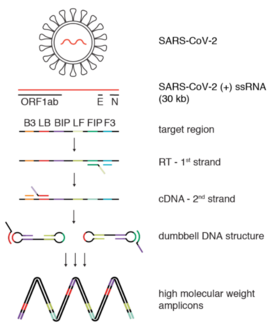
Reverse transcription loop-mediated isothermal amplification (RT-LAMP) is a one step nucleic acid amplification method to multiply specific sequences of RNA. It is used to diagnose infectious disease caused by RNA viruses. [1]
It combines LAMP [2] DNA-detection with reverse transcription, making cDNA from RNA before running the reaction. [3] RT-LAMP does not require thermal cycles (unlike PCR) and is performed at a constant temperature between 60 and 65 °C.
RT-LAMP is used in the detection of RNA viruses (groups II, IV, and V on the Baltimore Virus Classification system), such as the SARS-CoV-2 virus [4] and the Ebola virus. [5]
Applications
RT-LAMP is used to test for the presence of specific RNA-samples of viruses for the specific sequence of the virus, made possible by comparing the sequences against a large external database of references.
Detection of the SARS-CoV2-Virus
The RT-LAMP technique is being supported as a cheaper and easier alternative to RT-PCR for the early diagnostics of people that are infectious for COVID-19. [6] There are open access test designs (including the recombinant proteins) which makes it legally possible for anyone to produce a test. In contrast to classic rapid tests by lateral flow, RT-LAMP allows the early diagnosis of the disease by testing the viral RNA. [7]
The tests can be done without previous RNA-isolation, detecting the viruses directly from swabs [8] or from saliva. [9]
Detection of non-human viruses
One example of use case of RT-LAMP was as an experiment to detect a new duck Tembusu-like, BYD virus, named after the region, Baiyangdian, where it was first isolated [10] [11] [1] Another recent application of this method, was in a 2013 experiment to detect an Akabane virus using RT-LAMP. The experiment, done in China, isolated the virus from aborted calf fetuses. [12]
Detection of body fluids
RT-LAMP is also being used in Forensic Serology to identify body fluids. Researchers have done experiments to show that this method can effectively identify certain body fluids. Knowing there would be limitations, Su et al, come to the conclusion that RT-LAMP was only able to identify blood. [13] [14]
Methodology
Reverse transcription
A specific sequence of the cDNA is detected by 4 LAMP primers. Two of them are inner primers (FIP and BIP), which serve as base for the Bst enzyme copy the template into a new DNA. The outer primers(F3 and B3) anneal to the template strand and help the reaction to proceed.
As in the case of RT-PCR, the RT-LAMP procedure starts by making DNA from the sample RNA. This conversion is made by a reverse transcriptase, an enzyme derived from retroviruses capable of making such a conversion. [15] This DNA derived from RNA is called cDNA, or complementary DNA. The FIP primer is used by the reverse transcriptase to build a single-strand of copy DNA. The F3 primer binds to this side of the template strand as well, and displaces the previously made copy.
Amplification
This displaced, single-stranded copy is a mixture of target RNA and primers. The primers are designed to have a sequence that binds to the sequence itself, forming a loop.
The BIP primer binds to the other end of this single strand and is used by the Bst DNA polymerase to build a complementary strand, making double-strand DNA. The F3 primer binds to this end and displaces, once again, this newly generated single-stranded DNA molecule.
This new single strand that has been released will act as the starting point for the LAMP cycling amplification. This single-stranded DNA has a dumbbell-like structure as the ends fold and self-bind, forming two loops.
The DNA polymerase and the FIP or BIP primers keep amplifying this strand and the LAMP-reaction product is extended. This cycle can be started from either the forward or backward side of the strand using the appropriate primer. Once this cycle has begun, the strand undergoes self-primed DNA synthesis during the elongation stage of the amplification process. This amplification takes place in less an hour, under isothermal conditions between 60 and 65 °C.
Read out
The read out of RT-LAMP tests is frequently colorimetric. Two of the common ways are based on measuring either pH or magnesium ions. The amplification reaction causes pH to lower and Mg2+ levels to drop. This can be perceived by indicators, such as Phenol red, for pH, and hydroxynaphthol blue (HNB), for magnesium. [15] Another option is to use SYBR Green I, a DNA intercalating coloring agent. [16]

Advantages and disadvantages

This method is specifically advantageous because it can all be done quickly in one step. The sample is mixed with the primers, reverse transcriptase and DNA polymerase and the reaction takes place under a constant temperature. The required temperature can be achieved using a simple hot water bath.
PCR requires thermocycling; RT-LAMP does not, making it more time efficient and very cost effective. [3] This inexpensive and streamlined method can be more readily used in developing countries that do not have access to high tech laboratories.
A disadvantage of this method is generating the sequence specific primers. For each LAMP assay, primers must be specifically designed to be compatible with the target DNA. This can be difficult which discourages researchers from using the LAMP method in their work. [1] There is however, a free software called Primer Explorer, developed by Fujitsu in Japan, which can aid in the selection of these primers.
See also
References
- ^ a b c Mori Y, Notomi T (2009). "Loop-mediated isothermal amplification (LAMP): a rapid, accurate, and cost-effective diagnostic method for infectious diseases". J. Infect. Chemother. 15 (2): 62–9. doi: 10.1007/s10156-009-0669-9. PMC 7087713. PMID 19396514.
- ^ Notomi, Tsugunori; Okayama, Hiroto; Masubuchi, Harumi; Yonekawa, Toshihiro; Watanabe, Keiko; Amino, Nobuyuki; Hase, Tetsu (2000-06-15). "Loop-mediated isothermal amplification of DNA". Nucleic Acids Research. 28 (12): e63. doi: 10.1093/nar/28.12.e63. ISSN 0305-1048. PMC 102748. PMID 10871386.
- ^ a b Fu S, Qu G, Guo S, Ma L, Zhang N, Zhang S, Gao S, Shen Z (2011). "Applications of loop-mediated isothermal DNA amplification". Appl. Biochem. Biotechnol. 163 (7): 845–50. doi: 10.1007/s12010-010-9088-8. PMID 20844984. S2CID 45682156.
- ^ Habibzadeh, Parham; Mofatteh, Mohammad; Silawi, Mohammad; Ghavami, Saeid; Faghihi, Mohammad Ali (2021-02-17). "Molecular diagnostic assays for COVID-19: an overview". Critical Reviews in Clinical Laboratory Sciences. 58 (6): 385–398. doi: 10.1080/10408363.2021.1884640. ISSN 1549-781X. PMC 7898297. PMID 33595397.
- ^ Kurosaki, Yohei; Magassouba, N’Faly; Oloniniyi, Olamide K.; Cherif, Mahamoud S.; Sakabe, Saori; Takada, Ayato; Hirayama, Kenji; Yasuda, Jiro (2016-02-22). "Development and Evaluation of Reverse Transcription-Loop-Mediated Isothermal Amplification (RT-LAMP) Assay Coupled with a Portable Device for Rapid Diagnosis of Ebola Virus Disease in Guinea". PLOS Neglected Tropical Diseases. 10 (2): e0004472. doi: 10.1371/journal.pntd.0004472. ISSN 1935-2735. PMC 4764121. PMID 26900929.
- ^ "LAMP-based Test Could Enable Point-of-Care COVID-19 Testing". Diagnostics from Technology Networks. Retrieved 2020-07-31.
- ^ Alekseenko, Alisa; Barrett, Donal; Pareja-Sanchez, Yerma; Howard, Rebecca J.; Strandback, Emilia; Ampah-Korsah, Henry; Rovšnik, Urška; Zuniga-Veliz, Silvia; Klenov, Alexander; Malloo, Jayshna; Ye, Shenglong (2021-01-19). "Direct detection of SARS-CoV-2 using non-commercial RT-LAMP reagents on heat-inactivated samples". Scientific Reports. 11 (1): 1820. Bibcode: 2021NatSR..11.1820A. doi: 10.1038/s41598-020-80352-8. ISSN 2045-2322. PMC 7815738. PMID 33469065.
- ^ Lalli, Matthew A.; Langmade, S Joshua; Chen, Xuhua; Fronick, Catrina C.; Sawyer, Christopher S.; Burcea, Lauren C.; Wilkinson, Michael N.; Fulton, Robert S.; Heinz, Michael; Buchser, William J.; Head, Richard D.; Mitra, Robi D.; Milbrandt, Jeffrey (2020). "Rapid and extraction-free detection of SARS-CoV-2 from saliva with colorimetric LAMP". medRxiv 10.1101/2020.05.07.20093542.
- ^ Nagura-Ikeda, Mayu; Imai, Kazuo; Tabata, Sakiko; Miyoshi, Kazuyasu; Murahara, Nami; Mizuno, Tsukasa; Horiuchi, Midori; Kato, Kento; Imoto, Yoshitaka; Iwata, Maki; Mimura, Satoshi (2020-08-24). "Clinical Evaluation of Self-Collected Saliva by Quantitative Reverse Transcription-PCR (RT-qPCR), Direct RT-qPCR, Reverse Transcription–Loop-Mediated Isothermal Amplification, and a Rapid Antigen Test To Diagnose COVID-19". Journal of Clinical Microbiology. 58 (9). doi: 10.1128/JCM.01438-20. ISSN 0095-1137. PMC 7448663. PMID 32636214.
- ^ Vaidya NK, Wang FB, Zou X, Wahl LM (2012). "Transmission dynamics of the recently-identified BYD virus causing duck egg-drop syndrome". PLOS ONE. 7 (4): e35161. Bibcode: 2012PLoSO...735161V. doi: 10.1371/journal.pone.0035161. PMC 3329443. PMID 22529985.
- ^ Jiang T, Liu J, Deng YQ, Su JL, Xu LJ, Liu ZH, Li XF, Yu XD, Zhu SY, Gao GF, Qin ED, Qin CF (December 2012). "Development of RT-LAMP and real-time RT-PCR assays for the rapid detection of the new duck Tembusu-like BYD virus". Arch. Virol. 157 (12): 2273–80. doi: 10.1007/s00705-012-1431-7. PMID 22865206. S2CID 15573433.
- ^ Qiao J, Wang J, Meng Q, Wang G, Liu Y, He Z, Yang H, Zhang Z, Cai X, Chen C (2013). "Rapid detection of Akabane virus by a novel reverse transcription loop-mediated isothermal amplification assay (RT-LAMP)". Virol. J. 10: 288. doi: 10.1186/1743-422X-10-288. PMC 3848447. PMID 24034624.
- ^ Su, Chih-Wen; Li, Chiao-Yun; Lee, James Chun-I; Ji, Dar-Der; Li, Shu-Ying; Daniel, Barbara; Syndercombe-Court, Denise; Linacre, Adrian; Hsieh, Hsing-Mei (2015-06-01). "A novel application of real-time RT-LAMP for body fluid identification: using HBB detection as the model". Forensic Science, Medicine, and Pathology. 11 (2): 208–215. doi: 10.1007/s12024-015-9668-6. ISSN 1556-2891. PMID 25877518. S2CID 19288327.
- ^ Satoh, Tetsuya; Kouroki, Seiya; Ogawa, Keita; Tanaka, Yorika; Matsumura, Kazutoshi; Iwase, Susumu (2018-07-01). "Development of mRNA-based body fluid identification using reverse transcription loop-mediated isothermal amplification". Analytical and Bioanalytical Chemistry. 410 (18): 4371–4378. doi: 10.1007/s00216-018-1088-5. ISSN 1618-2650. PMID 29696299. S2CID 13823864.
- ^ a b Kellner, Max J.; Ross, James J.; Schnabl, Jakob; Dekens, Marcus P. S.; Heinen, Robert; Grishkovskaya, Irina; Bauer, Benedikt; Stadlmann, Johannes; Menéndez-Arias, Luis; Fritsche-Polanz, Robert; Traugott, Marianna (2020-07-23). "A rapid, highly sensitive and open-access SARS-CoV-2 detection assay for laboratory and home testing". bioRxiv: 2020.06.23.166397. doi: 10.1101/2020.06.23.166397. hdl: 10261/216969. S2CID 220835822.
- ^ Bokelmann, Lukas; Nickel, Olaf; Maricic, Tomislav; Paabo, Svante; Meyer, Matthias; Borte, Stephan; Riesenberg, Stephan (2020-08-06). "Rapid, reliable, and cheap point-of-care bulk testing for SARS-CoV-2 by combining hybridization capture with improved colorimetric LAMP (Cap-iLAMP)". medRxiv 10.1101/2020.08.04.20168617.