| Transcription factor glossary | |
|---|---|
| |

In molecular biology, a transcription factor (TF) (or sequence-specific DNA-binding factor) is a protein that controls the rate of transcription of genetic information from DNA to messenger RNA, by binding to a specific DNA sequence. [1] [2] The function of TFs is to regulate—turn on and off—genes in order to make sure that they are expressed in the desired cells at the right time and in the right amount throughout the life of the cell and the organism. Groups of TFs function in a coordinated fashion to direct cell division, cell growth, and cell death throughout life; cell migration and organization ( body plan) during embryonic development; and intermittently in response to signals from outside the cell, such as a hormone. There are approximately 1600 TFs in the human genome. [3] [4] [5] Transcription factors are members of the proteome as well as regulome.
TFs work alone or with other proteins in a complex, by promoting (as an activator), or blocking (as a repressor) the recruitment of RNA polymerase (the enzyme that performs the transcription of genetic information from DNA to RNA) to specific genes. [6] [7] [8]
A defining feature of TFs is that they contain at least one DNA-binding domain (DBD), which attaches to a specific sequence of DNA adjacent to the genes that they regulate. [9] [10] TFs are grouped into classes based on their DBDs. [11] [12] Other proteins such as coactivators, chromatin remodelers, histone acetyltransferases, histone deacetylases, kinases, and methylases are also essential to gene regulation, but lack DNA-binding domains, and therefore are not TFs. [13]
TFs are of interest in medicine because TF mutations can cause specific diseases, and medications can be potentially targeted toward them.
Number
Transcription factors are essential for the regulation of gene expression and are, as a consequence, found in all living organisms. The number of transcription factors found within an organism increases with genome size, and larger genomes tend to have more transcription factors per gene. [14]
There are approximately 2800 proteins in the human genome that contain DNA-binding domains, and 1600 of these are presumed to function as transcription factors, [3] though other studies indicate it to be a smaller number. [15] Therefore, approximately 10% of genes in the genome code for transcription factors, which makes this family the single largest family of human proteins. Furthermore, genes are often flanked by several binding sites for distinct transcription factors, and efficient expression of each of these genes requires the cooperative action of several different transcription factors (see, for example, hepatocyte nuclear factors). Hence, the combinatorial use of a subset of the approximately 2000 human transcription factors easily accounts for the unique regulation of each gene in the human genome during development. [13]
Mechanism
Transcription factors bind to either enhancer or promoter regions of DNA adjacent to the genes that they regulate. Depending on the transcription factor, the transcription of the adjacent gene is either up- or down-regulated. Transcription factors use a variety of mechanisms for the regulation of gene expression. [16] These mechanisms include:
- stabilize or block the binding of RNA polymerase to DNA[ citation needed]
- catalyze the
acetylation or deacetylation of
histone proteins. The transcription factor can either do this directly or recruit other proteins with this catalytic activity. Many transcription factors use one or the other of two opposing mechanisms to regulate transcription:
[17]
- histone acetyltransferase (HAT) activity – acetylates histone proteins, which weakens the association of DNA with histones, which make the DNA more accessible to transcription, thereby up-regulating transcription
- histone deacetylase (HDAC) activity – deacetylates histone proteins, which strengthens the association of DNA with histones, which make the DNA less accessible to transcription, thereby down-regulating transcription
- recruit coactivator or corepressor proteins to the transcription factor DNA complex [18]
Function
Transcription factors are one of the groups of proteins that read and interpret the genetic "blueprint" in the DNA. They bind to the DNA and help initiate a program of increased or decreased gene transcription. As such, they are vital for many important cellular processes. Below are some of the important functions and biological roles transcription factors are involved in:
Basal transcriptional regulation
In eukaryotes, an important class of transcription factors called general transcription factors (GTFs) are necessary for transcription to occur. [19] [20] [21] Many of these GTFs do not actually bind DNA, but rather are part of the large transcription preinitiation complex that interacts with RNA polymerase directly. The most common GTFs are TFIIA, TFIIB, TFIID (see also TATA binding protein), TFIIE, TFIIF, and TFIIH. [22] The preinitiation complex binds to promoter regions of DNA upstream to the gene that they regulate.
Differential enhancement of transcription
Other transcription factors differentially regulate the expression of various genes by binding to enhancer regions of DNA adjacent to regulated genes. These transcription factors are critical to making sure that genes are expressed in the right cell at the right time and in the right amount, depending on the changing requirements of the organism.[ citation needed]
Development
Many transcription factors in multicellular organisms are involved in development. [23] Responding to stimuli, these transcription factors turn on/off the transcription of the appropriate genes, which, in turn, allows for changes in cell morphology or activities needed for cell fate determination and cellular differentiation. The Hox transcription factor family, for example, is important for proper body pattern formation in organisms as diverse as fruit flies to humans. [24] [25] Another example is the transcription factor encoded by the sex-determining region Y (SRY) gene, which plays a major role in determining sex in humans. [26]
Response to intercellular signals
Cells can communicate with each other by releasing molecules that produce signaling cascades within another receptive cell. If the signal requires upregulation or downregulation of genes in the recipient cell, often transcription factors will be downstream in the signaling cascade. [27] Estrogen signaling is an example of a fairly short signaling cascade that involves the estrogen receptor transcription factor: Estrogen is secreted by tissues such as the ovaries and placenta, crosses the cell membrane of the recipient cell, and is bound by the estrogen receptor in the cell's cytoplasm. The estrogen receptor then goes to the cell's nucleus and binds to its DNA-binding sites, changing the transcriptional regulation of the associated genes. [28]
Response to environment
Not only do transcription factors act downstream of signaling cascades related to biological stimuli but they can also be downstream of signaling cascades involved in environmental stimuli. Examples include heat shock factor (HSF), which upregulates genes necessary for survival at higher temperatures, [29] hypoxia inducible factor (HIF), which upregulates genes necessary for cell survival in low-oxygen environments, [30] and sterol regulatory element binding protein (SREBP), which helps maintain proper lipid levels in the cell. [31]
Cell cycle control
Many transcription factors, especially some that are proto-oncogenes or tumor suppressors, help regulate the cell cycle and as such determine how large a cell will get and when it can divide into two daughter cells. [32] [33] One example is the Myc oncogene, which has important roles in cell growth and apoptosis. [34]
Pathogenesis
Transcription factors can also be used to alter gene expression in a host cell to promote pathogenesis. A well studied example of this are the transcription-activator like effectors ( TAL effectors) secreted by Xanthomonas bacteria. When injected into plants, these proteins can enter the nucleus of the plant cell, bind plant promoter sequences, and activate transcription of plant genes that aid in bacterial infection. [35] TAL effectors contain a central repeat region in which there is a simple relationship between the identity of two critical residues in sequential repeats and sequential DNA bases in the TAL effector's target site. [36] [37] This property likely makes it easier for these proteins to evolve in order to better compete with the defense mechanisms of the host cell. [38]
Regulation
It is common in biology for important processes to have multiple layers of regulation and control. This is also true with transcription factors: Not only do transcription factors control the rates of transcription to regulate the amounts of gene products (RNA and protein) available to the cell but transcription factors themselves are regulated (often by other transcription factors). Below is a brief synopsis of some of the ways that the activity of transcription factors can be regulated:
Synthesis
Transcription factors (like all proteins) are transcribed from a gene on a chromosome into RNA, and then the RNA is translated into protein. Any of these steps can be regulated to affect the production (and thus activity) of a transcription factor. An implication of this is that transcription factors can regulate themselves. For example, in a negative feedback loop, the transcription factor acts as its own repressor: If the transcription factor protein binds the DNA of its own gene, it down-regulates the production of more of itself. This is one mechanism to maintain low levels of a transcription factor in a cell. [39]
Nuclear localization
In eukaryotes, transcription factors (like most proteins) are transcribed in the nucleus but are then translated in the cell's cytoplasm. Many proteins that are active in the nucleus contain nuclear localization signals that direct them to the nucleus. But, for many transcription factors, this is a key point in their regulation. [40] Important classes of transcription factors such as some nuclear receptors must first bind a ligand while in the cytoplasm before they can relocate to the nucleus. [40]
Activation
Transcription factors may be activated (or deactivated) through their signal-sensing domain by a number of mechanisms including:
- ligand binding – Not only is ligand binding able to influence where a transcription factor is located within a cell but ligand binding can also affect whether the transcription factor is in an active state and capable of binding DNA or other cofactors (see, for example, nuclear receptors).
- phosphorylation [41] [42] – Many transcription factors such as STAT proteins must be phosphorylated before they can bind DNA.
- interaction with other transcription factors (e.g., homo- or hetero- dimerization) or coregulatory proteins[ citation needed]
Accessibility of DNA-binding site
In eukaryotes, DNA is organized with the help of histones into compact particles called nucleosomes, where sequences of about 147 DNA base pairs make ~1.65 turns around histone protein octamers. DNA within nucleosomes is inaccessible to many transcription factors. Some transcription factors, so-called pioneer factors are still able to bind their DNA binding sites on the nucleosomal DNA. For most other transcription factors, the nucleosome should be actively unwound by molecular motors such as chromatin remodelers. [43] Alternatively, the nucleosome can be partially unwrapped by thermal fluctuations, allowing temporary access to the transcription factor binding site. In many cases, a transcription factor needs to compete for binding to its DNA binding site with other transcription factors and histones or non-histone chromatin proteins. [44] Pairs of transcription factors and other proteins can play antagonistic roles (activator versus repressor) in the regulation of the same gene.[ citation needed]
Availability of other cofactors/transcription factors
Most transcription factors do not work alone. Many large TF families form complex homotypic or heterotypic interactions through dimerization. [45] For gene transcription to occur, a number of transcription factors must bind to DNA regulatory sequences. This collection of transcription factors, in turn, recruit intermediary proteins such as cofactors that allow efficient recruitment of the preinitiation complex and RNA polymerase. Thus, for a single transcription factor to initiate transcription, all of these other proteins must also be present, and the transcription factor must be in a state where it can bind to them if necessary. Cofactors are proteins that modulate the effects of transcription factors. Cofactors are interchangeable between specific gene promoters; the protein complex that occupies the promoter DNA and the amino acid sequence of the cofactor determine its spatial conformation. For example, certain steroid receptors can exchange cofactors with NF-κB, which is a switch between inflammation and cellular differentiation; thereby steroids can affect the inflammatory response and function of certain tissues. [46]
Interaction with methylated cytosine
Transcription factors and methylated cytosines in DNA both have major roles in regulating gene expression. (Methylation of cytosine in DNA primarily occurs where cytosine is followed by guanine in the 5' to 3' DNA sequence, a CpG site.) Methylation of CpG sites in a promoter region of a gene usually represses gene transcription, [47] while methylation of CpGs in the body of a gene increases expression. [48] TET enzymes play a central role in demethylation of methylated cytosines. Demethylation of CpGs in a gene promoter by TET enzyme activity increases transcription of the gene. [49]
The DNA binding sites of 519 transcription factors were evaluated. [50] Of these, 169 transcription factors (33%) did not have CpG dinucleotides in their binding sites, and 33 transcription factors (6%) could bind to a CpG-containing motif but did not display a preference for a binding site with either a methylated or unmethylated CpG. There were 117 transcription factors (23%) that were inhibited from binding to their binding sequence if it contained a methylated CpG site, 175 transcription factors (34%) that had enhanced binding if their binding sequence had a methylated CpG site, and 25 transcription factors (5%) were either inhibited or had enhanced binding depending on where in the binding sequence the methylated CpG was located.[ citation needed]
TET enzymes do not specifically bind to methylcytosine except when recruited (see DNA demethylation). Multiple transcription factors important in cell differentiation and lineage specification, including NANOG, SALL4A, WT1, EBF1, PU.1, and E2A, have been shown to recruit TET enzymes to specific genomic loci (primarily enhancers) to act on methylcytosine (mC) and convert it to hydroxymethylcytosine hmC (and in most cases marking them for subsequent complete demethylation to cytosine). [51] TET-mediated conversion of mC to hmC appears to disrupt the binding of 5mC-binding proteins including MECP2 and MBD ( Methyl-CpG-binding domain) proteins, facilitating nucleosome remodeling and the binding of transcription factors, thereby activating transcription of those genes. EGR1 is an important transcription factor in memory formation. It has an essential role in brain neuron epigenetic reprogramming. The transcription factor EGR1 recruits the TET1 protein that initiates a pathway of DNA demethylation. [52] EGR1, together with TET1, is employed in programming the distribution of methylation sites on brain DNA during brain development and in learning (see Epigenetics in learning and memory).
Structure


Transcription factors are modular in structure and contain the following domains: [1]
- DNA-binding domain (DBD), which attaches to specific sequences of DNA ( enhancer or promoter. Necessary component for all vectors. Used to drive transcription of the vector's transgene promoter sequences) adjacent to regulated genes. DNA sequences that bind transcription factors are often referred to as response elements.
- Activation domain (AD), which contains binding sites for other proteins such as transcription coregulators. These binding sites are frequently referred to as activation functions (AFs), Transactivation domain (TAD) or Trans-activating domain TAD, not to be confused with topologically associating domain ( TAD). [53]
- An optional signal-sensing domain (SSD) (e.g., a ligand-binding domain), which senses external signals and, in response, transmits these signals to the rest of the transcription complex, resulting in up- or down-regulation of gene expression. Also, the DBD and signal-sensing domains may reside on separate proteins that associate within the transcription complex to regulate gene expression.
DNA-binding domain
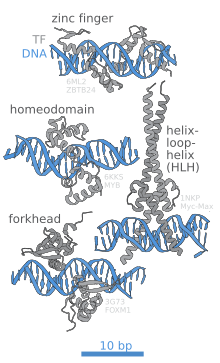
The portion ( domain) of the transcription factor that binds DNA is called its DNA-binding domain. Below is a partial list of some of the major families of DNA-binding domains/transcription factors:
| Family | InterPro | Pfam | SCOP |
|---|---|---|---|
| basic helix-loop-helix [54] | InterPro: IPR001092 | Pfam PF00010 | SCOP 47460 |
| basic-leucine zipper ( bZIP) [55] | InterPro: IPR004827 | Pfam PF00170 | SCOP 57959 |
| C-terminal effector domain of the bipartite response regulators | InterPro: IPR001789 | Pfam PF00072 | SCOP 46894 |
| AP2/ERF/GCC box | InterPro: IPR001471 | Pfam PF00847 | SCOP 54176 |
| helix-turn-helix [56] | |||
| homeodomain proteins, which are encoded by homeobox genes, are transcription factors. Homeodomain proteins play critical roles in the regulation of development. [57] [58] | InterPro: IPR009057 | Pfam PF00046 | SCOP 46689 |
| lambda repressor-like | InterPro: IPR010982 | SCOP 47413 | |
| srf-like ( serum response factor) | InterPro: IPR002100 | Pfam PF00319 | SCOP 55455 |
| paired box [59] | |||
| winged helix | InterPro: IPR013196 | Pfam PF08279 | SCOP 46785 |
| zinc fingers [60] | |||
| * multi-domain Cys2His2 zinc fingers [61] | InterPro: IPR007087 | Pfam PF00096 | SCOP 57667 |
| * Zn2/Cys6 | SCOP 57701 | ||
| * Zn2/Cys8 nuclear receptor zinc finger | InterPro: IPR001628 | Pfam PF00105 | SCOP 57716 |
Response elements
The DNA sequence that a transcription factor binds to is called a transcription factor-binding site or response element. [62]
Transcription factors interact with their binding sites using a combination of electrostatic (of which hydrogen bonds are a special case) and Van der Waals forces. Due to the nature of these chemical interactions, most transcription factors bind DNA in a sequence specific manner. However, not all bases in the transcription factor-binding site may actually interact with the transcription factor. In addition, some of these interactions may be weaker than others. Thus, transcription factors do not bind just one sequence but are capable of binding a subset of closely related sequences, each with a different strength of interaction.[ citation needed]
For example, although the consensus binding site for the TATA-binding protein (TBP) is TATAAAA, the TBP transcription factor can also bind similar sequences such as TATATAT or TATATAA.[ citation needed]
Because transcription factors can bind a set of related sequences and these sequences tend to be short, potential transcription factor binding sites can occur by chance if the DNA sequence is long enough. It is unlikely, however, that a transcription factor will bind all compatible sequences in the genome of the cell. Other constraints, such as DNA accessibility in the cell or availability of cofactors may also help dictate where a transcription factor will actually bind. Thus, given the genome sequence, it is still difficult to predict where a transcription factor will actually bind in a living cell.
Additional recognition specificity, however, may be obtained through the use of more than one DNA-binding domain (for example tandem DBDs in the same transcription factor or through dimerization of two transcription factors) that bind to two or more adjacent sequences of DNA.
Clinical significance
Transcription factors are of clinical significance for at least two reasons: (1) mutations can be associated with specific diseases, and (2) they can be targets of medications.
Disorders
Due to their important roles in development, intercellular signaling, and cell cycle, some human diseases have been associated with mutations in transcription factors. [63]
Many transcription factors are either tumor suppressors or oncogenes, and, thus, mutations or aberrant regulation of them is associated with cancer. Three groups of transcription factors are known to be important in human cancer: (1) the NF-kappaB and AP-1 families, (2) the STAT family and (3) the steroid receptors. [64]
Below are a few of the better-studied examples:
| Condition | Description | Locus |
|---|---|---|
| Rett syndrome | Mutations in the MECP2 transcription factor are associated with Rett syndrome, a neurodevelopmental disorder. [65] [66] | Xq28 |
| Diabetes | A rare form of diabetes called MODY (Maturity onset diabetes of the young) can be caused by mutations in hepatocyte nuclear factors (HNFs) [67] or insulin promoter factor-1 (IPF1/Pdx1). [68] | multiple |
| Developmental verbal dyspraxia | Mutations in the FOXP2 transcription factor are associated with developmental verbal dyspraxia, a disease in which individuals are unable to produce the finely coordinated movements required for speech. [69] | 7q31 |
| Autoimmune diseases | Mutations in the FOXP3 transcription factor cause a rare form of autoimmune disease called IPEX. [70] | Xp11.23-q13.3 |
| Li-Fraumeni syndrome | Caused by mutations in the tumor suppressor p53. [71] | 17p13.1 |
| Breast cancer | The STAT family is relevant to breast cancer. [72] | multiple |
| Multiple cancers | The HOX family are involved in a variety of cancers. [73] | multiple |
| Osteoarthritis | Mutation or reduced activity of SOX9 [74] |
Potential drug targets
Approximately 10% of currently prescribed drugs directly target the nuclear receptor class of transcription factors. [75] Examples include tamoxifen and bicalutamide for the treatment of breast and prostate cancer, respectively, and various types of anti-inflammatory and anabolic steroids. [76] In addition, transcription factors are often indirectly modulated by drugs through signaling cascades. It might be possible to directly target other less-explored transcription factors such as NF-κB with drugs. [77] [78] [79] [80] Transcription factors outside the nuclear receptor family are thought to be more difficult to target with small molecule therapeutics since it is not clear that they are "drugable" but progress has been made on Pax2 [81] [82] and the notch pathway. [83]
Role in evolution
Gene duplications have played a crucial role in the evolution of species. This applies particularly to transcription factors. Once they occur as duplicates, accumulated mutations encoding for one copy can take place without negatively affecting the regulation of downstream targets. However, changes of the DNA binding specificities of the single-copy Leafy transcription factor, which occurs in most land plants, have recently been elucidated. In that respect, a single-copy transcription factor can undergo a change of specificity through a promiscuous intermediate without losing function. Similar mechanisms have been proposed in the context of all alternative phylogenetic hypotheses, and the role of transcription factors in the evolution of all species. [84] [85]
Role in biocontrol activity
The transcription factors have a role in resistance activity which is important for successful biocontrol activity. The resistant to oxidative stress and alkaline pH sensing were contributed from the transcription factor Yap1 and Rim101 of the Papiliotrema terrestris LS28 as molecular tools revealed an understanding of the genetic mechanisms underlying the biocontrol activity which supports disease management programs based on biological and integrated control. [86]
Analysis
There are different technologies available to analyze transcription factors. On the genomic level, DNA- sequencing and database research are commonly used. [87] The protein version of the transcription factor is detectable by using specific antibodies. The sample is detected on a western blot. By using electrophoretic mobility shift assay (EMSA), [88] the activation profile of transcription factors can be detected. A multiplex approach for activation profiling is a TF chip system where several different transcription factors can be detected in parallel.[ citation needed]
The most commonly used method for identifying transcription factor binding sites is chromatin immunoprecipitation (ChIP). [89] This technique relies on chemical fixation of chromatin with formaldehyde, followed by co-precipitation of DNA and the transcription factor of interest using an antibody that specifically targets that protein. The DNA sequences can then be identified by microarray or high-throughput sequencing ( ChIP-seq) to determine transcription factor binding sites. If no antibody is available for the protein of interest, DamID may be a convenient alternative. [90]
Classes
As described in more detail below, transcription factors may be classified by their (1) mechanism of action, (2) regulatory function, or (3) sequence homology (and hence structural similarity) in their DNA-binding domains.
Mechanistic
There are two mechanistic classes of transcription factors:
- General transcription factors are involved in the formation of a preinitiation complex. The most common are abbreviated as TFIIA, TFIIB, TFIID, TFIIE, TFIIF, and TFIIH. They are ubiquitous and interact with the core promoter region surrounding the transcription start site(s) of all class II genes. [91]
- Upstream transcription factors are proteins that bind somewhere upstream of the initiation site to stimulate or repress transcription. These are roughly synonymous with specific transcription factors, because they vary considerably depending on what recognition sequences are present in the proximity of the gene. [92]
| Examples of specific transcription factors [92] | |||
|---|---|---|---|
| Factor | Structural type | Recognition sequence | Binds as |
| SP1 | Zinc finger | 5'-GGGCGG- 3' | Monomer |
| AP-1 | Basic zipper | 5'-TGA(G/C)TCA-3' | Dimer |
| C/EBP | Basic zipper | 5'-ATTGCGCAAT-3' | Dimer |
| Heat shock factor | Basic zipper | 5'-XGAAX-3' | Trimer |
| ATF/CREB | Basic zipper | 5'-TGACGTCA-3' | Dimer |
| c-Myc | Basic helix-loop-helix | 5'-CACGTG-3' | Dimer |
| Oct-1 | Helix-turn-helix | 5'-ATGCAAAT-3' | Monomer |
| NF-1 | Novel | 5'-TTGGCXXXXXGCCAA-3' | Dimer |
| (G/C) = G or C X = A, T, G or C | |||
Functional
Transcription factors have been classified according to their regulatory function: [13]
- I. constitutively active – present in all cells at all times – general transcription factors, Sp1, NF1, CCAAT
- II. conditionally active – requires activation
- II.A developmental (cell specific) – expression is tightly controlled, but, once expressed, require no additional activation – GATA, HNF, PIT-1, MyoD, Myf5, Hox, Winged Helix
- II.B signal-dependent – requires external signal for activation
- II.B.1 extracellular ligand ( endocrine or paracrine)-dependent – nuclear receptors
- II.B.2 intracellular ligand ( autocrine)-dependent – activated by small intracellular molecules – SREBP, p53, orphan nuclear receptors
- II.B.3 cell membrane receptor-dependent – second messenger signaling cascades resulting in the phosphorylation of the transcription factor
Structural
Transcription factors are often classified based on the sequence similarity and hence the tertiary structure of their DNA-binding domains: [93] [12] [94] [11]
- 1 Superclass: Basic Domains
- 1.1 Class: Leucine zipper factors ( bZIP)
- 1.2 Class: Helix-loop-helix factors (
bHLH)
- 1.2.1 Family: Ubiquitous (class A) factors
- 1.2.2 Family: Myogenic transcription factors ( MyoD)
- 1.2.3 Family: Achaete-Scute
- 1.2.4 Family: Tal/Twist/Atonal/Hen
- 1.3 Class: Helix-loop-helix / leucine zipper factors ( bHLH-ZIP)
- 1.4 Class: NF-1
- 1.5 Class: RF-X
- 1.6 Class: bHSH
- 2 Superclass: Zinc-coordinating DNA-binding domains
- 2.1 Class: Cys4
zinc finger of
nuclear receptor type
- 2.1.1 Family: Steroid hormone receptors
- 2.1.2 Family: Thyroid hormone receptor-like factors
- 2.2 Class: diverse Cys4 zinc fingers
- 2.2.1 Family: GATA-Factors
- 2.3 Class: Cys2His2 zinc finger domain
- 2.4 Class: Cys6 cysteine-zinc cluster
- 2.5 Class: Zinc fingers of alternating composition
- 2.1 Class: Cys4
zinc finger of
nuclear receptor type
- 3 Superclass:
Helix-turn-helix
- 3.1 Class:
Homeo domain
- 3.1.1 Family: Homeo domain only; includes Ubx
- 3.1.2 Family: POU domain factors; includes Oct
- 3.1.3 Family: Homeo domain with LIM region
- 3.1.4 Family: homeo domain plus zinc finger motifs
- 3.2 Class: Paired box
- 3.2.1 Family: Paired plus homeo domain
- 3.2.2 Family: Paired domain only
- 3.3 Class:
Fork head /
winged helix
- 3.3.1 Family: Developmental regulators; includes forkhead
- 3.3.2 Family: Tissue-specific regulators
- 3.3.3 Family: Cell-cycle controlling factors
- 3.3.0 Family: Other regulators
- 3.4 Class:
Heat Shock Factors
- 3.4.1 Family: HSF
- 3.5 Class: Tryptophan clusters
- 3.5.1 Family: Myb
- 3.5.2 Family: Ets-type
- 3.5.3 Family: Interferon regulatory factors
- 3.6 Class: TEA ( transcriptional enhancer factor) domain
- 3.1 Class:
Homeo domain
- 4 Superclass: beta-Scaffold Factors with Minor Groove Contacts
- 4.1 Class: RHR ( Rel homology region)
- 4.2 Class: STAT
- 4.2.1 Family: STAT
- 4.3 Class: p53
- 4.3.1 Family: p53
- 4.4 Class:
MADS box
- 4.4.1 Family: Regulators of differentiation; includes ( Mef2)
- 4.4.2 Family: Responders to external signals, SRF ( serum response factor) ( SRF)
- 4.4.3 Family: Metabolic regulators (ARG80)
- 4.5 Class: beta-Barrel alpha-helix transcription factors
- 4.6 Class:
TATA binding proteins
- 4.6.1 Family: TBP
- 4.7 Class: HMG-box
- 4.8 Class: Heteromeric CCAAT factors
- 4.8.1 Family: Heteromeric CCAAT factors
- 4.9 Class: Grainyhead
- 4.9.1 Family: Grainyhead
- 4.10 Class:
Cold-shock domain factors
- 4.10.1 Family: csd
- 4.11 Class: Runt
- 4.11.1 Family: Runt
- 0 Superclass: Other Transcription Factors
Transcription factor databases
There are numerous databases cataloging information about transcription factors, but their scope and utility vary dramatically. Some may contain only information about the actual proteins, some about their binding sites, or about their target genes. Examples include the following:
- footprintDB-- a metadatabase of multiple databases, including JASPAR and others
- JASPAR: database of transcription factor binding sites for eukaryotes
- PlantTFD: Plant transcription factor database [95]
- TcoF-DB: Database of transcription co-factors and transcription factor interactions [96]
- TFcheckpoint: database of human, mouse and rat TF candidates
- transcriptionfactor.org (now commercial, selling reagents)
- MethMotif.org: An integrative cell-specific database of transcription factor binding motifs coupled with DNA methylation profiles. [97]
See also
- Cdx protein family
- DNA-binding protein
- Inhibitor of DNA-binding protein
- Mapper(2)
- Nuclear receptor, a class of ligand activated transcription factors
- Open Regulatory Annotation Database
- Phylogenetic footprinting
- TRANSFAC database
- YeTFaSCo
References
- ^ a b Latchman DS (December 1997). "Transcription factors: an overview". The International Journal of Biochemistry & Cell Biology. 29 (12): 1305–12. doi: 10.1016/S1357-2725(97)00085-X. PMC 2002184. PMID 9570129.
- ^ Karin M (February 1990). "Too many transcription factors: positive and negative interactions". The New Biologist. 2 (2): 126–31. PMID 2128034.
- ^ a b Babu MM, Luscombe NM, Aravind L, Gerstein M, Teichmann SA (June 2004). "Structure and evolution of transcriptional regulatory networks" (PDF). Current Opinion in Structural Biology. 14 (3): 283–91. doi: 10.1016/j.sbi.2004.05.004. PMID 15193307.
- ^ How Genes are Regulated: Transcription Factors on YouTube
-
^ Lambert S, Jolma A, Campitelli L, Pratyush Z, Das K, Yin Y, et al. (2018).
"The Human Transcription Factors". Cell. 172 (4): 650–665.
doi:
10.1016/j.cell.2018.01.029.
PMID
29425488.
The final tally encompasses 1,639 known or likely human TFs.
- ^ Roeder RG (September 1996). "The role of general initiation factors in transcription by RNA polymerase II". Trends in Biochemical Sciences. 21 (9): 327–35. doi: 10.1016/S0968-0004(96)10050-5. PMID 8870495.
- ^ Nikolov DB, Burley SK (January 1997). "RNA polymerase II transcription initiation: a structural view". Proceedings of the National Academy of Sciences of the United States of America. 94 (1): 15–22. Bibcode: 1997PNAS...94...15N. doi: 10.1073/pnas.94.1.15. PMC 33652. PMID 8990153.
- ^ Lee TI, Young RA (2000). "Transcription of eukaryotic protein-coding genes". Annual Review of Genetics. 34: 77–137. doi: 10.1146/annurev.genet.34.1.77. PMID 11092823.
- ^ Mitchell PJ, Tjian R (July 1989). "Transcriptional regulation in mammalian cells by sequence-specific DNA binding proteins". Science. 245 (4916): 371–8. Bibcode: 1989Sci...245..371M. doi: 10.1126/science.2667136. PMID 2667136.
- ^ Ptashne M, Gann A (April 1997). "Transcriptional activation by recruitment". Nature. 386 (6625): 569–77. Bibcode: 1997Natur.386..569P. doi: 10.1038/386569a0. PMID 9121580. S2CID 6203915.
- ^ a b Jin J, Zhang H, Kong L, Gao G, Luo J (January 2014). "PlantTFDB 3.0: a portal for the functional and evolutionary study of plant transcription factors". Nucleic Acids Research. 42 (Database issue): D1182-7. doi: 10.1093/nar/gkt1016. PMC 3965000. PMID 24174544.
- ^ a b Matys V, Kel-Margoulis OV, Fricke E, Liebich I, Land S, Barre-Dirrie A, et al. (January 2006). "TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes". Nucleic Acids Research. 34 (Database issue): D108-10. doi: 10.1093/nar/gkj143. PMC 1347505. PMID 16381825.
- ^ a b c Brivanlou AH, Darnell JE (February 2002). "Signal transduction and the control of gene expression". Science. 295 (5556): 813–8. Bibcode: 2002Sci...295..813B. doi: 10.1126/science.1066355. PMID 11823631. S2CID 14954195.
- ^ van Nimwegen E (September 2003). "Scaling laws in the functional content of genomes". Trends in Genetics. 19 (9): 479–84. arXiv: physics/0307001. doi: 10.1016/S0168-9525(03)00203-8. PMID 12957540. S2CID 15887416.
- ^ "List Of All Transcription Factors In Human". biostars.org.
- ^ Gill G (2001). "Regulation of the initiation of eukaryotic transcription". Essays in Biochemistry. 37: 33–43. doi: 10.1042/bse0370033. PMID 11758455.
- ^ Narlikar GJ, Fan HY, Kingston RE (February 2002). "Cooperation between complexes that regulate chromatin structure and transcription". Cell. 108 (4): 475–87. doi: 10.1016/S0092-8674(02)00654-2. PMID 11909519. S2CID 14586791.
- ^ Xu L, Glass CK, Rosenfeld MG (April 1999). "Coactivator and corepressor complexes in nuclear receptor function". Current Opinion in Genetics & Development. 9 (2): 140–7. doi: 10.1016/S0959-437X(99)80021-5. PMID 10322133.
- ^ Robert O. J. Weinzierl (1999). Mechanisms of Gene Expression: Structure, Function and Evolution of the Basal Transcriptional Machinery. World Scientific Publishing Company. ISBN 1-86094-126-5.
- ^ Reese JC (April 2003). "Basal transcription factors". Current Opinion in Genetics & Development. 13 (2): 114–8. doi: 10.1016/S0959-437X(03)00013-3. PMID 12672487.
- ^ Shilatifard A, Conaway RC, Conaway JW (2003). "The RNA polymerase II elongation complex". Annual Review of Biochemistry. 72: 693–715. doi: 10.1146/annurev.biochem.72.121801.161551. PMID 12676794.
- ^ Thomas MC, Chiang CM (2006). "The general transcription machinery and general cofactors". Critical Reviews in Biochemistry and Molecular Biology. 41 (3): 105–78. doi: 10.1080/10409230600648736. PMID 16858867. S2CID 13073440.
- ^ Lobe CG (1992). Transcription factors and mammalian development. Current Topics in Developmental Biology. Vol. 27. pp. 351–83. doi: 10.1016/S0070-2153(08)60539-6. ISBN 978-0-12-153127-0. PMID 1424766.
- ^ Lemons D, McGinnis W (September 2006). "Genomic evolution of Hox gene clusters". Science. 313 (5795): 1918–22. Bibcode: 2006Sci...313.1918L. doi: 10.1126/science.1132040. PMID 17008523. S2CID 35650754.
- ^ Moens CB, Selleri L (March 2006). "Hox cofactors in vertebrate development". Developmental Biology. 291 (2): 193–206. doi: 10.1016/j.ydbio.2005.10.032. PMID 16515781.
- ^ Ottolenghi C, Uda M, Crisponi L, Omari S, Cao A, Forabosco A, et al. (January 2007). "Determination and stability of sex". BioEssays. 29 (1): 15–25. doi: 10.1002/bies.20515. PMID 17187356. S2CID 23824870.
- ^ Pawson T (1993). "Signal transduction--a conserved pathway from the membrane to the nucleus". Developmental Genetics. 14 (5): 333–8. doi: 10.1002/dvg.1020140502. PMID 8293575.
- ^ Osborne CK, Schiff R, Fuqua SA, Shou J (December 2001). "Estrogen receptor: current understanding of its activation and modulation". Clinical Cancer Research. 7 (12 Suppl): 4338s–4342s, discussion 4411s–4412s. PMID 11916222.
- ^ Shamovsky I, Nudler E (March 2008). "New insights into the mechanism of heat shock response activation". Cellular and Molecular Life Sciences. 65 (6): 855–61. doi: 10.1007/s00018-008-7458-y. PMID 18239856. S2CID 9912334.
- ^ Benizri E, Ginouvès A, Berra E (April 2008). "The magic of the hypoxia-signaling cascade". Cellular and Molecular Life Sciences. 65 (7–8): 1133–49. doi: 10.1007/s00018-008-7472-0. PMID 18202826. S2CID 44049779.
- ^ Weber LW, Boll M, Stampfl A (November 2004). "Maintaining cholesterol homeostasis: sterol regulatory element-binding proteins". World Journal of Gastroenterology. 10 (21): 3081–7. doi: 10.3748/wjg.v10.i21.3081. PMC 4611246. PMID 15457548.
- ^ Wheaton K, Atadja P, Riabowol K (1996). "Regulation of transcription factor activity during cellular aging". Biochemistry and Cell Biology. 74 (4): 523–34. doi: 10.1139/o96-056. PMID 8960358.
- ^ Meyyappan M, Atadja PW, Riabowol KT (1996). "Regulation of gene expression and transcription factor binding activity during cellular aging". Biological Signals. 5 (3): 130–8. doi: 10.1159/000109183. PMID 8864058.
- ^ Evan G, Harrington E, Fanidi A, Land H, Amati B, Bennett M (August 1994). "Integrated control of cell proliferation and cell death by the c-myc oncogene". Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 345 (1313): 269–75. Bibcode: 1994RSPTB.345..269E. doi: 10.1098/rstb.1994.0105. PMID 7846125.
- ^ Boch J, Bonas U (2010). "Xanthomonas AvrBs3 family-type III effectors: discovery and function". Annual Review of Phytopathology. 48: 419–36. doi: 10.1146/annurev-phyto-080508-081936. PMID 19400638.
- ^ Moscou MJ, Bogdanove AJ (December 2009). "A simple cipher governs DNA recognition by TAL effectors". Science. 326 (5959): 1501. Bibcode: 2009Sci...326.1501M. doi: 10.1126/science.1178817. PMID 19933106. S2CID 6648530.
- ^ Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, et al. (December 2009). "Breaking the code of DNA binding specificity of TAL-type III effectors". Science. 326 (5959): 1509–12. Bibcode: 2009Sci...326.1509B. doi: 10.1126/science.1178811. PMID 19933107. S2CID 206522347.
- ^ Voytas DF, Joung JK (December 2009). "Plant science. DNA binding made easy". Science. 326 (5959): 1491–2. Bibcode: 2009Sci...326.1491V. doi: 10.1126/science.1183604. PMC 7814878. PMID 20007890. S2CID 33257689.
- ^ Pan G, Li J, Zhou Y, Zheng H, Pei D (August 2006). "A negative feedback loop of transcription factors that controls stem cell pluripotency and self-renewal". FASEB Journal. 20 (10): 1730–2. doi: 10.1096/fj.05-5543fje. PMID 16790525. S2CID 44783683.
- ^ a b Whiteside ST, Goodbourn S (April 1993). "Signal transduction and nuclear targeting: regulation of transcription factor activity by subcellular localisation". Journal of Cell Science. 104 (4): 949–55. doi: 10.1242/jcs.104.4.949. PMID 8314906.
- ^ Bohmann D (November 1990). "Transcription factor phosphorylation: a link between signal transduction and the regulation of gene expression". Cancer Cells. 2 (11): 337–44. PMID 2149275.
- ^ Weigel NL, Moore NL (October 2007). "Steroid receptor phosphorylation: a key modulator of multiple receptor functions". Molecular Endocrinology. 21 (10): 2311–9. doi: 10.1210/me.2007-0101. PMID 17536004.
- ^ Teif VB, Rippe K (September 2009). "Predicting nucleosome positions on the DNA: combining intrinsic sequence preferences and remodeler activities". Nucleic Acids Research. 37 (17): 5641–55. doi: 10.1093/nar/gkp610. PMC 2761276. PMID 19625488.
- ^ Teif VB, Rippe K (October 2010). "Statistical-mechanical lattice models for protein-DNA binding in chromatin". Journal of Physics: Condensed Matter. 22 (41): 414105. arXiv: 1004.5514. Bibcode: 2010JPCM...22O4105T. doi: 10.1088/0953-8984/22/41/414105. PMID 21386588. S2CID 103345.
- ^ Amoutzias GD, Robertson DL, Van de Peer Y, Oliver SG (May 2008). "Choose your partners: dimerization in eukaryotic transcription factors". Trends in Biochemical Sciences. 33 (5): 220–9. doi: 10.1016/j.tibs.2008.02.002. PMID 18406148.
- ^ Copland JA, Sheffield-Moore M, Koldzic-Zivanovic N, Gentry S, Lamprou G, Tzortzatou-Stathopoulou F, et al. (June 2009). "Sex steroid receptors in skeletal differentiation and epithelial neoplasia: is tissue-specific intervention possible?". BioEssays. 31 (6): 629–41. doi: 10.1002/bies.200800138. PMID 19382224. S2CID 205469320.
- ^ Weber M, Hellmann I, Stadler MB, Ramos L, Pääbo S, Rebhan M, et al. (April 2007). "Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome". Nat. Genet. 39 (4): 457–66. doi: 10.1038/ng1990. PMID 17334365. S2CID 22446734.
- ^ Yang X, Han H, De Carvalho DD, Lay FD, Jones PA, Liang G (October 2014). "Gene body methylation can alter gene expression and is a therapeutic target in cancer". Cancer Cell. 26 (4): 577–90. doi: 10.1016/j.ccr.2014.07.028. PMC 4224113. PMID 25263941.
- ^ Maeder ML, Angstman JF, Richardson ME, Linder SJ, Cascio VM, Tsai SQ, et al. (December 2013). "Targeted DNA demethylation and activation of endogenous genes using programmable TALE-TET1 fusion proteins". Nat. Biotechnol. 31 (12): 1137–42. doi: 10.1038/nbt.2726. PMC 3858462. PMID 24108092.
- ^ Yin Y, Morgunova E, Jolma A, Kaasinen E, Sahu B, Khund-Sayeed S, et al. (May 2017). "Impact of cytosine methylation on DNA binding specificities of human transcription factors". Science. 356 (6337): eaaj2239. doi: 10.1126/science.aaj2239. PMC 8009048. PMID 28473536. S2CID 206653898.
- ^ Lio CJ, Rao A (2019). "TET Enzymes and 5hmC in Adaptive and Innate Immune Systems". Front Immunol. 10: 210. doi: 10.3389/fimmu.2019.00210. PMC 6379312. PMID 30809228.
- ^ Sun Z, Xu X, He J, Murray A, Sun MA, Wei X, Wang X, McCoig E, Xie E, Jiang X, Li L, Zhu J, Chen J, Morozov A, Pickrell AM, Theus MH, Xie H. EGR1 recruits TET1 to shape the brain methylome during development and upon neuronal activity. Nat Commun. 2019 Aug 29;10(1):3892. doi: 10.1038/s41467-019-11905-3. PMID 31467272
- ^ Wärnmark A, Treuter E, Wright AP, Gustafsson JA (October 2003). "Activation functions 1 and 2 of nuclear receptors: molecular strategies for transcriptional activation". Molecular Endocrinology. 17 (10): 1901–9. doi: 10.1210/me.2002-0384. PMID 12893880. S2CID 31314461.
- ^ Littlewood TD, Evan GI (1995). "Transcription factors 2: helix-loop-helix". Protein Profile. 2 (6): 621–702. PMID 7553065.
- ^ Vinson C, Myakishev M, Acharya A, Mir AA, Moll JR, Bonovich M (September 2002). "Classification of human B-ZIP proteins based on dimerization properties". Molecular and Cellular Biology. 22 (18): 6321–35. doi: 10.1128/MCB.22.18.6321-6335.2002. PMC 135624. PMID 12192032.
- ^ Wintjens R, Rooman M (September 1996). "Structural classification of HTH DNA-binding domains and protein-DNA interaction modes". Journal of Molecular Biology. 262 (2): 294–313. doi: 10.1006/jmbi.1996.0514. PMID 8831795.
- ^ Gehring WJ, Affolter M, Bürglin T (1994). "Homeodomain proteins". Annual Review of Biochemistry. 63: 487–526. doi: 10.1146/annurev.bi.63.070194.002415. PMID 7979246.
- ^ Bürglin TR, Affolter M (June 2016). "Homeodomain proteins: an update". Chromosoma. 125 (3): 497–521. doi: 10.1007/s00412-015-0543-8. PMC 4901127. PMID 26464018.
- ^ Dahl E, Koseki H, Balling R (September 1997). "Pax genes and organogenesis". BioEssays. 19 (9): 755–65. doi: 10.1002/bies.950190905. PMID 9297966. S2CID 23755557.
- ^ Laity JH, Lee BM, Wright PE (February 2001). "Zinc finger proteins: new insights into structural and functional diversity". Current Opinion in Structural Biology. 11 (1): 39–46. doi: 10.1016/S0959-440X(00)00167-6. PMID 11179890.
- ^ Wolfe SA, Nekludova L, Pabo CO (2000). "DNA recognition by Cys2His2 zinc finger proteins". Annual Review of Biophysics and Biomolecular Structure. 29: 183–212. doi: 10.1146/annurev.biophys.29.1.183. PMID 10940247.
- ^ Wang JC (March 2005). "Finding primary targets of transcriptional regulators". Cell Cycle. 4 (3): 356–8. doi: 10.4161/cc.4.3.1521. PMID 15711128.
- ^ Semenza, Gregg L. (1999). Transcription factors and human disease. Oxford [Oxfordshire]: Oxford University Press. ISBN 978-0-19-511239-9.
- ^ Libermann TA, Zerbini LF (February 2006). "Targeting transcription factors for cancer gene therapy". Current Gene Therapy. 6 (1): 17–33. doi: 10.2174/156652306775515501. PMID 16475943.
- ^ Moretti P, Zoghbi HY (June 2006). "MeCP2 dysfunction in Rett syndrome and related disorders". Current Opinion in Genetics & Development. 16 (3): 276–81. doi: 10.1016/j.gde.2006.04.009. PMID 16647848.
- ^ Chadwick LH, Wade PA (April 2007). "MeCP2 in Rett syndrome: transcriptional repressor or chromatin architectural protein?". Current Opinion in Genetics & Development. 17 (2): 121–5. doi: 10.1016/j.gde.2007.02.003. PMID 17317146. Archived from the original on 2 October 2023 – via Zenodo.
- ^ Maestro MA, Cardalda C, Boj SF, Luco RF, Servitja JM, Ferrer J (2007). "Distinct Roles of HNF1 Β , HNF1 α , and HNF4 α in Regulating Pancreas Development, Β -Cell Function and Growth". Distinct roles of HNF1beta, HNF1alpha, and HNF4alpha in regulating pancreas development, beta-cell function and growth. Endocrine Development. Vol. 12. pp. 33–45. doi: 10.1159/000109603. ISBN 978-3-8055-8385-5. PMID 17923767.
- ^ Al-Quobaili F, Montenarh M (April 2008). "Pancreatic duodenal homeobox factor-1 and diabetes mellitus type 2 (review)". International Journal of Molecular Medicine. 21 (4): 399–404. doi: 10.3892/ijmm.21.4.399. PMID 18360684. Archived from the original on 2 October 2023.
- ^ Lennon PA, Cooper ML, Peiffer DA, Gunderson KL, Patel A, Peters S, et al. (April 2007). "Deletion of 7q31.1 supports involvement of FOXP2 in language impairment: clinical report and review". American Journal of Medical Genetics. Part A. 143A (8): 791–8. doi: 10.1002/ajmg.a.31632. PMID 17330859. S2CID 22021740.
- ^ van der Vliet HJ, Nieuwenhuis EE (2007). "IPEX as a result of mutations in FOXP3". Clinical & Developmental Immunology. 2007: 1–5. doi: 10.1155/2007/89017. PMC 2248278. PMID 18317533.
- ^ Iwakuma T, Lozano G, Flores ER (July 2005). "Li-Fraumeni syndrome: a p53 family affair". Cell Cycle. 4 (7): 865–7. doi: 10.4161/cc.4.7.1800. PMID 15917654.
- ^ Clevenger CV (November 2004). "Roles and Regulation of Stat Family Transcription Factors in Human Breast Cancer". American Journal of Pathology (Review). 165 (5): 1449–1460. doi: 10.1016/S0002-9440(10)63403-7. PMC 1618660. PMID 15509516.
- ^ ""Transcription factors as targets and markers in cancer" Workshop 2007". Archived from the original on 25 May 2012. Retrieved 14 December 2009.
- ^ Govindaraj K, Hendriks J, Lidke DS, Karperien M, Post JN (January 2019). "Changes in Fluorescence Recovery After Photobleaching (FRAP) as an indicator of SOX9 transcription factor activity". Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms. 1862 (1): 107–117. doi: 10.1016/j.bbagrm.2018.11.001. PMID 30465885.
- ^ Overington JP, Al-Lazikani B, Hopkins AL (December 2006). "How many drug targets are there?". Nature Reviews. Drug Discovery. 5 (12): 993–6. doi: 10.1038/nrd2199. PMID 17139284. S2CID 11979420.
- ^ Gronemeyer H, Gustafsson JA, Laudet V (November 2004). "Principles for modulation of the nuclear receptor superfamily". Nature Reviews. Drug Discovery. 3 (11): 950–64. doi: 10.1038/nrd1551. PMID 15520817. S2CID 205475111.
- ^ Bustin SA, McKay IA (June 1994). "Transcription factors: targets for new designer drugs". British Journal of Biomedical Science. 51 (2): 147–57. PMID 8049612.
- ^ Butt TR, Karathanasis SK (1995). "Transcription factors as drug targets: opportunities for therapeutic selectivity". Gene Expression. 4 (6): 319–36. PMC 6134363. PMID 7549464.
- ^ Papavassiliou AG (August 1998). "Transcription-factor-modulating agents: precision and selectivity in drug design". Molecular Medicine Today. 4 (8): 358–66. doi: 10.1016/S1357-4310(98)01303-3. PMID 9755455.
- ^ Ghosh D, Papavassiliou AG (2005). "Transcription factor therapeutics: long-shot or lodestone". Current Medicinal Chemistry. 12 (6): 691–701. doi: 10.2174/0929867053202197. PMID 15790306.
- ^ Grimley E, Liao C, Ranghini E, Nikolovska-Coleska Z, Dressler G (2017). "Inhibition of Pax2 Transcription Activation with a Small Molecule that Targets the DNA Binding Domain". ACS Chemical Biology. 12 (3): 724–734. doi: 10.1021/acschembio.6b00782. PMC 5761330. PMID 28094913.
- ^ Grimley E, Dressler GR (2018). "Are Pax proteins potential therapeutic targets in kidney disease and cancer?". Kidney International. 94 (2): 259–267. doi: 10.1016/j.kint.2018.01.025. PMC 6054895. PMID 29685496.
-
^ Moellering RE, Cornejo M, Davis TN, Del Bianco C, Aster JC, Blacklow SC, et al. (November 2009).
"Direct inhibition of the NOTCH transcription factor complex". Nature. 462 (7270): 182–8.
Bibcode:
2009Natur.462..182M.
doi:
10.1038/nature08543.
PMC
2951323.
PMID
19907488.
- Lay summary in: Katherine Bagley (11 November 2009). "New drug target for cancer". The Scientist. Archived from the original on 16 November 2009.
- ^ Sayou C, Monniaux M, Nanao MH, Moyroud E, Brockington SF, Thévenon E, et al. (February 2014). "A promiscuous intermediate underlies the evolution of LEAFY DNA binding specificity". Science. 343 (6171): 645–8. Bibcode: 2014Sci...343..645S. doi: 10.1126/science.1248229. hdl: 1885/64773. PMID 24436181. S2CID 207778924.
- ^ Jin J, He K, Tang X, Li Z, Lv L, Zhao Y, et al. (July 2015). "An Arabidopsis Transcriptional Regulatory Map Reveals Distinct Functional and Evolutionary Features of Novel Transcription Factors". Molecular Biology and Evolution. 32 (7): 1767–73. doi: 10.1093/molbev/msv058. PMC 4476157. PMID 25750178.
- ^ Castoria R, Miccoli C, Barone G, Palmieri D, De Curtis F, Lima G, et al. (March 2021). Cann I (ed.). "Molecular Tools for the Yeast Papiliotrema terrestris LS28 and Identification of Yap1 as a Transcription Factor Involved in Biocontrol Activity". Applied and Environmental Microbiology. 87 (7). Bibcode: 2021ApEnM..87E2910C. doi: 10.1128/AEM.02910-20. PMC 8091616. PMID 33452020.
- ^ Grau J, Ben-Gal I, Posch S, Grosse I (July 2006). "VOMBAT: prediction of transcription factor binding sites using variable order Bayesian trees" (PDF). Nucleic Acids Research. 34 (Web Server issue): W529-33. doi: 10.1093/nar/gkl212. PMC 1538886. PMID 16845064.
- ^ Wenta N, Strauss H, Meyer S, Vinkemeier U (July 2008). "Tyrosine phosphorylation regulates the partitioning of STAT1 between different dimer conformations". Proceedings of the National Academy of Sciences of the United States of America. 105 (27): 9238–43. Bibcode: 2008PNAS..105.9238W. doi: 10.1073/pnas.0802130105. PMC 2453697. PMID 18591661.
- ^ Furey TS (December 2012). "ChIP-seq and beyond: new and improved methodologies to detect and characterize protein-DNA interactions". Nature Reviews. Genetics. 13 (12): 840–52. doi: 10.1038/nrg3306. PMC 3591838. PMID 23090257.
- ^ Aughey GN, Southall TD (January 2016). "Dam it's good! DamID profiling of protein-DNA interactions". Wiley Interdisciplinary Reviews: Developmental Biology. 5 (1): 25–37. doi: 10.1002/wdev.205. PMC 4737221. PMID 26383089.
- ^ Orphanides G, Lagrange T, Reinberg D (November 1996). "The general transcription factors of RNA polymerase II". Genes & Development. 10 (21): 2657–83. doi: 10.1101/gad.10.21.2657. PMID 8946909.
- ^ a b Boron WF (2003). Medical Physiology: A Cellular And Molecular Approaoch. Elsevier/Saunders. pp. 125–126. ISBN 1-4160-2328-3.
- ^ Stegmaier P, Kel AE, Wingender E (2004). "Systematic DNA-binding domain classification of transcription factors". Genome Informatics. International Conference on Genome Informatics. 15 (2): 276–86. PMID 15706513. Archived from the original on 19 June 2013.
- ^ "TRANSFAC database". Retrieved 5 August 2007.
- ^ Jin J, Tian F, Yang DC, Meng YQ, Kong L, Luo J, et al. (January 2017). "PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants". Nucleic Acids Research. 45 (D1): D1040–D1045. doi: 10.1093/nar/gkw982. PMC 5210657. PMID 27924042.
- ^ Schmeier S, Alam T, Essack M, Bajic VB (January 2017). "TcoF-DB v2: update of the database of human and mouse transcription co-factors and transcription factor interactions". Nucleic Acids Research. 45 (D1): D145–D150. doi: 10.1093/nar/gkw1007. PMC 5210517. PMID 27789689.
- ^ Xuan Lin QX, Sian S, An O, Thieffry D, Jha S, Benoukraf T (January 2019). "MethMotif: an integrative cell specific database of transcription factor binding motifs coupled with DNA methylation profiles". Nucleic Acids Research. 47 (D1): D145–D154. doi: 10.1093/nar/gky1005. PMC 6323897. PMID 30380113.
Further reading
- Carretero-Paulet, Lorenzo; Galstyan, Anahit; Roig-Villanova, Irma; Martínez-García, Jaime F.; Bilbao-Castro, Jose R. «Genome-Wide Classification and Evolutionary Analysis of the bHLH Family of Transcription Factors in Arabidopsis, Poplar, Rice, Moss, and Algae». Plant Physiology, 153, 3, 2010-07, pàg. 1398–1412. doi:10.1104/pp.110.153593. ISSN 0032-0889
- Jin J, He K, Tang X, Li Z, Lv L, Zhao Y, et al. (2015). "An Arabidopsis Transcriptional Regulatory Map Reveals Distinct Functional and Evolutionary Features of Novel Transcription Factors". Molecular Biology and Evolution. 32 (7): 1767–73. doi: 10.1093/molbev/msv058. PMC 4476157. PMID 25750178.
External links
- Transcription+Factors at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Transcription factor database Archived 4 December 2008 at the Wayback Machine