temperatures would reduce beam-induced radiation damage. [6] Both liquid helium (−269 °C or 4 K or −452.2 °F) and liquid nitrogen (−195.79 °C or 77 K or −320 °F) were considered as cryogens. In 1980, Erwin Knapek and Jacques Dubochet published comments on beam damage at cryogenic temperatures sharing observations that:
Thin crystals mounted on carbon film were found to be from 30 to 300 times more beam-resistant at 4 K than at room temperature... Most of our results can be explained by assuming that cryoprotection in the region of 4 K is strongly dependent on the temperature. [7]
However, these results were not reproducible and amendments were published in Nature just two years later informing that the beam resistance was less significant than initially anticipated. The protection gained at 4 K was closer to "tenfold for standard samples of L- valine", [8] than what was previously stated.
In 1981, Alasdair McDowall and Jacques Dubochet, scientists at the European Molecular Biology Laboratory, reported the first successful implementation of cryo-EM. [9] McDowall and Dubochet vitrified pure water in a thin film by spraying it onto a hydrophilic carbon film that was rapidly plunged into cryogen (liquid propane or liquid ethane cooled to 77 K). The thin layer of amorphous ice was less than 1 µm thick and an electron diffraction pattern confirmed the presence of amorphous/vitreous ice. In 1984, Dubochet's group demonstrated the power of cryo-EM in structural biology with analysis of vitrified adenovirus type 2, T4 bacteriophage, Semliki Forest virus, Bacteriophage CbK, and Vesicular-Stomatitis-Virus. [10]
Recent advancements
The 2010s were marked with drastic advancements of electron cameras. Notably, the improvements made to direct electron detectors have led to a "resolution revolution" [11] pushing the resolution barrier beneath the crucial ~2-3 Å limit to resolve amino acid position and orientation. [12]
Henderson ( MRC Laboratory of Molecular Biology, Cambridge, UK) formed a consortium with engineers at the Rutherford Appleton Laboratory and scientists at the Max Planck Society to fund and develop a first prototype. The consortium then joined forces with the electron microscope manufacturer FEI to roll out and market the new design. At about the same time, Gatan Inc. of Pleasanton, California came out with a similar detector designed by Peter Denes ( Lawrence Berkeley National Laboratory) and David Agard ( University of California, San Francisco). A third type of camera was developed by Nguyen-Huu Xuong at the Direct Electron company ( San Diego, California). [11]
More recently, advancements in the use of protein-based imaging scaffolds are helping to solve the problems of sample orientation bias and size limit. Proteins smaller than ~50 kDa generally have insufficient SNR to be able to resolve protein particles in the image, making 3D reconstruction difficult or impossible. [13] Imaging scaffolds boost the SNR of smaller proteins by binding them to a larger object, the scaffold. The Yeates group at UCLA were able to create a clearer image of three variants of KRAS (roughly 19 kDa in size) by utilising a rigidified imaging scaffold, and using DARPins as modular binding domain between the scaffold and the protein-of-interest. [14]
2017 Nobel Prize in Chemistry
In recognition of the impact cryo-EM has had on biochemistry, three scientists, Jacques Dubochet, Joachim Frank and Richard Henderson, were awarded the Nobel Prize in Chemistry "for developing cryo-electron microscopy for the high-resolution structure determination of biomolecules in solution." [4]
Comparisons to X-ray crystallography
Traditionally, X-ray crystallography has been the most popular technique for determining the 3D structures of biological molecules. [15] However, the aforementioned improvements in cryo-EM have increased its popularity as a tool for examining the details of biological molecules. Since 2010, yearly cryo-EM structure deposits have outpaced X-ray crystallography. [16] Though X-ray crystallography has drastically more total deposits due to a decades-longer history, total deposits of the two methods are projected to eclipse around 2035. [16]
The resolution of X-ray crystallography is limited by crystal homogeneity, [17] and coaxing biological molecules with unknown ideal crystallization conditions into a crystalline state can be very time-consuming, in extreme cases taking months or even years. [18] To contrast, sample preparation in cryo-EM may require several rounds of screening and optimization to overcome issues such as protein aggregation and preferred orientations, [19] [20] but it does not require the sample to form a crystal, rather samples for cryo-EM are flash-frozen and examined in their near-native states. [21]
According to Proteopedia, the median resolution achieved by X-ray crystallography (as of May 19, 2019) on the Protein Data Bank is 2.05 Å, [17] and the highest resolution achieved on record (as of September 30, 2022) is 0.48 Å. [22] As of 2020, the majority of the protein structures determined by cryo-EM are at a lower resolution of 3–4 Å. [23] However, as of 2020, the best cryo-EM resolution has been recorded at 1.22 Å, [20] making it a competitor in resolution in some cases.
Correlative light cryo-TEM and cryo-ET
In 2019, correlative light cryo-TEM and cryo-ET were used to observe tunnelling nanotubes (TNTs) in neuronal cells. [24]
Scanning electron cryomicroscopy
Scanning electron cryomicroscopy (cryoSEM) is a scanning electron microscopy technique with a scanning electron microscope's cold stage in a cryogenic chamber.
Cryogenic transmission electron microscopy
Cryogenic transmission electron microscopy (cryo-TEM) is a transmission electron microscopy technique that is used in structural biology and materials science. Colloquially, the term "cryogenic electron microscopy" or its shortening "cryo-EM" refers to cryogenic transmission electron microscopy by default, as the vast majority of cryo-EM is done in transmission electron microscopes, rather than scanning electron microscopes.
Centers
The Federal Institute of Technology, the University of Lausanne and the University of Geneva opened the Dubochet Center For Imaging (DCI) at the end of November 2021, for the purposes of applying and further developing cryo-EM. [25] Less than a month after the first identification of the SARS-CoV-2 Omicron variant, researchers at the DCI were able to define its structure, identify the crucial mutations to circumvent individual vaccines and provide insights for new therapeutic approaches. [26]
The Danish National cryo-EM Facility also known as EMBION was inaugurated on December 1, 2016. EMBION is a cryo-EM consortium between Danish Universities (Aarhus University host and University of Copenhagen co-host).

Advanced methods
- Cryogenic electron tomography (cryo-ET), a specialized application where many images are taken of individual samples at various tilt angles, resulting in a 3D reconstruction of a single sample. [27]
- Electron crystallography, method to determine the arrangement of atoms in solids using a TEM
- MicroED, [28] method to determine the structure of proteins, peptides, organic molecules, and inorganic compounds using electron diffraction from 3D crystals [29] [30] [31]
- Single particle analysis cryo-EM, an averaging method to determine protein structure from monodisperse samples. [32]
-
Structure of alcohol oxidase from Pichia pastoris by Cryo-EM
-
Cryo-EM image of an intact ARMAN cell from an Iron Mountain biofilm. Image width is 576 nm.
See also
References
- ^ Tivol WF, Briegel A, Jensen GJ (October 2008). "An improved cryogen for plunge freezing". Microscopy and Microanalysis. 14 (5): 375–379. Bibcode: 2008MiMic..14..375T. doi: 10.1017/S1431927608080781. PMC 3058946. PMID 18793481.
- ^ Cheng Y, Grigorieff N, Penczek PA, Walz T (April 2015). "A primer to single-particle cryo-electron microscopy". Cell. 161 (3): 438–449. doi: 10.1016/j.cell.2015.03.050. PMC 4409659. PMID 25910204.
- ^ Stoddart C (1 March 2022). "Structural biology: How proteins got their close-up". Knowable Magazine. doi: 10.1146/knowable-022822-1. S2CID 247206999. Retrieved 25 March 2022.
- ^ a b "The Nobel Prize in Chemistry 2017". NobelPrize.org. Retrieved 2022-09-30.
- ^ Doerr A (January 2017). "Cryo-electron tomography". Nature Methods. 14 (1): 34. doi: 10.1038/nmeth.4115. ISSN 1548-7091. S2CID 27162203.
- ^ Dubochet J, Knapek E (April 2018). "Ups and downs in early electron cryo-microscopy". PLOS Biology. 16 (4): e2005550. doi: 10.1371/journal.pbio.2005550. PMC 5929567. PMID 29672565.
- ^ Knapek E, Dubochet J (August 1980). "Beam damage to organic material is considerably reduced in cryo-electron microscopy". Journal of Molecular Biology. 141 (2): 147–161. doi: 10.1016/0022-2836(80)90382-4. PMID 7441748.
- ^ Newmark P (30 September 1982). "Cryo-transmission microscopy Fading hopes". Nature. 299 (5882): 386–387. Bibcode: 1982Natur.299..386N. doi: 10.1038/299386c0.
- ^ Dubochet J, McDowall AW (December 1981). "Vitrification of Pure Water for Electron Microscopy". Journal of Microscopy. 124 (3): 3–4. doi: 10.1111/j.1365-2818.1981.tb02483.x.
- ^ Adrian M, Dubochet J, Lepault J, McDowall AW (March 1984). "Cryo-electron microscopy of viruses". Nature. 308 (5954): 32–36. Bibcode: 1984Natur.308...32A. doi: 10.1038/308032a0. PMID 6322001. S2CID 4319199.
- ^ a b Kühlbrandt, Werner (2014-03-28). "The Resolution Revolution". Science. 343 (6178): 1443–1444. Bibcode: 2014Sci...343.1443K. doi: 10.1126/science.1251652. ISSN 0036-8075. PMID 24675944. S2CID 35524447.
- ^ Kuster, Daniel J.; Liu, Chengyu; Fang, Zheng; Ponder, Jay W.; Marshall, Garland R. (2015-04-20). "High-Resolution Crystal Structures of Protein Helices Reconciled with Three-Centered Hydrogen Bonds and Multipole Electrostatics". PLOS ONE. 10 (4): e0123146. Bibcode: 2015PLoSO..1023146K. doi: 10.1371/journal.pone.0123146. ISSN 1932-6203. PMC 4403875. PMID 25894612.
- ^ Herzik, Mark A.; Wu, Mengyu; Lander, Gabriel C. (2019-03-04). "High-resolution structure determination of sub-100 kDa complexes using conventional cryo-EM". Nature Communications. 10 (1): 1032. Bibcode: 2019NatCo..10.1032H. doi: 10.1038/s41467-019-08991-8. ISSN 2041-1723. PMC 6399227. PMID 30833564.
- ^ Castells-Graells R, Meador K, Arbing MA, Sawaya MR, Gee M, Cascio D, et al. (September 2023). "Cryo-EM structure determination of small therapeutic protein targets at 3 Å-resolution using a rigid imaging scaffold". Proceedings of the National Academy of Sciences of the United States of America. 120 (37): e2305494120. Bibcode: 2023PNAS..12005494C. doi: 10.1073/pnas.2305494120. PMC 10500258. PMID 37669364.
- ^ Smyth MS, Martin JH (February 2000). "x ray crystallography". Molecular Pathology. 53 (1): 8–14. doi: 10.1136/mp.53.1.8. PMC 1186895. PMID 10884915.
- ^ a b Chiu, Wah; Schmid, Michael F.; Pintilie, Grigore D.; Lawson, Catherine L. (January 2021). "Evolution of standardization and dissemination of cryo-EM structures and data jointly by the community, PDB, and EMDB". Journal of Biological Chemistry. 296: 100560. doi: 10.1016/j.jbc.2021.100560. ISSN 0021-9258. PMC 8050867. PMID 33744287.
- ^ a b "Resolution - Proteopedia, life in 3D". proteopedia.org. Retrieved 2020-10-27.
- ^ Callaway E (February 2020). "Revolutionary cryo-EM is taking over structural biology". Nature. 578 (7794): 201. Bibcode: 2020Natur.578..201C. doi: 10.1038/d41586-020-00341-9. PMID 32047310.
- ^ Lyumkis, Dmitry (2019-03-29). "Challenges and opportunities in cryo-EM single-particle analysis". Journal of Biological Chemistry. 294 (13): 5181–5197. doi: 10.1074/jbc.rev118.005602. ISSN 0021-9258. PMC 6442032. PMID 30804214.
- ^ a b Nakane T, Kotecha A, Sente A, McMullan G, Masiulis S, Brown PM, et al. (November 2020). "Single-particle cryo-EM at atomic resolution". Nature. 587 (7832): 152–156. Bibcode: 2020Natur.587..152N. doi: 10.1038/s41586-020-2829-0. PMC 7611073. PMID 33087931.
- ^ Wang HW, Wang JW (January 2017). "How cryo-electron microscopy and X-ray crystallography complement each other". Protein Science. 26 (1): 32–39. doi: 10.1002/pro.3022. PMC 5192981. PMID 27543495.
- ^ Schmidt A, Teeter M, Weckert E, Lamzin VS (April 2011). "Crystal structure of small protein crambin at 0.48 Å resolution". Acta Crystallographica. Section F, Structural Biology and Crystallization Communications. 67 (Pt 4): 424–428. doi: 10.1107/S1744309110052607. PMC 3080141. PMID 21505232.
- ^ Yip KM, Fischer N, Paknia E, Chari A, Stark H (November 2020). "Atomic-resolution protein structure determination by cryo-EM". Nature. 587 (7832): 157–161. Bibcode: 2020Natur.587..157Y. doi: 10.1038/s41586-020-2833-4. PMID 33087927. S2CID 224823207.
- ^ Sartori-Rupp A, Cordero Cervantes D, Pepe A, Gousset K, Delage E, Corroyer-Dulmont S, et al. (January 2019). "Correlative cryo-electron microscopy reveals the structure of TNTs in neuronal cells". Nature Communications. 10 (1): 342. Bibcode: 2019NatCo..10..342S. doi: 10.1038/s41467-018-08178-7. PMC 6341166. PMID 30664666.
- ^ "Inauguration of the Dubochet Center for Imaging (DCI) on the campuses of UNIGE, UNIL and EPFL". unige.ch. 2021-11-30. Retrieved 2022-04-30.
- ^ "Scientists uncover Omicron variant mysteries using microscopes". swissinfo.ch. 2021-12-30. Retrieved 2022-04-30.
- ^ Bäuerlein, Felix J. B.; Baumeister, Wolfgang (2021-10-01). "Towards Visual Proteomics at High Resolution". Journal of Molecular Biology. From Protein Sequence to Structure at Warp Speed: How Alphafold Impacts Biology. 433 (20): 167187. doi: 10.1016/j.jmb.2021.167187. ISSN 0022-2836. PMID 34384780.
- ^ Nannenga BL, Shi D, Leslie AG, Gonen T (September 2014). "High-resolution structure determination by continuous-rotation data collection in MicroED". Nature Methods. 11 (9): 927–930. doi: 10.1038/nmeth.3043. PMC 4149488. PMID 25086503.
- ^ Jones CG, Martynowycz MW, Hattne J, Fulton TJ, Stoltz BM, Rodriguez JA, et al. (November 2018). "The CryoEM Method MicroED as a Powerful Tool for Small Molecule Structure Determination". ACS Central Science. 4 (11): 1587–1592. doi: 10.1021/acscentsci.8b00760. PMC 6276044. PMID 30555912.
- ^ de la Cruz MJ, Hattne J, Shi D, Seidler P, Rodriguez J, Reyes FE, et al. (February 2017). "Atomic-resolution structures from fragmented protein crystals with the cryoEM method MicroED". Nature Methods. 14 (4): 399–402. doi: 10.1038/nmeth.4178. PMC 5376236. PMID 28192420.
- ^ Gruene T, Wennmacher JT, Zaubitzer C, Holstein JJ, Heidler J, Fecteau-Lefebvre A, et al. (December 2018). "Rapid Structure Determination of Microcrystalline Molecular Compounds Using Electron Diffraction". Angewandte Chemie. 57 (50): 16313–16317. doi: 10.1002/anie.201811318. PMC 6468266. PMID 30325568.
- ^ Cheng Y (August 2018). "Single-particle cryo-EM-How did it get here and where will it go". Science. 361 (6405): 876–880. Bibcode: 2018Sci...361..876C. doi: 10.1126/science.aat4346. PMC 6460916. PMID 30166484.
- ^ Xiao, C., Fischer, M.G., Bolotaulo, D.M., Ulloa-Rondeau, N., Avila, G.A., and Suttle, C.A. (2017) "Cryo-EM reconstruction of the Cafeteria roenbergensis virus capsid suggests novel assembly pathway for giant viruses". Scientific Reports, 7: 5484. doi: 10.1038/s41598-017-05824-w.

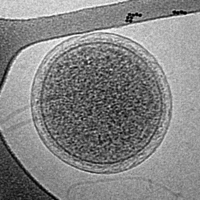
![Cryo-EM image of the CroV giant marine virus (scale bar represents 200 nm)[33]](https://upload.wikimedia.org/wikipedia/commons/thumb/6/6e/CroV_TEM_%28cropped%29.jpg/200px-CroV_TEM_%28cropped%29.jpg)