| Nucleocapsid protein | |||||||||
|---|---|---|---|---|---|---|---|---|---|
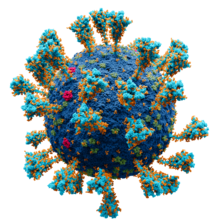 Model of the external structure of the SARS-CoV-2
virion.
[1] The N protein, contained entirely within the virion, is not visible.
● Blue: envelope ● Turquoise: spike glycoprotein (S) ● Red: envelope proteins (E) ● Green: membrane proteins (M) ● Orange: glycans | |||||||||
| Identifiers | |||||||||
| Symbol | CoV_nucleocap | ||||||||
| Pfam | PF00937 | ||||||||
| InterPro | IPR001218 | ||||||||
| |||||||||
The nucleocapsid (N) protein is a protein that packages the positive-sense RNA genome of coronaviruses to form ribonucleoprotein structures enclosed within the viral capsid. [2] [3] The N protein is the most highly expressed of the four major coronavirus structural proteins. [2] In addition to its interactions with RNA, N forms protein-protein interactions with the coronavirus membrane protein (M) during the process of viral assembly. [2] [3] N also has additional functions in manipulating the cell cycle of the host cell. [3] [4] The N protein is highly immunogenic and antibodies to N are found in patients recovered from SARS and COVID-19. [5]
History
COVID-19 was first identified in January 2020. A patient in the state of Washington was given a diagnosis of coronavirus infection on 20 January. A group of scientists based at the Centers for Disease Control and Prevention in Atlanta, Georgia isolated the virus from nasopharyngeal and oropharyngeal swabs and were able to characterize the genomic sequence, replication properties and cell culture tropism from the swabs. They made available the virus to the wider scientific community shortly thereafter "by depositing it into two virus reagent repositories". [6]
Structure
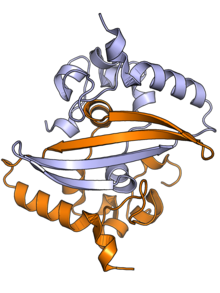
The N protein is composed of two main protein domains connected by an intrinsically disordered region (IDR) known as the linker region, with additional disordered segments at each terminus. [2] [3] A third small domain at the C-terminal tail appears to have an ordered alpha helical secondary structure and may be involved in the formation of higher-order oligomeric assemblies. [7] In SARS-CoV, the causative agent of SARS, the N protein is 422 amino acid residues long [2] and in SARS-CoV-2, the causative agent of COVID-19, it is 419 residues long. [7] [8]
Both the N-terminal and C-terminal domains are capable of binding RNA. The C-terminal domain forms a dimer that is likely to be the native functional state. [2] Parts of the IDR, particularly a conserved sequence motif rich in serine and arginine residues (the SR-rich region), may also be implicated in dimer formation, though reports on this vary. [2] [3] Although higher-order oligomers formed through the C-terminal domain have been observed crystallographically, it is unclear if these structures have a physiological role. [2] [9]
The C-terminal dimer has been structurally characterized by X-ray crystallography for several coronaviruses and has a highly conserved structure. [7] The N-terminal domain - sometimes known as the RNA-binding domain, though other parts of the protein also interact with RNA - has also been crystallized and has been studied by nuclear magnetic resonance spectroscopy in the presence of RNA. [10]
Post-translational modifications
The N protein is post-translationally modified by phosphorylation at sites located in the IDR, particularly in the SR-rich region. [2] [11] SARS-CoV-2 nucleocapsid (N) protein is arginine methylated by protein arginine methyltransferase 1 (PRMT1) at residues R95 and R177. Type I PRMT inhibitor (MS023) or substitution of R95 or R177 with lysine inhibited interaction of N protein with the 5’-UTR of SARS-CoV-2 genomic RNA, a property required for viral packaging | doi: 10.1016/j.jbc.2021.100821 | PMID 34029587. In several coronaviruses, ADP-ribosylation of the N protein has also been reported. [12] [11] With unclear functional significance, the SARS-CoV N protein has been observed to be SUMOylated and the N proteins of several coronaviruses including SARS-CoV-2 have been observed to be proteolytically cleaved. [11] [13] [14]
Expression and localization
The N protein is the most highly expressed in host cells of the four major structural proteins. [2] Like the other structural proteins, the gene encoding the N protein is located toward the 3' end of the genome. [3]
N protein is localized primarily to the cytoplasm. [3] In many coronaviruses, a population of N protein is localized to the nucleolus, [3] [4] [15] thought to be associated with its effects on the cell cycle. [4]
Function
Genome packaging and viral assembly

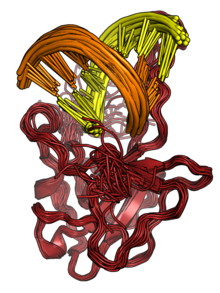
The N protein binds to RNA to form ribonucleoprotein (RNP) structures for packaging the genome into the viral capsid. [2] [3] The RNP particles formed are roughly spherical and are organized in flexible helical structures inside the virus. [2] [3] Formation of RNPs is thought to involve allosteric interactions between RNA and multiple RNA-binding regions of the protein. [2] [9] Dimerization of N is important for assembly of RNPs. Encapsidation of the genome occurs through interactions between N and M. [2] [3] N is essential for viral assembly. [3] N also serves as a chaperone protein for the formation of RNA structure in the genomic RNA. [3] [9]
Genomic and subgenomic RNA synthesis
Synthesis of genomic RNA appears to involve participation by the N protein. N is physically colocalized with the viral RNA-dependent RNA polymerase early in the replication cycle and forms interactions with non-structural protein 3, a component of the replicase-transcriptase complex. [3] Although N appears to facilitate efficient replication of genomic RNA, it is not required for RNA transcription in all coronaviruses. [3] [17] In at least one coronavirus, transmissible gastroenteritis virus (TGEV), N is involved in template switching in the production of subgenomic mRNAs, a process that is a distinctive feature of viruses in the order Nidovirales. [3] [17] [18]
Cell cycle effects
Coronaviruses manipulate the cell cycle of the host cell through various mechanisms. In several coronaviruses, including SARS-CoV, the N protein has been reported to cause cell cycle arrest in S phase through interactions with cyclin-CDK. [3] [4] In SARS-CoV, a cyclin box-binding region in the N protein can serve as a cyclin-CDK phosphorylation substrate. [3] Trafficking of N to the nucleolus may also play a role in cell cycle effects. [4] More broadly, N may be involved in reduction of host cell protein translation activity. [3]
Immune system effects
The N protein is involved in viral pathogenesis via its effects on components of the immune system. In SARS-CoV, [3] [19] [20] MERS-CoV, [21] and SARS-CoV-2, [22] N has been reported as suppressing interferon responses.
Evolution and conservation
The sequences and structures of N proteins from different coronaviruses, particularly the C-terminal domains, appear to be well conserved. [2] [7] [23] Similarities between the structure and topology of the N proteins of coronaviruses and arteriviruses suggest a common evolutionary origin and supports the classification of these two groups in the common order Nidovirales. [2] [3]
Examination of SARS-CoV-2 sequences collected during the COVID-19 pandemic found that missense mutations were most common in the central linker region of the protein, suggesting this relatively unstructured region is more tolerant of mutations than the structured domains. [7] A separate study of SARS-CoV-2 sequences identified at least one site in the N protein under positive selection. [24]
The N protein's properties of being well conserved, not appearing to recombine frequently, and producing a strong T-cell response have led to it being studied as a potential target for coronavirus vaccines. [25] [26] [23] [27] The vaccine candidate UB-612 is one such experimental vaccine that targets the N protein, along with other viral proteins, to attempt to induce broad immunity. [28] [29]
References
- ^ Solodovnikov, Alexey; Arkhipova, Valeria (2021-07-29). "Достоверно красиво: как мы сделали 3D-модель SARS-CoV-2" [Truly beautiful: how we made the SARS-CoV-2 3D model] (in Russian). N+1. Archived from the original on 2021-07-30. Retrieved 30 July 2021.
- ^ a b c d e f g h i j k l m n o p Chang, Chung-ke; Hou, Ming-Hon; Chang, Chi-Fon; Hsiao, Chwan-Deng; Huang, Tai-huang (March 2014). "The SARS coronavirus nucleocapsid protein – Forms and functions". Antiviral Research. 103: 39–50. doi: 10.1016/j.antiviral.2013.12.009. PMC 7113676. PMID 24418573.
- ^ a b c d e f g h i j k l m n o p q r s t u McBride, Ruth; van Zyl, Marjorie; Fielding, Burtram (7 August 2014). "The Coronavirus Nucleocapsid Is a Multifunctional Protein". Viruses. 6 (8): 2991–3018. doi: 10.3390/v6082991. PMC 4147684. PMID 25105276.
- ^ a b c d e Su, Mingjun; Chen, Yaping; Qi, Shanshan; Shi, Da; Feng, Li; Sun, Dongbo (5 November 2020). "A Mini-Review on Cell Cycle Regulation of Coronavirus Infection". Frontiers in Veterinary Science. 7: 586826. doi: 10.3389/fvets.2020.586826. PMC 7674852. PMID 33251267.
- ^ Li, Dandan; Li, Jinming (20 April 2021). "Immunologic Testing for SARS-CoV-2 Infection from the Antigen Perspective". Journal of Clinical Microbiology. 59 (5): e02160-20. doi: 10.1128/JCM.02160-20. PMC 8091849. PMID 33318065.
- ^ Harcourt, Jennifer; et al. (2020). "Severe Acute Respiratory Syndrome Coronavirus 2 from Patient with Coronavirus Disease, United States". Emerging Infectious Diseases. 26 (6): 1266–1273. doi: 10.3201/eid2606.200516. PMC 7258473. PMID 32160149.
- ^ a b c d e f Ye, Qiaozhen; West, Alan M. V.; Silletti, Steve; Corbett, Kevin D. (September 2020). "Architecture and self‐assembly of the SARS‐CoV ‐2 nucleocapsid protein". Protein Science. 29 (9): 1890–1901. doi: 10.1002/pro.3909. PMC 7405475. PMID 32654247.
- ^ Shah, Vibhuti Kumar; Firmal, Priyanka; Alam, Aftab; Ganguly, Dipyaman; Chattopadhyay, Samit (7 August 2020). "Overview of Immune Response During SARS-CoV-2 Infection: Lessons From the Past". Frontiers in Immunology. 11: 1949. doi: 10.3389/fimmu.2020.01949. PMC 7426442. PMID 32849654.
- ^ a b c Chang, Chung-ke; Lo, Shou-Chen; Wang, Yong-Sheng; Hou, Ming-Hon (April 2016). "Recent insights into the development of therapeutics against coronavirus diseases by targeting N protein". Drug Discovery Today. 21 (4): 562–572. doi: 10.1016/j.drudis.2015.11.015. PMC 7108309. PMID 26691874.
- ^ a b Dinesh, Dhurvas Chandrasekaran; Chalupska, Dominika; Silhan, Jan; Koutna, Eliska; Nencka, Radim; Veverka, Vaclav; Boura, Evzen (2 December 2020). "Structural basis of RNA recognition by the SARS-CoV-2 nucleocapsid phosphoprotein". PLOS Pathogens. 16 (12): e1009100. doi: 10.1371/journal.ppat.1009100. PMC 7735635. PMID 33264373.
- ^ a b c Fung, To Sing; Liu, Ding Xiang (June 2018). "Post-translational modifications of coronavirus proteins: roles and function". Future Virology. 13 (6): 405–430. doi: 10.2217/fvl-2018-0008. PMC 7080180. PMID 32201497.
- ^ Grunewald, Matthew E.; Fehr, Anthony R.; Athmer, Jeremiah; Perlman, Stanley (April 2018). "The coronavirus nucleocapsid protein is ADP-ribosylated". Virology. 517: 62–68. doi: 10.1016/j.virol.2017.11.020. PMC 5871557. PMID 29199039.
- ^ Lutomski, Corinne A.; El-Baba, Tarick J.; Bolla, Jani R.; Robinson, Carol V. (2021-08-23). "Multiple Roles of SARS-CoV-2 N Protein Facilitated by Proteoform-Specific Interactions with RNA, Host Proteins, and Convalescent Antibodies". JACS Au. 1 (8): 1147–1157. doi: 10.1021/jacsau.1c00139. ISSN 2691-3704. PMC 8231660. PMID 34462738.
- ^ Meyer, Bjoern; Chiaravalli, Jeanne; Gellenoncourt, Stacy; Brownridge, Philip; Bryne, Dominic P.; Daly, Leonard A.; Grauslys, Arturas; Walter, Marius; Agou, Fabrice; Chakrabarti, Lisa A.; Craik, Charles S. (December 2021). "Characterising proteolysis during SARS-CoV-2 infection identifies viral cleavage sites and cellular targets with therapeutic potential". Nature Communications. 12 (1): 5553. Bibcode: 2021NatCo..12.5553M. doi: 10.1038/s41467-021-25796-w. ISSN 2041-1723. PMC 8455558. PMID 34548480.
- ^ Masters, Paul S. (2006). "The Molecular Biology of Coronaviruses". Advances in Virus Research. 66: 193–292. doi: 10.1016/S0065-3527(06)66005-3. ISBN 9780120398690. PMC 7112330. PMID 16877062.
- ^ Goodsell, David S.; Voigt, Maria; Zardecki, Christine; Burley, Stephen K. (6 August 2020). "Integrative illustration for coronavirus outreach". PLOS Biology. 18 (8): e3000815. doi: 10.1371/journal.pbio.3000815. PMC 7433897. PMID 32760062.
- ^ a b Zúñiga, Sonia; Cruz, Jazmina L. G.; Sola, Isabel; Mateos-Gómez, Pedro A.; Palacio, Lorena; Enjuanes, Luis (15 February 2010). "Coronavirus Nucleocapsid Protein Facilitates Template Switching and Is Required for Efficient Transcription". Journal of Virology. 84 (4): 2169–2175. doi: 10.1128/JVI.02011-09. PMC 2812394. PMID 19955314.
- ^ Sola, Isabel; Almazán, Fernando; Zúñiga, Sonia; Enjuanes, Luis (9 November 2015). "Continuous and Discontinuous RNA Synthesis in Coronaviruses". Annual Review of Virology. 2 (1): 265–288. doi: 10.1146/annurev-virology-100114-055218. PMC 6025776. PMID 26958916.
- ^ Spiegel, Martin; Pichlmair, Andreas; Martínez-Sobrido, Luis; Cros, Jerome; García-Sastre, Adolfo; Haller, Otto; Weber, Friedemann (15 February 2005). "Inhibition of Beta Interferon Induction by Severe Acute Respiratory Syndrome Coronavirus Suggests a Two-Step Model for Activation of Interferon Regulatory Factor 3". Journal of Virology. 79 (4): 2079–2086. doi: 10.1128/JVI.79.4.2079-2086.2005. PMC 546554. PMID 15681410.
- ^ Kopecky-Bromberg, Sarah A.; Martínez-Sobrido, Luis; Frieman, Matthew; Baric, Ralph A.; Palese, Peter (15 January 2007). "Severe Acute Respiratory Syndrome Coronavirus Open Reading Frame (ORF) 3b, ORF 6, and Nucleocapsid Proteins Function as Interferon Antagonists". Journal of Virology. 81 (2): 548–557. doi: 10.1128/JVI.01782-06. PMC 1797484. PMID 17108024.
- ^ Chang, Chi-You; Liu, Helene Minyi; Chang, Ming-Fu; Chang, Shin C. (16 June 2020). "Middle East Respiratory Syndrome Coronavirus Nucleocapsid Protein Suppresses Type I and Type III Interferon Induction by Targeting RIG-I Signaling". Journal of Virology. 94 (13): e00099-20. doi: 10.1128/JVI.00099-20. PMC 7307178. PMID 32295922.
- ^ Mu, Jingfang; Fang, Yaohui; Yang, Qi; Shu, Ting; Wang, An; Huang, Muhan; Jin, Liang; Deng, Fei; Qiu, Yang; Zhou, Xi (December 2020). "SARS-CoV-2 N protein antagonizes type I interferon signaling by suppressing phosphorylation and nuclear translocation of STAT1 and STAT2". Cell Discovery. 6 (1): 65. doi: 10.1038/s41421-020-00208-3. PMC 7490572. PMID 32953130.
- ^ a b Nikolaidis, Marios; Markoulatos, Panayotis; Van de Peer, Yves; Oliver, Stephen G; Amoutzias, Grigorios D (2021-10-12). Hepp, Crystal (ed.). "The neighborhood of the Spike gene is a hotspot for modular intertypic homologous and non-homologous recombination in Coronavirus genomes". Molecular Biology and Evolution. 39: msab292. doi: 10.1093/molbev/msab292. ISSN 0737-4038. PMC 8549283. PMID 34638137.
- ^ Cagliani, Rachele; Forni, Diego; Clerici, Mario; Sironi, Manuela (June 2020). "Computational Inference of Selection Underlying the Evolution of the Novel Coronavirus, Severe Acute Respiratory Syndrome Coronavirus 2". Journal of Virology. 94 (12): e00411-20. doi: 10.1128/JVI.00411-20. PMC 7307108. PMID 32238584.
- ^ Grifoni, Alba; Weiskopf, Daniela; Ramirez, Sydney I.; Mateus, Jose; Dan, Jennifer M.; Moderbacher, Carolyn Rydyznski; Rawlings, Stephen A.; Sutherland, Aaron; Premkumar, Lakshmanane; Jadi, Ramesh S.; Marrama, Daniel (June 2020). "Targets of T Cell Responses to SARS-CoV-2 Coronavirus in Humans with COVID-19 Disease and Unexposed Individuals". Cell. 181 (7): 1489–1501.e15. doi: 10.1016/j.cell.2020.05.015. PMC 7237901. PMID 32473127.
- ^ Dutta, Noton K.; Mazumdar, Kaushiki; Gordy, James T. (2020-06-16). Dutch, Rebecca Ellis (ed.). "The Nucleocapsid Protein of SARS–CoV-2: a Target for Vaccine Development". Journal of Virology. 94 (13). doi: 10.1128/JVI.00647-20. ISSN 0022-538X. PMC 7307180. PMID 32546606.
- ^ Hajnik RL, Plante JA, Liang Y, Alameh MG, Tang J, Bonam SR, Zhong C, Adam A, Scharton D, Rafael GH, Liu Y, Hazell NC, Sun J, Soong L, Shi PY, Wang T, Walker DH, Sun J, Weissman D, Weaver SC, Plante KS, Hu H (September 2022). "Dual spike and nucleocapsid mRNA vaccination confer protection against SARS-CoV-2 Omicron and Delta variants in preclinical models". Science Translational Medicine. 14 (662): eabq1945. doi: 10.1126/scitranslmed.abq1945. PMC 9926941. PMID 36103514. S2CID 252283038.
- ^ Martínez-Flores D, Zepeda-Cervantes J, Cruz-Reséndiz A, Aguirre-Sampieri S, Sampieri A, Vaca L (2021). "SARS-CoV-2 Vaccines Based on the Spike Glycoprotein and Implications of New Viral Variants". Frontiers in Immunology. 12: 701501. doi: 10.3389/fimmu.2021.701501. PMC 8311925. PMID 34322129.
- ^ Wang CY, Hwang KP, Kuo HK, Peng WJ, Shen YH, Kuo BS, Huang JH, Liu H, Ho YH, Lin F, Ding S, Liu Z, Wu HT, Huang CT, Lee YJ, Liu MC, Yang YC, Lu PL, Tsai HC, Lee CH, Shi ZY, Liu CE, Liao CH, Chang FY, Chen HC, Wang FD, Hou KL, Cheng J, Wang MS, Yang YT, Chiu HC, Jiang MH, Shih HY, Shen HY, Chang PY, Lan YR, Chen CT, Lin YL, Liang JJ, Liao CC, Chou YC, Morris MK, Hanson CV, Guirakhoo F, Hellerstein M, Yu HJ, King CC, Kemp T, Heppner DG, Monath TP (May 2022). "A multitope SARS-CoV-2 vaccine provides long-lasting B cell and T cell immunity against Delta and Omicron variants". Journal of Clinical Investigation. 132 (10). doi: 10.1172/JCI157707. PMC 9106357. PMID 35316221.
